Abstract
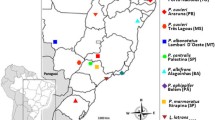
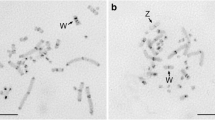
The PIM357 satellite DNA family is present in 26 Pimelia taxa (Tenebrionidae, Coleoptera) with endemic congeneric species from the Canary Islands showing higher interrepeat variability than continental ones. In this paper, we compare the repetitive DNA sequences of a Canarian species that has distinct subfamilies of repeat units, P. radula ascendens, with another without such subfamilies, P. sparsa sparsa. The chromosomal localization of the repeat units and the comparison of the variability of randomly cloned monomers to the one estimated by comparing repeat units from dimers and trimers suggest the absence of satellite subfamilies in P. sparsa sparsa. Hence, the repeat units of this species seem to be uniformly and randomly distributed throughout all chromosomes out of one chromosomal pair. On the contrary, P. radula ascendens shows four divergent subfamilies of repeat units supported by several diagnostic nucleotide substitutions. These subfamilies seem to form four distinct repeat units: monomer subfamily 1, monomer subfamily 4 and two higher-order units (dimer linking subfamily 1 and 4, and dimer linking subfamily 2 and 3). Moreover, monomers of subfamily 1 are present in three chromosomal pairs only. We discuss the effect of different potential factors acting in the concerted evolution and the genomic organization of stDNA sequences in these taxa.
Similar content being viewed by others
References
Biessmann H, Zurovcova M, Yao JG, Lozovskaya E, Walter MF (2000) A telomeric satellite in Drosophila virilis and its sibling species. Chromosoma 109: 372-380.
Bruvo B, Plohl M, Ugarkovic D (1995) Uniform distribution of satellite DNA variants on chromosomes of tenebrionid species Alphitobius diaperinus and Tenebrio molitor. Hereditas 123: 69-75.
Chaves R, Guedes-Pinto H, Heslop-Harrison J, Schwarzacher T (2000) The species and chromosomal distribution of the centromeric alpha-satellite I sequence from sheep in the tribe Caprini and other Bovidae. Cytogenet Cell Genet 91: 62-66.
Cox CB, Moore PD (1993) Biogeography: An Ecological and Evolutionary Approach. Oxford: Blackwell Science Ltd.
DeSalle R, Templeton AR (1988) Founder effects and the rate of mitochondrial DNA evolution in Hawaiian Drosophila. Evolution 42: 1076-1084.
Do GS, Seo BB, Yamamoto M, Suzuki G, Mukai Y (2001) Identification and chromosomal location of tandemly repeated DNA sequences in Allium cepa. Genes Genet Syst 76(1): 53-60.
Dover G (1986) Molecular drive in multigene families: how biological novelties arise, spread and are assimilated. Trends Genet 2: 159-165.
Elder F, Turner BJ (1995) Concerted evolution of repetitive DNA sequences in eukaryotes. Quart Rev Biol 70(3): 297-323.
Greig GM, Warburton PE, Willard HF (1993) Organization and evolution of an alpha satellite DNA subset shared by human chromosomes 13 and 21. J Mol Evol 37(5): 464-475.
Higgins DG, Thompson JD, Gibson TJ (1996) Using clustal for multiple sequence alignments. Meth Enzymol 266: 383-402.
Juan C, Pons J, Petitpierre E (1993) Localization of tandemly repeated DNA sequences in beetle chromosomes by fluorescent in situ hybridisation. Chromosome Res 1: 167-174.
Juan C, Oromí P, Hewitt GM (1995) Mitochondrial DNA phylogeny and sequential colonization of Canary Islands by darkling beetles of the genus Pimelia (Tenebrionidae). Proc R Soc B 261: 173-180.
Juan C, Ibrahim KM, Oromí P, Hewitt GM (1996) Mitochondrial DNA sequence variation and phylogeography of Pimelia darkling beetles on the island of Tenerife (Canary Islands). Heredity 77: 589-598.
Nietzel A, Rocchi M, Starke H et al. (2001) A new multicolor-FISH approach for the characterization of marker chromosomes: centromere-specific multicolor-FISH (cenM-FISH). Hum Genet 108(3): 199-204.
Plohl M, Borstnik B, Lucijanic-Justic V, Ugarkovic D (1992) Evidence for random distribution of sequence variants in Tenebrio molitor satellite DNA. Genet Res 60: 7-13.
Plohl M, Mestrovic N, Bruvo B, Ugarkovic D (1998) Similarity of structural features and evolution of satellite DNAs from Palorus subdepressus (Coleoptera) and related species. J Mol Evol 46: 234-239.
Pons J (1999) Evolucion del DNA Satelite en el Genero Pimelia. PhD Thesis. University of Balearic Islands, Palma de Mallorca (Spain).
Pons J, Petitpierre E, Juan C (2002) Evolutionary dynamics of satellite DNA family PIM357 in species of the genus Pimelia (Tenebrionidae, Coleoptera). Mol Biol Evol 19(8): 1329-1340.
Rozas J, Rozas (1997) DnaSP version 2.0 a novel software package for extensive molecular population genetic analysis. Comput Applic Biosci 13: 307-311.
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191: 528-535.
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75: 304-306.
Swofford DL (1999) PAUP*. Phylogenetics Analysis Using Parsimony (*and other methods). Version 4. Sunderland, MA: Sinauer Associates.
Trick M, Dover G (1984) Unexpectedly slow homogenisation within a repetitive DNA family shared between two species of tsetse fly. J Mol Evol 20: 322-329.
Willard HF, Wayne JS (1987) Hierarchical order in chromosome-specific human alpha satellite DNA. Trends Genet 3: 192-198.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Pons, J., Juan, C. & Petitpierre, E. Higher-order organization and compartmentalization of satellite DNA PIM357 in species of the coleopteran genus Pimelia . Chromosome Res 10, 597–606 (2002). https://doi.org/10.1023/A:1020918803675
Issue Date:
DOI: https://doi.org/10.1023/A:1020918803675