Abstract
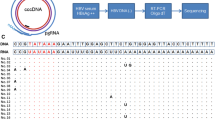
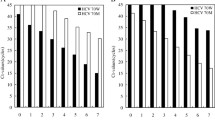
Reports suggest that response tointerferon-alpha therapy is influenced by both hepatitisC viral genotype and titer. Our aim was to determine ifdirect, automated, cycle sequencing of the PCR productfrom an HCV RNA detection assay could be used toreliably determine HCV genotype. In addition, theapproach was used to determine the HCV genotypedistribution in our patient population and to learn ifthere was a correlation between HCV genotype and RNAtiter that could be used to predict response totreatment. In all 143 consecutive patients were testedfor both HCV RNA titer and genotype. Automated, cycle sequencing of PCR product was highly effectiveand failed to yield a genotype in only 3 (2%) patients.The distribution of HCV genotypes was: 1a (40%), 1b(39%), 2a (2%), 2b (6%), 3a (4%). There were significant differences in the median HCV RNA titersbetween genotypes 1, 2, and 3. 6 High HCV RNA titers>4.4 × 106 copies/ml were only seenin genotype 1. However, the HCV RNA level should not beused as a surrogate marker of genotype because of a significantoverlap of titers within the genotypes.
Similar content being viewed by others
REFERENCES
Ogata N, Alter HJ, Miller RH, Purcell RH: Nucleotide sequence and mutation rate of the H strain of hepatitis C virus. Proc Natl Acad Sci USA 88:3391-3396, 1991
Schaluder GG, Dawson GJ, Simons JN, Pilot-Matias TJ, Gutierrez RA, Heynen CA, Knigge MF, Kurpiewski GS, Leary TP, Muerhoff AS, Desai SM, Mushahwar IK: Molecular and serologic analysis in the transmission of the GB hepatitis agents. J Med Virol 46:81-90, 1995
Chan SW, McOmish F, Holmes EC, Dow B, Peutherer JF, Follett E, Yap PL, Simmonds P: Analysis of a new hepatitis C virus type and its phylogenetic relationship to existing variants. J Gen Virol 73:1131-1141, 1992
Simmonds P, Holmes EC, Cha TA, Chan SW, McOmish F, Irvine B, Beall E, Yap PL, Kolberg J, Urdea MS: Classification of the hepatitis C virus into six major genotypes and a series of subtypes by phylogenetic analysis of the NS-5 region. J Gen Virol 74:2391-2399, 1993
Stuyver L, Wyseur A, van Arnhem W, Lunel F, Laurent-Puig P, Pawlotsky J-M, Kleter B, Bassit L, Nkengasong J, van Doorn L-J, Maertens G: Hepatitis C virus genotyping by means of 5′-UR/core line probe assays and molecular analysis of untypeable samples. Virus Res 38:137-157, 1995
Romeo R, Rumi M, Colombo M: Alpha interferon treatment of chronic hepatitis C. Biomed Pharmacother 49:111-115, 1995
Hino K, Sainokami S, Shimoda K, Iino S, Wang Y, Okamoto H, Miyakawa Y, Mayumi M: Genotypes and titers of hepatitis C virus for predicting response to interferon in patients with chronic hepatitis C. J Med Virol 42:299-305, 1994
Ansorge WB, Sproat J, Stegemann C, Schwager C, Zenke M: Automated DNA sequencing: Ultrasensitive detection of fluorescent bands during electrophoresis. Nucleic Acids Res 15:4593-4602, 1987
Mahaney K, Tedeschi V, Maertens G, Di Bisceglie AM, Vergalla J, Hoofnagle JH, Sallie R: Genotypic analysis of hepatitis C virus in American patients. Hepatology 20:1405-1411, 1994
Chemello L, Bonetti P, Canalletto L, Talato F, Conadon V, Casarin P, Belussi F, Frezza M, Noventa F, Pontisso P, Benvegnu L, Casarin C, Alberti A, TriVeneto Viral Hepatitis Group: Randomized trial comparing three different regimens of alpha-2a interferon in chronic hepatitis C. Hepatology 22:700-706, 1995
Davidson F, Simmonds P, Ferguson JC, Jarvis LM, Cow BC, Follett EA, Seed CR, Krusius T, Lin C, Medgyesi GA: Survey of major genotypes and subtypes of hepatitis C virus using RFLP of sequences amplified from the 5′ non-coding region. J Gen Virol 76:1197-1204, 1995
Stuyver L, Rossau R, Wyseur A, Duhamel M, Vanderborght B, Van Heuverswyn H, Maertens G: Typing of hepatitis C virus isolates and characterization of new subtypes using a line probe assay. J Gen Virol 74:1093-1102, 1993
Okamoto H, Tokita H, Sakamoto M, Horikita M, Kojima M, Iizuka H, and Mishiro S: Characterization of the genomic sequence of type V (or 3a) hepatitis C isolates and PCR primers for specific detection. J Gen Virol 74:2385-2390, 1993
Mita E, Hayashi N, Kanazawa Y, Hagiwara H, Ueda K, Kasahara A, Fusamoto H, Kamada T: Hepatitis C virus genotype and RNA titer in the progression of type C chronic liver disease. J Hepatol 21:468-473, 1994
Andonov A, Chaudhary RK: Subtyping of hepatitis C virus isolates by a line probe assay using hybridization. J Clin Microbiol 254-256, 1995
Lau JY, Mizokami M, Kolberg JA, Davis GL, Prescott LE, Ohno T, Perrillo RP, Lindsay KL, Gish RG, Qian KP, Kohara M, Simmonds P, Urdea MS: Application of six hepatitis C virus genotyping systems to sera from chronic hepatitis C patients in the United States. J Infect Dis 171:281-289, 1995
Bukh J, Purcell RH, Miller RH: Sequence analysis of the 5′ non-coding region of hepatitis C virus. Proc Natl Acad Sci USA 89:4942-4946, 1992
Sanger F, Nicklen S, Coulson AR: DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463-5467, 1977
Rights and permissions
About this article
Cite this article
O'Brien, C.B., Henzel, B.S., Wolfe, L. et al. cDNA Sequencing of the 5′ Noncoding Region (5′ NCR) to Determine Hepatitis C Genotypes in Patients with Chronic Hepatitis C. Dig Dis Sci 42, 1087–1093 (1997). https://doi.org/10.1023/A:1018813825486
Issue Date:
DOI: https://doi.org/10.1023/A:1018813825486