Abstract
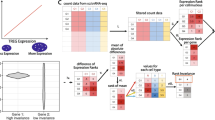
Microarray-based genomic techniques allow the simultaneous determination of relative levels of expression of a large number of genes. Studies of the transcriptome in complex neurobiological systems are uniquely demanding due to the heterogeneous nature of these cells. Most brain regions contain a large variety of cell populations that are closely intermingled. The expression of any specific gene may be restricted to a subpopulation of cells, and changes in gene expression may occur in only a small fraction of the cells expressing that transcript. Due to this dilution effect, many genes of interest are expected to have relatively low levels of expression in tissue homogenates. Furthermore, biologically significant differences in expression may result in only small fold-changes. Therefore genomic approaches using brain dissections must be optimized to identify potentially regulated transcripts and differential expression should be confirmed using quantitative assays. We evaluated the effects of increasing tissue complexity on detection of regulated transcripts in focused microarray studies using a mouse cell line, mouse hypothalamus and mouse cortex. Regulated transcripts were confirmed by quantitative real-time PCR. As tissue complexity increased, distinguishing significantly regulated genes from background variation became increasingly more difficult. However, we found that cDNA microarray studies using regional brain dissections and appropriate numbers of replicates could identify genes showing less than 2-fold regulation and that most regulated genes identified fell within this range.
Similar content being viewed by others
REFERENCES
Alizadeh, A. A., Eisen, M. B., Davis, R. E., Ma, C., Lossos, I. S., Rosenwald, A., Boldrick, J. C., Sabet, H., Tran, T., Yu, X., Powell, J. I., Yang, L., Marti, G. E., Moore, T., Hudson, J., Jr., Lu, L., Lewis, D. B., Tibshirani, R., Sherlock, G., Chan, W. C., Greiner, T. C., Weisenburger, D. D., Armitage, J. O., Warnke, R., and Staudt, L. M. 2000. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403:503-511.
Golub, T. R., Slonim, D. K., Tamayo, P., Huard, C., Gaasenbeek, M., Mesirov, J. P., Coller, H., Loh, M. L., Downing, J. R., Caligiuri, M. A., Bloomfield, C. D., and Lander, E. S. 1999. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286:531-537.
Pomeroy, S. L., Tamayo, P., Gaasenbeek, M., Sturla, L. M., Angelo, M., Mclaughlin, M. E., Kim, J. Y., Goumnerova, L. C., Black, P. M., Lau, C., Allen, J. C., Zagzag, D., Olson, J. M., Curran, T., Wetmore, C., Biegel, J. A., Poggio, T., Mukherjee, S., Rifkin, R., Califano, A., Stolovitzky, G., Louis, D. N., Mesirov, J. P., Lander, E. S., and Golub, T. R. 2002. Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature 415: 436-442.
Van T Veer, L. I., Dai, H., Van De Vijver, M. J., He, Y. D., Hart, A. A., Mao, M., Peterse, H. L., Van Der Kooy, K., Marton, M. J., Witteveen, A. T., Schreiber, G. J., Kerkhoven, R. M., Roberts, C., Linsley, P. S., Bernards, R., and Friend, S. H. 2002. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530-536.
Lashkari, D. A., Derisi, J. L., Mccusker, J. H., Namath, A. F., Gentile, C., Hwang, S. Y., Brown, P. O., and Davis, R. W. 1997. Yeast microarrays for genome wide parallel genetic and gene expression analysis. Proc. Natl. Acad. Sci. USA 94:13057-13062.
Iyer, V. R., Eisen, M. B., Ross, D. T., Schuler, G., Moore, T., Lee, J. C. F., Trent, J. M., Staudt, L. M., Hudson, J., Jr., Boguski, M. S., Lashkari, D., Shalon, D., Botstein, D., and Brown, P. O. 1999. The transcriptional program in the response of human fibroblasts to serum. Science 283:83-87.
Schena, M., Shalon, D., Davis, R. W., and Brown, P. O. 1995. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270:467-470.
