Abstract
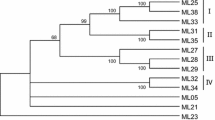
Sequence analyses of numerous plant disease resistance genes have revealed the presence of conserved motifs common to this class of genes, namely a nucleotide binding site (NBS) and leucine rich repeat region. In this study, thirty-three resistance gene analogs (RGAs) were cloned and sequenced from cotton (Gossypium hirsutum L.) following PCR with degenerate primers designed from the conserved NBS motif of plant resistance (R) genes. Phylogenetic analysis of the predicted amino acid sequences grouped the RGAs into four distinct classes from which several subgroups were delineated based on nucleic acid sequences. Gene database searches with the consensus protein sequences of each of the four classes and respective subgroups of cotton RGAs revealed their conserved NBS domains and homology to RGAs and known resistance genes from a variety of plant genera. Given the complete lack of knowledge regarding molecular organization of R genes in cotton, the cloned RGAs described here may be useful as probes to map, characterize, and manipulate R genes of the cotton genome.
Similar content being viewed by others
References
Altschul, S.F., T.L. Madden, A.A. Schäffer, J. Zhang, Z. Zhang, W. Miller & D.J. Lipman, 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nuc Acids Res 25: 3389-3402.
Baker, B., P. Zambryski, B. Staskawicz & S.P. Dinesh-Kumar, 1997. Signaling in plant-microbe interactions. Science 276: 726-733.
Bent, A.F., 1996. Plant disease resistance genes: function meets structure. Plant Cell 8: 1757-1771.
Bent, A.F., B.N. Kundel, D. Dahlbeck, K.L. Brown, R. Schmidt, J. Giraudat, J. Leung & B.J. Staskawicz, 1994. RPS2 of Arabdopsis thaliana: a leucine-rich repeat class of plant disease resistance genes. Sci 265: 1856-1860.
Collins, N.C., C.A. Webb, S. Seah, J.G. Ellis, S.H. Hulbert & A. Pryor, 1998. The isolation and mapping of disease resistance gene analogs in maize. Mol Plant-Microbe Interact 11: 968-978.
Dangl, J.L., R.A. Dietrich & M.H. Richberg, 1996. Death don't have no mercy: cell death programs in plant-microbe interactions. Plant Cell 8: 1793-1807.
Deng, Z., S. Huang, P. Ling, C. Chen, C. Yu, C.A. Weber, G.A. Moore & F.G. Gmitter Jr., 2000. Cloning and characterization of NBS-LRR class resistance-gene candidate sequences in citrus. Theor Appl Genet 101: 814-822.
Di Gaspero, G. & G. Cipriani, 2002. Resistance gene analogs are candidate markers for disease-resistance genes in grape (Vitis spp.). Theor Appl Genet 106: 163-172.
Dixon, M.S., D.A. Jones, J.S. Keddie, C.M. Thomas, K. Harrison & J.D.G. Jones, 1996. The tomato Cf-2 disease resistance locus comprises two functional genes encoding leucine-rich repeat proteins. Cell 84: 451-459.
Dodds, P.N., G.J. Lawrence & J.G. Ellis, 2001. Six amino acid changes confined to the leucine-rich repeat beta-strand/betaturn motif determine the difference between the P and P2 rust resistance specificities in flax. Plant Cell 13: 163-178.
Ellis, J., P. Dodds & T. Pryor, 2000. Structure, function and evolution of plant disease resistance genes. Curr Opinion Plant Biol 3: 278-284.
Hammond-Kosack, K.E. & J.D.G. Jones, 1997. Plant disease resistance genes. Annu Rev Plant Physiol Plant Mol Biol 48: 575-607.
Joyeux, A., M.G. Fortin, R. Mayerhofer & A.G. Good, 1999. Genetic mapping of plant disease resistance gene homologues using a minimal Brassica napus L. population. Genome 42: 735-743.
Kanazin, V., L.F. Marek & R.C. Shoemaker, 1996. Resistance gene analogs are conserved and clustered in soybean. Proc Natl Acad Sci USA 93: 11746-11750.
Karlin, S. & S.F. Altschul, 1990. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc Natl Acad Sci USA 87: 2264-2268.
Kobe, B. & J. Deisenhofer, 1994. The leucine-rich repeat: a versatile binding motif. Trends Biochem Sci 19: 415-421.
Lawrence, G.J., E.J. Finnegan, M.A. Ayliffe & J.G. Ellis, 1995. The L6 gene for flax rust resistance is related to the Arabidopsis bacterial resistance gene RPS2 and the tobacco viral resistance gene N. Plant Cell 7: 1195-1206.
Leister, D., A. Ballvora, F. Salamini & C. Gebhardt, 1996. A PCR based approach for isolating pathogen resistance genes from potato with potential for wide application in plants. Nature Genet 14: 421-429.
Meyers, B.C., D.B. Chin, K.A. Shen, S. Sivaramakrishnan, D.O. Lavelle, Z. Zhang & R.W. Michelmore, 1998. The major resistance gene cluster in lettuce is highly duplicated and spans several megabases. Plant Cell 10: 1817-1832.
