Abstract
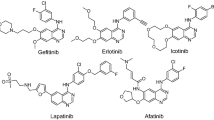
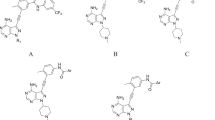
The overexpression and/or mutation of the epidermal growth factor receptor (EGFR) tyrosine kinase has been observed in many human solid tumors, and is under intense investigation as a novel anticancer molecular target. Comparative 3D-QSAR analyses using different alignments were undertaken employing comparative molecular field analysis (CoMFA) and comparative molecular similarity analysis (CoMSIA) for 122 anilinoquinazoline and 50 anilinoquinoline inhibitors of EGFR kinase. The SYBYL multifit alignment rule was applied to three different conformational templates, two obtained from a MacroModel Monte Carlo conformational search, and one from the bound conformation of erlotinib in complex with EGFR in the X-ray crystal structure. In addition, a flexible ligand docking alignment obtained with the GOLD docking program, and a novel flexible receptor-guided consensus dynamics alignment obtained with the DISCOVER program in the INSIGHTII modeling package were also investigated. 3D-QSAR models with q2 values up to 0.70 and r2 values up to 0.97 were obtained. Among the 4-anilinoquinazoline set, the q2 values were similar, but the ability of the different conformational models to predict the activities of an external test set varied considerably. In this regard, the model derived using the X-ray crystallographically determined bioactive conformation of erlotinib afforded the best predictive model. Electrostatic, hydrophobic and H-bond donor descriptors contributed the most to the QSAR models of the 4-anilinoquinazolines, whereas electrostatic, hydrophobic and H-bond acceptor descriptors contributed the most to the 4-anilinoquinoline QSAR, particularly the H-bond acceptor descriptor. A novel receptor-guided consensus dynamics alignment has also been introduced for 3D-QSAR studies. This new alignment method may incorporate to some extent ligand-receptor induced fit effects into 3D-QSAR models.
Similar content being viewed by others
References
Hong, W.K. and Ullrich, A., Oncol. Biotherap. 1 (2000) 1.
Salomon, D.S., Brandt, R., Cardiello, F. and Normanno, N., Crit. Rev. Oncol. Hematol. 19 (1995) 183.
Woodburn, J.R., Pharmacol. Ther., 82 (1999) 241.
Gosh, S., Liu, X.-P., Zheng, Y. and Uckun, F.M., Curr. Cancer Drug Targets, 1 (2001) 129.
Mendelsohn, J. and Baselga, J., Oncogene, 19 (2000) 6550.
Rewcastle, G.W., Palmer, B.D., Bridges, A.J., Showalter, H.D.H., Sun, L., Nelson, J., McMichael, A., Kraker, A.J., Fry, D.W. and Denny, W.A., J. Med. Chem., 39 (1996) 918.
Bridges, A.J., Zhou, H., Cody, D.R., Rewcastle, G.W., McMichael, A., Showalter, H.D.H., Fry, D.W., Kraker, A.J. and Denny, W.A., J. Med. Chem., 39 (1996) 267.
Rewcastle, G.W., Denny, W.A., Bridges, A.J., Zhou, H., Cody, D.R., McMichael, A. and Fry, D.W., J. Med. Chem., 38 (1995) 3482.
Palmer, B.D., Trumpp-Kallmeyer, S., Fry, D.W., Nelson, J.M., Showalter, H.D. and Denny, W.A., J. Med. Chem., 40 (1997) 1519.
Baselga, J. and Averbuch, S.D., Drugs, 60 (Suppl. 1) (2000) 33.
Ciardiello, F., Caputo, R., Bianco, R. et al., Clin. Cancer Res., 6 (2000) 2053.
de Bono, J.S. and Rowinsky, E.K., Trend Mol. Med., 8 (2002) S19.
Stamos, J., Sliwkowski, M.X. and Eigenbrot, C.J., Biol. Chem., 277 (2002), 46265.
Cramer III, R.D., Patterson, D.E. and Bunce, J.D., J. Am. Chem. Soc., 110 (1988) 5959.
Klebe, G., Abraham, U. and Mietzner, T.J., J. Med. Chem., 37 (1994) 4130.
Wissner, A., Berger, D.M., Boschelli, D.H., Floyd, M.B. Jr., Greenberger, L.M., Graber, B.C., Johnson, B.D., Mamuya, N., Nilakantan, R., Reich, M.F., Shen, R., Tsou, H.R., Upeslacis, E., Wang, Y.F., Wu, B., Ye, F. and Zhang, N., J. Med. Chem., 43 (2000) 3244.
Clark, M., Cramer III, R.D. and Van Opdenbosch, N., J. Comput. Chem., 10 (1989) 982.
Stewart, J.J., J. Comput.-Aided Mol. Design, 4 (1990) 1.
Jones, G., Willet, P., Glen, R.C., Leach, A.R. and Taylor, R., J. Mol. Biol., 267 (1997) 727.
Kamath, S. and Coutinho, E., J. Biosci., 22 (1997) 315.
Viswanadhan, V.N., Ghoise, A.K., Revenkar, G.R. and Robins, R., J. Chem. Inf. Comput. Sci., 29 (1989) 163.
Klebe, G., J. Mol. Biol., 237 (1994) 212.
Buolamwini, J.K., Raghavan, K., Fesen, M.R., Pommier, Y., Kohn, K.W. and Weinstein J.N., Pharm. Res., 13 (1996) 1891.
Buolamwini, J.K. and Assefa, H., CoMFA and CoMSIA 3D-QSAR Studies of Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitors. 221st ACS National Meeting, San Diego, CA, April 1-5, 2001.
Bohm, M., Sturzebecher, J. and Klebe, G., J. Med. Chem., 42 (1999) 458.
Bohm, M. and Klebe, G., J. Med. Chem., 45 (2002) 1585.
Pierce, A.C., Sandretto, K.L. and Bemis G.W., Proteins, 49 (2001) 567.
Vedani, A. and Dobbler, M., J. Med. Chem., 45 (2002) 2139.
Datar, P., Desai, P., Coutinho, E. and Iyer, K.J., J.Mol.Model., 8 (2002) 290.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Assefa, H., Kamath, S. & Buolamwini, J.K. 3D-QSAR and docking studies on 4-anilinoquinazoline and 4-anilinoquinoline epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors. J Comput Aided Mol Des 17, 475–493 (2003). https://doi.org/10.1023/B:JCAM.0000004622.13865.4f
Issue Date:
DOI: https://doi.org/10.1023/B:JCAM.0000004622.13865.4f