Abstract
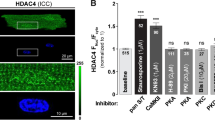
The myocyte enhancer factor-2 (MEF2) family of transcription factors regulates transcription of muscle-dependent genes in cardiac, skeletal and smooth muscle. They are activated by calcium/calmodulin (CaM)-dependent protein kinases I and IV and silenced by CaM KIIδC. MEF2 is held in an inactive form by the class II histone deacetylases (HDAC) until phosphorylated by either CaM kinase I or IV. Upon phosphorylation, HDAC is transported out of the nucleus via a 14-3-3 dependent mechanism freeing MEF2 to drive transcription. The 14-3-3 chaperone protein exists as a homodimer. In the region of homodimerization, there are two canonical CaM kinase II phosphorylation sites (ser60 and ser65). In vitro phosphorylation assay results indicate that 14-3-3β is indeed a substrate for CaM kinase II. We hypothesize that CaM kinase IIδC phosphorylation of 14-3-3β will disrupt homodimer formation resulting in the return of HDAC to the nucleus and their reassociation with MEF2. To test this, we mutated serines 60 and 65 of 14-3-3β to aspartates to mimic the phosphorylated state. In MEF2 enhancer-reporter assays in smooth muscle cells, expression of the 14-3-3β double mutant attenuated MEF2-enhancer activity driven by CaM kinase I or IV. The intracellular fate of HDAC4 was followed by transfection of smooth muscle cells with an HDAC4-Green Fluorescent Protein fusion hybrid. The 14-3-3β double mutant prevented HDAC4 cytoplasmic localization in the presence of active CaM kinase I or IV. These data suggest that the mechanism of CaM kinase IIδC silencing of MEF-2-dependent genes is by phosphorylation of 14-3-3β, which allows HDAC to return to the nucleus to reform a complex with MEF2, thereby silencing MADS box-dependent gene induction in smooth muscle.
Similar content being viewed by others
References
Katoh Y, Molkentin JD, Dave V, Olson EN, Periasamy M: MEF2B is a component of a smooth muscle-specific complex that binds an A/T-rich element important for smooth muscle myosin heavy chain gene expression. J Biol Chem 273: 1511-1518, 1998
Lin Q, Lu J, Yanagisawa H, Webb R, Lyons GE, Richardson JA, Olson EN: Requirement of the MADS-box transcription factor MEF2C for vascular development. Development 125: 4565-4574, 1998
Lu J, McKinsey TA, Nicol RL, Olson EN: Signal-dependent activation of the MEF2 transcription factor by dissociation from histone deacetylases. Proc Natl Acad Sci USA 97: 4070-4075, 2000
Lu J, McKinsey TA, Zhang CL, Olson EN: Regulation of skeletal myogenesis by association of the MEF2 transcription factor with class II histone deacetylases. Mol Cell 6: 233-244, 2000
Kao HY, Verdel A, Tsai CC, Simon C Juguilon H, Khochbin S: Mechanism for nucleocytoplasmic shuttling of histone deacetylase 7. J Biol Chem 276: 47496-47507, 2001
Passier R, Zeng H, Frey N, Naya FJ, Nicol RL, McKinsey TA, Overbeek P, Richardson JA, Grant SR, Olson EN: CaM kinase signaling induces cardiac hypertrophy and activates the MEF2 transcription factor in vivo. J Clin Invest 105: 1339-1342, 2000
McKinsey TA, Zhang CL, Olson EN: Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc Natl Acad Sci USA 97: 14400-14405, 2000
Wang AH, Kruhlak MJ, Wu J, Bertos NR, Vezmar M, Posner BI, Bazett-Jones DP, Yang XJ: Regulation of histone deacetylase 4 by binding of 14-3-3 proteins. Mol Cell Biol 20: 6904-6912, 2000
Vincenz C, Dixit VM: 14-3-3 proteins associate with A20 in an isoform-specific manner and function both as chaperone and adapter molecules. J Biol Chem 271: 20029-20034, 1996
Rittinger K, Budman J, Xu J, Volinia S, Cantley LC, Smerdon SJ, Gamblin SJ, Yaffe MB: Structural analysis of 14-3-3 phosphopeptide complexes identifies a dual role for the nuclear export signal of 14-3-3 in ligand binding. Mol Cell 4: 153-166, 1999
Grozinger CM, Schreiber SL: Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3-3-dependent cellular localization. Proc Natl Acad Sci USA 97: 7835-7840, 2000
Fu H, Subramanian RR, Masters SC: 14-3-3 proteins: Structure, function, and regulation. Annu Rev Pharmacol Toxicol 40: 617-647, 2000