Meyers, B.C., A.W. Dickerman, R.W. Michelmore, S. Sivaramakrishnan, B.W. Sobral & N.D. Young, 1999. Plant disease resistance genes encode members of an ancient and diverse protein family within the nucleotide-binding superfamily. Plant J 20: 317-332.
Mindrinos, M., F. Katagiri, G.L. Yu & F.M. Ausubel, 1994. The A. thaliana disease resistance gene RPS2 encodes a protein containing a nucleotide-binding site and leucine-rich repeats. Cell 78: 1089-1099.
Ori, N., Y. Eshed, I. Paran, G. Presting, D. Aviv, S. Tanksley, D. Zamir & R. Fluhr, 1997. The I2C family from the wilt disease resistance locus 12 belongs to the nucleotide binding, leucine-rich repeat superfamily of plant resistance genes. Plant Cell 9: 521-532.
Pan, Q., J. Wendel & R. Fluhr, 2000. Divergent evolution of plant NBS-LRR resistance gene homologues in dicot and cereal genomes. J Mol Evol 50: 203-213.
Parker, J.E., M.J. Coleman, V. Szabo, L.N. Frost, R. Schmidt, E.A. van der Biezen, T. Moores, C. Dean, M.J. Daniels & J.D.G. Jones, 1997. The Arabidopsis downy mildew resistance gene RPP5 shares similarity to the toll and interleukin-1 receptors with N and L6. Plant Cell 9: 879-894.
Paterson, A.H., C.L. Brubaker & J.F. Wendel, 1993. A rapid method for extraction of cotton (Gossypium sp) genomic DNA suitable for RFLP or PCR analysis. Plant Mol Biol Report 11: 122-127.
Rivkin, M.I., C.E. Vallejos & P.E. McClean, 1999. Disease-resistance related sequences in common bean. Genome 42: 41-47.
Saitou, N. & M. Nei, 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406-425.
Scofield, S.R., C.M. Tobias, J.P. Rathjen, J.H. Chang, D.T. Lavelle, R.W. Michelmore & B.J. Staskawicz, 1996. Molecular basis of gene-for-gene specificity in bacterial speck disease of tomato. Sci 274: 2063-2065.
Seah, S., H. Bariana, J. Jahier, K. Sivasithamparam & E.S. Lagudah, 2001. The introgressed segment carrying rust resistance genes Yr17, Lr37 and Sr38 in wheat can be assayed by a cloned disease resistance gene-like sequence. Theor Appl Genet 102: 600-605.
Seah, S., K. Sivasithamparam, A. Karakousis & E.S. Lagudah, 1998. Cloning and characterisation of a family of disease resistance gene analogs from wheat and barley. Theor Appl Genet 97: 937-945.
Shen, K.A., B.C. Meyers, M.N. Islam-Faridi, D.B. Chin, D.M. Stelly & R.W. Michelmore, 1998. Resistance gene candidates identified by PCR with degenerate oligonucleotide primers map to clusters of resistance genes in lettuce. Mol Plant-Microbe Interact 11: 815-823.
Shepherd, R.L., 1974. Transgressive segregation for root-knot nematode resistance in cotton. Crop Sci 14: 872-875.
Shepherd, R.L., J.C. McCarty, J.N. Jenkins & W.L. Parrott, 1996. Registration of nine cotton germplasm lines resistant to root-knot nematode. Crop Sci 36: 820.
Shepherd, R.L., J.C. McCarty, W.L. Parrott & J.N. Jenkins, 1988. Resistance of cotton cultivars and elite breeding lines to root-knot nematode. Mississippi Agricultural and Forestry Exp Stn Tech Bulletin 158. Mississippi State University, Mississippi State, MS.
Simons, G., J. Groenendijik, J. Wijbrandi, M. Reijans, J. Groenen, P. Diergaarde, T. Van der Lee, M. Bleeker, J. Onstenk, M. de Both, M. Haring, J. Mes, B. Cornelissen, M. Zabeau & P. Vos, 1998. Dissection of the fusarium I2 gene cluster in tomato reveals six homologs and one active gene copy. Plant Cell 10: 1055-1068.
Speulman, E., D. Bouchez, E.B. Holub & J.L. Beynon, 1998. Disease resistance gene homologs correlate with disease resistance loci of Arabidopsis thaliana. Plant J 14: 467-474.
Staskawicz, B.J., F.M. Ausubel, B.J. Baker, J.G. Ellis & J.D.G. Jones, 1995. Molecular genetics of plant disease resistance. Sci 268: 661-667.
Whitham, S., S.P. Dinesh-Kumar, D. Choi, R. Hehl, C. Corr & B. Baker, 1994. The product of the tobacco mosaic virus resistance gene N: similarity to Toll and the Interleukin-1 receptor. Cell 78: 1101-1115.
Yu, Y.G., G.R. Buss & M.A. Saghai Maroof, 1996. Isolation of a superfamily of candidate disease-resistance genes in soybean based on a conserved nucleotide-binding site. Proc Natl Acad Sci USA 93: 11751-11756.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Tan, H., Callahan, F., Zhang, XD. et al. Identification of resistance gene analogs in cotton (Gossypium hirsutum L.). Euphytica 134, 1–7 (2003). https://doi.org/10.1023/A:1026114327168
Issue Date:
DOI: https://doi.org/10.1023/A:1026114327168