Abstract
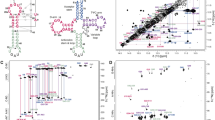
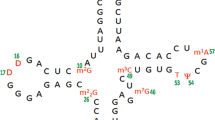
tRNA (m5U54)-methyltransferase (RUMT) catalyzes the S-adenosylmethionine-dependentmethylation of uridine-54 in the TΨC-loop of all transfer RNAs in E. coli to form the 54-ribosylthymine residue. However, in all tRNA structures, residue 54 is completely buried andthe question arises as to how RUMT gains access to the methylation site. A 17-mer RNAhairpin consisting of nucleotides 49–65 of the TΨ-loop is a substrate for RUMT.Homonuclear NMR methods in conjunction with restrained molecular dynamics (MD)methods were used to determine the solution structure of the 17-mer T-arm fragment. Theloop of the hairpin exhibits enhanced flexibility which renders the conventional NMR averagestructure less useful compared to the more commonly found situation where a molecule existsin predominantly one major conformation. However, when resorting to softer refinementmethods such as MD with time-averaged restraints, the conflicting restraints in the loop canbe satisfied much better. The dynamic structure of the T-arm is represented as an ensembleof 10 time-clusters. In all of these, U54 is completely exposed. The flexibility of the TΨ-loop in solution in conjunction with extensive binding studies of RUMT with the TΨC-loop and tRNA suggest that the specificity of the RUMT/tRNA recognition is associated withtRNA tertiary structure elements. For the methylation, RUMT would simply have to breakthe tertiary interactions between the D- and T-loops, leading to a melting of the T-armstructure and making U54 available for methylation.
Similar content being viewed by others
References
Amano, M. and Kawakami, M. (1992) Eur. J. Biochem., 210, 671–681.
Bax, A. and Davis, D.G. (1985) J. Magn. Reson., 65, 355–372.
Berendsen, H.J.C., Postma, J.P.M., van Gunsteren, W.F., Di Nola, A. and Haak, J.R. (1984) J. Chem. Phys., 81, 3684–3690.
Bonvin, A.M.J.J. and Brünger, A.T. (1995) J. Mol. Biol., 250, 80–93.
Bonvin, A.M.J.J. and Brünger, A.T. (1996) J. Biomol. NMR, 7, 72–76.
Fennen, J., Torda, A.E. and van Gunsteren, W.F. (1995) J. Biomol. NMR, 6, 163–170.
Gallo, K., Huang, C., Ferrin, T.F. and Langridge, R. (1985, 1989) Molecular Interactive Display and Simulation (MidasPlus), University of California, San Francisco, CA, U.S.A.
González, C., Stec, W., Reynolds, M. and James, T.L. (1995) Biochemistry, 34, 4969–4982.
Gu, X., Ivanetich, K.M. and Santi, D.V. (1996) Biochemistry, 35, 11652–11659.
Hall, K.B., Sampson, J.R., Uhlenbeck, O.C. and Redfield, A.G. (1989) Biochemistry, 28, 5791–5801.
Heerschap, A., Haasnoot, C.A.G. and Hilbers, C.W. (1983a) Nucleic Acids Res., 11, 4483–4499.
Heerschap, A., Haasnoot, C.A.G. and Hilbers, C.W. (1983b) Nucleic Acids Res., 11, 4501–4520.
Heus, H.A. and Pardi, A. (1991) J. Am. Chem. Soc., 113, 4360–4361.
Holbrook, S.R., Sussman, J.L., Warrant, R.W. and Kim, S.-H. (1978) J. Mol. Biol., 123, 631–660.
Hyde, E.I. and Reid, B.R. (1985) Biochemistry, 24, 4307–4314.
Jaeger, J.A. and Tinoco Jr., I. (1993) Biochemistry, 32, 12522–12530.
James, T.L. (1991) Curr. Opin. Struct. Biol., 1, 1042–1053.
Kealey, J.T., Gu, X. and Santi, D.V. (1994) Biochimie, 76, 1133–1142.
Keepers, J.W. and James, T.L. (1984) J. Magn. Reson., 57, 404–426.
Kemmink, J. and Scheek, R.M. (1995) J. Biomol. NMR, 6, 33–40.
Kneller, D.G., Sparky, NMR display and processing program, University of California, San Francisco, CA, U.S.A., © 1992.
Liu, H., Borgias, B., Kumar, A. and James, T.L. (1990, 1994) MARDIGRAS, University of California, San Francisco, CA, U.S.A.
Liu, H., Spielmann, H.P., Ulyanov, N.B., Wemmer, D.E. and James, T.L. (1995) J. Biomol. NMR, 6, 390–402.
