Abstract
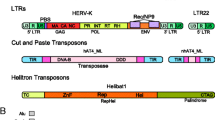
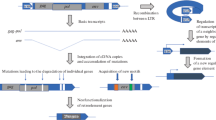
Phylogenetic analysis of transposable elements (TEs) allows us to define the relationships between the domains or gene(s) that compose them. Moreover, modules of a few amino-acids can be detected within gag, pol, envgenes or within the integrase domain of retrotransposons and transposase of DNA elements. The combination of these observations clearly shows that the evolutionary history of TEs is the outcome of the acquisition and loss of modules with differing origins and histories. This raises the question of the origin of TEs: are they derived from viruses? Do the basic building bricks come from the prokaryotes, and can they be assembled in the eukaryotes? Are the TEs found in prokaryotes the result of the disintegration of complex elements such as retroelements? Do they evolve from the simplest to the more complex, or are they opportunistic sequences evolving by acquiring and/or losing modules which may be either important or superfluous to their fitness (i.e., their ability to transpose). These are some of the questions that are addressed and discussed in the light of the comparative structures of TEs.
Similar content being viewed by others
REFERENCES
Capy, P. et al., Relationships between Transposable Elements Based upon the Integrase-Transposase Domains: Is There a Common Ancestor?, J. Mol. Evol., 1996, vol. 42, pp. 359-369.
Capy, P. et al., Does the Integrase of LTR-retrotransposons and Most of the Transposases of Class II Elements Share a Common Ancestor?, Genetica, 1997, vol. 100, pp. 63-72.
Lerat, E. and Capy, P., Retrotransposons and Retroviruses: Analysis of the Envelope Gene, Mol. Biol. Evol., 1999, vol. 16, pp. 1198-1207.
Lerat, E. et al., Is the Evolution of Transposable Elements Modular?, Genetica, 2000, vol. 107, pp. 15-25.
Malik, H.S. and Eickbush, T.H., Modular Evolution of the Integrase Domain in the Ty3/Gypsy Class of LTR Retrotransposons, Virol., 1999, vol. 73, no. 6, pp. 5186-5190.
McClure, M.A., Evolution of Retroposons by Acquisition or Deletion of Retrovirus-like Genes, Mol. Biol. Evol., 1991, vol. 8, pp. 835-856.
Xiong, Y. and Eickbush, T.H., Origin and Evolution of Retroelements Based upon Their Reverse Transcriptase Sequences, EMBO J., 1990, vol. 9, pp. 3353-3362.
Fayet, O. et al., Functional Similarities between Retroviruses and the IS3 Family of Bacterial Insertion Sequences?, Mol. Microbiol., 1990, vol. 4, pp. 1771-1777.
Doak, T.G. et al., A Proposed Superfamily of Transposase-related Genes: New Members in Transposon-like Elements of Cilliated Protozoa and a Common "D35E" Motif, Proc. Natl. Acad. Sci. USA, 1994, vol. 91, pp. 942-946.
Khan, E. et al., Retroviral Integrase Domains: DNA Binding and the Recognition of LTR Sequences, Nucleic Acids Res., 1991, vol. 19, pp. 851-860.
Lerat, E. et al., Is the Evolution of Transposable Element Modular?, Genetica, 1999, vol. 107, pp. 15-25.
Serre, M.C. et al., Mutagenesis of the IS1 Transposase: Importance of a His-Arg-Tyr Triad for Activity, J. Bacteriol., 1995, vol. 177, pp. 5070-5077.
Feschotte, C. and Wessler, S.R., Treasures in the Attic: Rolling Circle Transposons Discovered in Eukaryotic Genomes, Proc. Natl. Acad. Sci. USA, 2001, vol. 98, pp. 8923-8924.
Kapitonov, V.V. and Jurka, J., Rolling-circle Transposons in Eukaryotes, Proc. Natl. Acad. Sci. USA, 2001, vol. 98, pp. 8714-8719.
ICTV VIIth Report, Murphy, F.A., Ed., New York: Springer-Verlag, 1998.
Malik, H.S., Burke, W.D., and Eickbush, T.H., The Age and Evolution of Non-LTR Retrotransposable Elements, Mol. Biol. Evol., 1999, vol. 16, no. 6, pp. 793-805.
Malik, H.S. and Eickbush, T.H., Phylogenetic Analysis of Ribonuclease H Domains Suggests a Late, Chimeric Origin of LTR Retrotransposable Elements and Retroviruses, Genome Res., 2001, vol. 11, pp. 1187-1197.
Malik, H.S., Henikoff, S., and Eickbush, T.H., Poised for Contagion: Evolutionary Origins of the Infectious Abilities of Invertebrate Retroviruses, Genome Res., 2000, vol. 10, pp. 1307-1318.