Dudoit, S., Yang, Y. H., Callow, M. J., and Speed, T. P. 2000. Statistical methods for identifying differentially expressed genes in replicated cDNA microarray experiments. http://stat-www. berkeley.edu/users/terry/zarray/TechReport/578.pdf.
Kerr, M. K., Afshari, C. A., Bennett, L., Bushel, P., Martinez, J., Walker, N. J., and Churchill, G. A. 2001. Statistical analysis of a gene expression microarray experiment with replication. www.jax.org/research/churchill/research/expression/niebs.pdf.
Wurmbach, E., Yuen, T., Ebersole, B. J., and Sealfon, S. C. 2001. Gonadotropin-releasing hormone receptor-coupled gene network organization. J. Biol. Chem. 276:47195-47201.
Yuen, T., Zhang, W., Ebersole, B. J., and Sealfon, S. C. 2002. Monitoring G-protein-coupled receptor signaling with DNA microarrays and real-time polymerase chain reaction. Methods Enzymol. 345:556-569.
Yuen, T., Wurmbach, E., Pfeffer, R. L., Ebersole, B. J., and Sealfon, S. C. 2002. Accuracy and calibration of commercial oligonucleotide and custom cDNA microarrays. Nucleic Acids Res. 30:e48.
Alarid, E. T., Windle, J. J., Whyte, D. B., and Mellon, P. L. 1996. Immortalization of pituitary cells at discrete stages of development by directed oncogenesis in transgenic mice. Development 122:3319-3329.
Barnes, N. M. and Sharp, T. 1999. A review of central 5-HT receptors and their function. Neuropharmacology 38:1083-1152.
Lopez-Gimenez, J. F., Mengod, G., Palacios, J. M., and Vilaro, M. T. 1997. Selective visualization of rat brain 5-HT2A receptors by autoradiography with [3H]MDL 100,907. Naunyn Schmiedeberg's Arch. Pharmacol. 356:446-454.
Pazos, A., Cortes, R., and Palacios, J. M. 1985. Quantitative autoradiographic mapping of serotonin receptors in the rat brain. II. Serotonin-2 receptors. Brain Res. 346:231-249.
Jakab, R. L. and Goldman-Rakic, P. S. 1998. 5-Hydroxytryptamine2A serotonin receptors in the primate cerebral cortex: possible site of action of hallucinogenic and antipsychotic drugs in pyramidal cell apical dendrites. Proc. Natl. Acad. Sci. USA 95:735-740.
Ramakrishnan, R., Dorris, D., Lublinsky, A., Nguyen, A., Domanus, M., Prokhorova, A., Gieser, L., Touma, E., Lockner, R., Tata, M., Zhu, X., Patterson, M., Shippy, R., Sendera, T. J., and Mazumder, A. 2002. An assessment of Motorola CodeLink microarray performance for gene expression profiling applications. Nucleic Acids Res. 30:e30.
Trogan, E., Choudhury, R. P., Dansky, H. M., Rong, J. K., Breslow, J. L., and Fisher, E. A. 2002. Laser capture microdissection analysis of gene expression in macrophages from atherosclerotic lesions of apolipoprotein E-deficient mice. Proc. Natl. Acad. Sci. U S A 99:2234-2239.
Dent, G. W., O'dell, D. M., and Eberwine, J. H. 2001. Gene expression profiling in the amygdala: an approach to examine the molecular substrates of mammalian behavior. Physiol. Behav. 73:841-847.
Yuen, T., Wurmbach, E., Ebersole, B. J., Pfeffer, R. L., Ruf, F., and Sealfon, S. C. 2002. Coupling of GnRH concentration and the GnRHR-activated gene program. Mol. Endocrinol. 16:1145-1153.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wurmbach, E., González-Maeso, J., Yuen, T. et al. Validated Genomic Approach to Study Differentially Expressed Genes in Complex Tissues. Neurochem Res 27, 1027–1033 (2002). https://doi.org/10.1023/A:1020900720328
Issue Date:
DOI: https://doi.org/10.1023/A:1020900720328