Louise-May, S., Auffinger, P. and Westhof, E. (1996) In Biological Structure and Dynamics(Eds., Sarma, R.H. and Sarma, M.H.), Vol. 2, Adenine Press, Schenectady, NY, U.S.A., pp. 73–91.
Milligan, J.F., Groebe, D.R., Wilherell, G.W. and Uhlenbeck, O.C. (1987) Nucleic Acids Res., 15, 8783–8798.
Nanzer, A.P., van Gunsteren, W.F. and Torda, A.E. (1995) J. Biomol. NMR, 6, 313–320.
Pan, T., Long, D.M. and Uhlenbeck, O. (1993) In The RNA World (Eds., Gesteland, R.F. and Atkins, J.F.), Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, U.S.A., pp. 271–302.
Pearlman, D.A. and Kollman, P.A. (1991) J. Mol. Biol., 220, 429–457.
Pearlman, D.A. (1994) J. Biomol. NMR, 4, 1–16.
Pearlman, D.A., Case, D.A., Caldwell, J.C., Ross, W.S., Cheatham III, T.E., Ferguson, D.N., Seibel, G.L., Singh, U.C., Weiner, P.K. and Kollman, P.A. (1995) AMBER v. 4.1, University of San Francisco, San Francisco, CA, U.S.A.
Puglisi, J.D. and Tinoco, I.J. (1989) Methods Enzymol., 180, 304–325.
Quigley, G.J. and Rich, A. (1976) Science, 194, 796–806.
Romby, P., Acabon, P., Westhof, E., Ehresmann, C., Ebel, J.-P., Ehresmann, B. and Giege, R. (1987) J. Biomol. Struct. Dyn., 5, 669–687.
Roy, S. and Redfield, A.G. (1983) Biochemistry, 22, 1386–1390.
Ryckaert, J.P., Cicotti, G. and Berendsen, H.J.C. (1977) J. Comput. Phys., 23, 327–341.
Schmitz, U., Kumar, A. and James, T.L. (1992) J. Am. Chem. Soc., 114, 10564–10566.
Schmitz, U. and James, T.L. (1993) In Structural Biology: The State of the Art(Eds., Sarma, R.H. and Sarma, M.H.), Vol. 2, Adenine Press, Schenectady, NY, U.S.A., pp. 251–272.
Schmitz, U., Ulyanov, N.B., Kumar, A. and James, T.L. (1993) J. Mol. Biol., 234, 373–389.
Schmitz, U. and James, T.L. (1995) Methods Enzymol., 261, 1–43.
Schmitz, U., González, C., Liu, H., Ulyanov, N.B., Blocker, F. and James, T.L. (1996) In Biological Structure and Dynamics(Eds., Sarma, R.H. and Sarma, M.H.), Vol. 2, Adenine Press, Schenectady, NY, U.S.A., pp. 165–189.
Seibel, G.L., Singh, U.C. and Kollman, P.A. (1985) Proc. Natl. Acad. Sci. USA, 82, 6537–6540.
Shen, L.X., Cai, Z. and Tinoco, I. (1995) FASEB J., 9, 1023–1033.
Simmerling, C., Elber, R. and Zhang, J. (1995) In Modeling of Biomolecular Structures and Mechanisms(Ed., Pullman, A.), Kluwer Academic Publishers, Dordrecht, The Netherlands, pp. 241–265.
Smallcombe, S.H. (1993) J. Am. Chem. Soc., 115, 4776–4785.
States, D.J., Haberkorn, R.A. and Ruben, D.J. (1982) J. Magn. Reson., 48, 286–292.
Torda, A.E., Scheek, R.M. and van Gunsteren, W.F. (1990) J. Mol. Biol., 214, 223–235.
Torda, A.E., Brunne, R.M., Huber, T., Kessler, H. and van Gunsteren, W.F. (1993) J. Biomol. NMR, 3, 55–66.
Ulyanov, N.B., Schmitz, U., Kumar, A. and James, T.L. (1995) Biophys. J., 68, 13–24.
Varani, G. and Tinoco, I.J. (1991) Q. Rev. Biophys., 24, 479–532.
Wijmenga, S.S., Mooren, M.M.W. and Hilbers, C.W. (1993) In NMR of Macromolecules(Ed., Roberts, G.C.K.), Vol. 134, Oxford University Press, New York, NY, U.S.A., pp. 217–288.
Wüthrich, K. (1986) NMR of Proteins and Nucleic Acids, Wiley, New York, NY, U.S.A.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Yao, L.J., James, T.L., Kealey, J.T. et al. The dynamic NMR structure of the TΨC-loop: Implications for the specificity of tRNA methylation. J Biomol NMR 9, 229–244 (1997). https://doi.org/10.1023/A:1018618606857
Issue Date:
DOI: https://doi.org/10.1023/A:1018618606857