Abstract
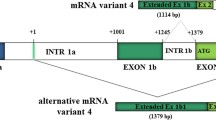
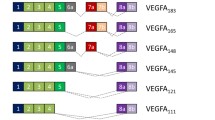
In order to characterise the candidate α2,3-sialyltransferases necessary for biosynthesis of the selectin ligand SLex and related antigens we have cloned and sequenced, from peripheral blood leukocytes of single individuals, various transcripts from the human ST3Gal III, IV and VI genes. Our clones have revealed a considerable heterogeneity in transcript isoforms. Among our ST3Gal IV clones we isolated nine alternatively spliced transcripts covering the coding region of the human ST3Gal IV gene (A1, A1 − 12, A1 + 18, A2, A2 − 12, A2 + 18, B, B − 12 and B + 18). Five of these isotranscripts A1 − 12, A1 + 18, A2 − 12, A2 + 18 and B + 18 have not been described before. In order to investigate if the alternatively spliced isotranscripts were specific for human PBL, we analysed the expression by RT-PCR and laser-induced fluorescent capillary electrophoresis (LIF-CE) in twenty other human tissues. We found a tissue specific expression of ST3Gal IV A1, A1 − 12, A1 + 18, A2, A2 − 12, A2 + 18 and B + 18 as well as a general expression of ST3Gal IV B and B − 12 isotranscripts in all tissues examined.
Similar content being viewed by others
References
Phillips ML, Nudelman E, Gaeta FC, Perez M, Singhal AK, Hakomori S, Paulson JC, ELAM-1 mediates cell adhesion by recognition of a carbohydrate ligand, sialyl-Lex, Science 250, 1130–2 (1990).
Polley MJ, Phillips ML, Wayner E, Nudelman E, Singhal AK, Hakomori S, Paulson JC, CD62 and endothelial cell-leukocyte adhesion molecule 1 (ELAM-1) recognize the same carbohydrate ligand, sialyl-Lewis x, Proc Natl Acad Sci USA 88, 6224–8 (1991).
Tyrrell D, James P, Rao N, Foxall C, Abbas S, Dasgupta F, Nashed M, Hasegawa A, Kiso M, Asa D, et al., Structural requirements for the carbohydrate ligand of E-selectin, Proc Natl Acad Sci USA 88, 10372–6 (1991).
Springer TA, Traffic signals for lymphocyte recirculation and leukocyte emigration: The multistep paradigm, Cell 76, 301–14 (1994).
Varki A, Selectin ligands, Proc Natl Acad Sci USA 91, 7390–7 (1994).
Lasky LA, Selectin-carbohydrate interactions and the initiation of the inflammatory response, Annu Rev Biochem 64, 113–39 (1995).
Kansas GS, Selectins and their ligands: Current concepts and controversies, Blood 88, 3259–87 (1996).
Modrek B, Resch A, Grasso C, Lee C, Genome-wide detection of alternative splicing in expressed sequences of human genes, Nucleic Acids Res 29, 2850–9 (2001).
Brett D, Hanke J, Lehmann G, Haase S, Delbruck S, Krueger S, Reich J, Bork P, EST comparison indicates 38% of humanmRNAs contain possible alternative splice forms, FEBS Lett 474, 83–6 (2000).
Sasaki K, Watanabe E, Kawashima K, Sekine S, Dohi T, Oshima M, Hanai N, Nishi T, Hasegawa M, Expression cloning of a novel Gal beta (1–3/1–4) GlcNAc alpha 2,3-sialyltransferase using lectin resistance selection, J Biol Chem 268, 22782–7 (1993).
Kitagawa H, Paulson JC, Cloning of a novel alpha 2,3-sialyltransferase that sialylates glycoprotein and glycolipid carbohydrate groups, J Biol Chem 269, 1394–401 (1994).
Kitagawa H, Mattei MG, Paulson JC, Genomic organization and chromosomal mapping of the Gal beta 1,3GalNAc/Gal beta 1,4GlcNAc alpha 2,3-sialyltransferase, J Biol Chem 271, 931–8 (1996).
Taniguchi A, Matsumoto K, Down-regulation of human sialyltransferase gene expression during in vitro human keratinocyte cell line differentiation, Biochem Biophys Res Commun 243, 177–83 (1998).
Hanauer A, Mandel JL, The glyceraldehyde 3 phosphate dehydrogenase gene family: Structure of a human cDNA and of an X chromosome linked pseudogene; amazing complexity of the gene family in mouse, Embo J 3, 2627–33 (1984).
