Abstract
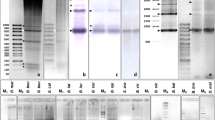
We have isolated and sequenced a member of tandem repetitive DNA containing BamHI site (BamHI family satellite DNA) from bluegill sunfish Lepomis macrochirus. PCR amplification with specific primers was performed to define the size of unit length repeat of the BamHI family satellite DNA, revealing that there were two distinct size of DNA fragments (0.9 kb and 1.3 kb) in the PCR products. The longer fragment (1.3 kb) consisted of internal sub-duplication of shorter fragment (0.9 kb). We have compared the size of PCR products among four fish populations, and found that both fragments co-existed in one population whereas the longer fragment was dominant in other three populations. The results may reflect ongoing homogenization of satellite DNA type over a short evolutionary time scale.
Similar content being viewed by others
References
Masumoto H, Masukata H, Muro Y, Nozaki N & Okazaki T (1989) A human centromere antigen (CENP-B) interact with a short specific sequence in alphoid DNA, a human centromeric alphoid. J. Cell Biol. 109: 1963–1973
Willard HF (1985) Chromosome-specific organization of human alpha satellite DNA. Amer. J. Human Genet. 37: 524–532
Willard HF (1990) Centromeres of mammalian chromosomes. Trends Genet. 6: 410–416
Zinkowski RP, Meyne J & Brinkley BR (1991) The centromere-kinetochore complex: a repeat subunit model. J. Cell Biol. 113: 1091–1110
Ikeno M, Masumoto H & Okazaki T (1994) Distribution of CENP-B boxes reflected in CREST centromere antigenic sites on long-range a-satellite DNA arrays of human chromosome 21. Human Mol. Genet. 3: 1245–1257
Laursen HB, Jørgensen AL, Jones C & Bak AL (1992) Higher rate of evolution of X chromosome α-repeat DNA in human than in the great apes. EMBO J. 11: 2367–2372
Franck JPC, Kornfield I & Wright JM (1994) The utility of SATA satellite DNA sequences for inferring phylogenetic relationships among the three major genera of Tilapiine Ciclid fishes. Mol. Phylogenet. Evol. 3: 10–16
Garrido-Ramos MA, de la Herran R, Jamilena M, Lozano R, Rejon CR & Rejon MR (1999) Evolution of centromeric satellite DNA and its use in phylogenetic studies of the Sparidae family (Pisces, Perciformes). Mol. Phylogenet. Evol. 12: 200–204
Kato M, Kato A & Shimizu N (1999) A method for evaluating phylogenetic relationship of α-satellite DNA suprachromosomal family by nucleotide frequency calculation. Mol. Phylogenet. Evol. 13: 329–335
Arnheim N (1983) Concerted evolution of multigene families. In Evolution of Genes and Proteins (Eds by Nei M & Koehn RK.) pp 38–61. Sinauer Associates, Inc., ISBN 0-87893-604-1
Elder JF Jr & Turner BJ (1994) Concerted evolution at the population level: pupfish HindIII satellite DNA sequences. Proc. Natl. Acad. Sci. USA 91: 994–998
Perbal B (1988) A Practical Guide to Molecular Cloning, John Wiley & Sons, New York
Kato M & Yoshida M (1995) Nucleotide sequence of a highly repetitive element isolated from Opsariichthys uncirostris (Osteichthyes). Mol. Biol. Rep. 21: 85–86.
Azuma M (1992) Ecological release in feeding behaviour: the case of bluegills in Japan. Hydrobiologia 243/244: 269–276
Kato M (1999) Structural bistability of repetitive DNA elements featuring CA/TG dinucleotide steps and mode of evolution of satellite DNA. Eur. J. Biochem. 265, 204–209
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Takahashi, T., Kawamura, Y., Sakata, N. et al. Nucleotide sequence of BamHI family satellite DNA and its unit length polymorphism in bluegill sunfish Lepomis macrochirus . Mol Biol Rep 28, 119–122 (2001). https://doi.org/10.1023/A:1017980227068
Issue Date:
DOI: https://doi.org/10.1023/A:1017980227068