Rothman A, Kulik TJ, Taubman MB, Berk BC, Smith CW, Nadal-Ginard B: Development and characterization of a cloned rat pulmonary arterial smooth muscle cell line that maintains differentiated properties through multiple subcultures. Circulation 86: 1977-1986, 1992
Haribabu B, Hook SS, Selbert MA, Goldstein EG, Tomhave ED, Edelman AM, Snyderman R, Means AR: Human calcium-calmodulin dependent protein kinase I: cDNA cloning, domain structure and activation by phosphorylation at threonine-177 by calcium-calmodulin dependent protein kinase I kinase. Embo J 14: 3679-3686, 1995
Cruzalegui FH, Means AR: Biochemical characterization of the multifunctional Ca2+/calmodulin-dependent protein kinase type IV expressed in insect cells. J Biol Chem 268: 26171-26178, 1993
Srinivasan M, Edman CF, Schulman H: Alternative splicing introduces a nuclear localization signal that targets multifunctional CaM kinase to the nucleus. J Cell Biol 126: 839-852, 1994
White SL, Low RB: Identification of promoter elements involved in cell-specific regulation of rat smooth muscle myosin heavy chain gene transcription. J Biol Chem 271: 15008-15017, 1996
Wu H, Rothermel B, Kanatous S, Rosenberg P, Naya FJ, Shelton JM, Hutcheson KA, DiMaio JM, Olson EN, Bassel-Duby R, Williams RS: Activation of MEF2 by muscle activity is mediated through a calcineurin-dependent pathway. Embo J 20: 6414-6423, 2001
Zeng H, Valencia T, Bai Y, Grant SR: Calcim/calmodulin-dependent enzymes regulate cardiac hypertrophy-sensitive gene expression. Miami Nature Biotechnol Short Reports 10: 1999
Valencia TG, Roberts LD, Zeng H, Grant SR: Tetracycline-inducible CaM kinase II silences hypertrophy-sensitive gene expression in rat neonate cardiomyocytes. Biochem Biophys Res Commun 274: 803-810, 2000
McDonough PM, Stella SL, Glembotski CC: Involvement of cytoplasmic calcium and protein kinases in the regulation of atrial natriuretic factor secretion by contraction rate and endothelin. J Biol Chem 269: 9466-9472, 1994
Ramirez MT, Zhao XL, Schulman H, Brown JH: The nuclear deltaB isoform of Ca2+/calmodulin-dependent protein kinase II regulates atrial natriuretic factor gene expression in ventricular myocytes. J Biol Chem 272: 31203-31208, 1997
Luo ZJ, Zhang XF, Rapp U, Avruch J: Identification of the 14.3.3 zeta domains important for self-association and Raf binding. J Biol Chem 270: 23681-23687, 1995
Chan TA, Hermeking H, Lengauer C, Kinzler KW, Vogelstein B: 14-3-3 Sigma is required to prevent mitotic catastrophe after DNA damage. Nature 401: 616-620, 1999
Liu D, Bienkowska J, Petosa C, Collier RJ, Fu H, Liddington R: Crystal structure of the zeta isoform of the 14-3-3 protein. Nature 376: 191-194, 1995
Obsil T, Ghirlando R, Klein DC, Ganguly S, Dyda F: Crystal structure of the 14-3-3zeta:serotonin N-acetyltransferase complex. A role for scaffolding in enzyme regulation. Cell 105: 257-267, 2001
Alessandrini A, Greulich H, Huang W, Erikson RL: Mek1 phosphorylation site mutants activate Raf-1 in NIH 3T3 cells. J Biol Chem 271: 31612-31618, 1996
Huang W, Kessler DS, Erikson RL: Biochemical and biological analysis of Mek1 phosphorylation site mutants. Mol Biol Cell 6: 237-245, 1995
Hurley JH, Dean AM, Sohl JL, Koshland DE Jr, Stroud RM: Regulation of an enzyme by phosphorylation at the active site. Science 249: 1012-1016, 1990
Mansour SJ, Candia JM, Matsuura JE, Manning MC, Ahn NG: Interdependent domains controlling the enzymatic activity of mitogen-activated protein kinase kinase 1. Biochemistry 35: 15529-15536, 1996
Orr JW, Newton AC: Requirement for negative charge on ‘activation loop’ of protein kinase C. J Biol Chem 269: 27715-27718, 1994
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ellis, J.J., Valencia, T.G., Zeng, H. et al. CaM kinase IIδC phosphorylation of 14-3-3β in vascular smooth muscle cells: Activation of class II HDAC repression. Mol Cell Biochem 242, 153–161 (2003). https://doi.org/10.1023/A:1021158216582
Issue Date:
DOI: https://doi.org/10.1023/A:1021158216582