Ercolani L, Florence B, Denaro M, Alexander M, Isolation and complete sequence of a functional human glyceraldehyde-3-phosphate dehydrogenase gene, J Biol Chem 263, 15335–41 (1988).
Gassen U, Kelm S, Schauer R, Differential gene expression of a human alpha 2,3-sialyltransferase in leukaemic cell lines and leucocytes, FEBS Lett 427, 91–5 (1998).
Kitagawa H, Paulson JC, Differential expression of five sialyltransferase genes in human tissues, J Biol Chem 269, 17872–8 (1994).
Murata T, Takizawa T, Funaba M, Fujimura H, Murata E, Torii K, Quantitation of mouse and rat beta-actin mRNA by competitive polymerase chain reaction using capillary electrophoresis, Anal Biochem 244, 172–4 (1997).
Bor MV, Sorensen BS, Nexo E, Simultaneous quantitation of several mRNA species by calibrated reverse transcription polymerase chain reaction and capillary electrophoresis: Analysis of the epidermal growth factor receptor and its activating ligands EGF, TGFalpha, and HB-EGF in rat liver, Lab Invest 80, 983–6 (2000).
Personett DA, Chouinard M, Sugaya K, McKinney M, Simplified RT/PCR quantitation of gene transcripts in cultured neuroblastoma (SN49) and microglial (BV-2) cells using capillary electrophoresis and laser-induced fluorescence, J Neurosci Methods 65, 77–91 (1996).
Shammas FV, Van Eekelen JA, Wee L, Heikkila R, Osland A, Sensitive and quantitative one-step polymerase chain reaction using capillary electrophoresis and fluorescence detection for measuring cytokeratin 19 expression, Scand J Clin Lab Invest 59, 635–42 (1999).
Moeseneder MM, Arrieta JM, Muyzer G, Winter C, Herndl GJ, Optimization of terminal-restriction fragment length polymorphism analysis for complex marine bacterioplankton communities and comparison with denaturing gradient gel electrophoresis, Appl Environ Microbiol 65, 3518–25 (1999).
Odin E, Wettergren Y, Larsson L, Larsson PA, Gustavsson B, Rapid method for relative gene expression determination in human tissues using automated capillary gel electrophoresis and multicolor detection, J Chromatogr B Biomed Sci Appl 734, 47–53 (1999).
van Eekelen JA, Shammas FV, Wee L, Heikkila R, Osland A, Quantitative analysis of cytokeratin 20 gene expression using RTPCRand capillary electrophoresis with fluorescentDNA detection, Clin Biochem 33, 457–64 (2000).
Richards MP, Ashwell CM, McMurtry JP, Quantitative analysis of leptin mRNA using competitive reverse transcription polymerase chain reaction and capillary electrophoresis with laser-induced fluorescence detection, Electrophoresis 21, 792–8 (2000).
Smith CW, Valcarcel J, Alternative pre-mRNA splicing: The logic of combinatorial control, Trends Biochem Sci 25, 381–8 (2000).
Reed R, Initial splice-site recognition and pairing during premRNA splicing, Curr Opin Genet Dev 6, 215–20 (1996).
Colley KJ, Lee EU, Adler B, Browne JK, Paulson JC, Conversion of a Golgi apparatus sialyltransferase to a secretory protein by replacement of the NH2-terminal signal anchor with a signal peptide, J Biol Chem 264, 17619–22 (1989).
Paulson JC, Colley KJ, Glycosyltransferases. Structure, localization, and control of cell type-specific glycosylation, J Biol Chem 264, 17615–8 (1989).
Grabenhorst E, Conradt HS, The cytoplasmic, transmembrane, and stem regions of glycosyltransferases specify their in vivo functional sublocalization and stability in the Golgi, J Biol Chem 274, 36107–16 (1999).
Chen C, Colley KJ, Minimal structural and glycosylation requirements for ST6Gal I activity and trafficking, Glycobiology 10, 531–83 (2000).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Grahn, A., Larson, G. Identification of nine alternatively spliced α2,3-sialyltransferase, ST3Gal IV, transcripts and analysis of their expression by RT-PCR and laser-induced fluorescent capillary electrophoresis (LIF-CE) in twenty-one human tissues. Glycoconj J 18, 759–767 (2001). https://doi.org/10.1023/A:1021199300718
Issue Date:
DOI: https://doi.org/10.1023/A:1021199300718