Feschotte, C. and Mouches, C., Evidence That a Family of Miniature Inverted-repeat Transposable Elements (MITEs) from the Arabidopsis thaliana Genome Has Arisen from a pogo-like DNA Transposon, Mol. Biol. Evol., 2000, vol. 17, no. 5, pp. 730-737.
Feschotte, C. and Mouches, C., Recent Amplification of Miniature Inverted-repeat Transposable Elements in the Vector Mosquito Culex pipiens: Characterization of the Mimo Family, Gene, 2000, vol. 250, nos. 1–2, pp. 109-116.
Malik, H.S. and Eickbush, T.H., The RTE Class of Non-LTR Retrotransposons Is Widely Distributed in Animals and Is the Origin of Many SINEs, Mol. Biol. Evol., 1998, vol. 15, pp. 1123-1134.
Ohshima, K. et al., The 3′ End of tRNA-derived Short Interspersed Repetitive Elements Are Derived from the 3′ Ends of Long Interspersed Repetitive Elements, Mol. Cell. Biol., 1996, vol. 16, pp. 3756-3764.
Okada, N. et al., SINEs and LINEs Share Common 3' Sequences: a Review, Gene, 1997, vol. 205, pp. 229-243.
Okada, N. and Hamada, M., The 3' Ends of tRNA-derived SINEs Originated from the 3' Ends of LINEs: a New Example from the Bovine Genome, J. Mol. Evol., 1997, vol. 44, suppl. 1, pp. S52-S56.
Eickbush, T.H., Transposing without Ends: the Non-LTR Retrotransposable Elements, New Biol., 1992, vol. 4, pp. 430-440.
Luan, D.D. et al., Reverse Transcription of R2Bm RNA Is Primed by a Nick at the Chromosomal Target Site: a Mechanism for Non-LTR Retrotransposition, Cell, 1993, vol. 72, pp. 595-605.
Petrov, D.A., Lozovskaya, E.R., and Hartl, D.L., High Intrinsic Rate of DNA Loss in Drosophila, Nature, 1996, vol. 384, no. 6607, pp. 346-349.
Capy, P., Perspectives: Evolution. Is Bigger Better in Cricket? [comment], Science, 2000, vol. 287, no. 5455, pp. 985-986.
Petrov, D.A. et al., Evidence for DNA Loss as a Determinant of Genome Size, Science, 2000, vol. 287, pp. 1060-1062.
Champion, S. et al., Characterization of the Reverse Transcriptase of 1731, a Drosophila melanogaster Retrotransposon, Eur. J. Biochem., 1992, vol. 209, no. 2, pp. 523-531.
Shiba, T. and Saigo, K., Retrovirus-like Particles Containing RNA Homologous to the Transposable Element copia in Drosophila melanogaster, Nature, 1983, vol. 302, no. 5904, pp. 119-124.
McClure, M.A., Vasi, T.K., and Fitch, W.M., Comparative Analysis of Multiple Protein-sequence Alignment Methods, Mol. Biol. Evol., 1994, vol. 11, pp. 571-592.
Shigenobu, S. et al., Genome Sequence of the Endocellular Bacterial Symbiont of Aphids Buchnera sp. APS, Nature, 2000, vol. 407, no. 6800, pp. 81-86.
Kurland, C.G. and Andersson, S.G., Origin and Evolution of the Mitochondrial Proteome, Microbiol. Mol. Biol. Rev., 2000, vol. 64, no. 4, pp. 786-820.
Brennicke, A. et al., The Mitochondrial Genome on Its Way to the Nucleus: Different Stages of Gene Transfer in Higher Plants, FEBS Lett., 1993, vol. 325, no. 1–2, pp. 140-145.
Harington, A. and Thornley, A.L., Biochemical and Genetic Consequences of Gene Transfer from Endosymbiont to Host Genome, J. Mol. Evol., 1982, vol. 18, no. 5, pp. 287-292.
Jordan, I.K. and McDonald, J.F., Phylogenetic Perspective Reveals Abundant Ty1/Ty2 Hybrid Elements in the Saccharomyces cerevisiae Genome (Letter), Mol. Biol. Evol., 1999, vol. 16(3), pp. 419-422.
Arkhipova, I. and Meselson, M., Transposable Elements in Sexual and Ancient Asexual Taxa, Proc. Natl. Acad. Sci. USA, 2000, vol. 97, no. 26, pp. 14473-14477.
Wright, S. and Finnegan, D., Genome Evolution: Sex and the Transposable Element, Curr. Biol., 2001, vol. 11, no. 8, pp. R296-R299.
Zeyl, C., Bell, G., and Green, D.M., Sex and the Spread of Retrotransposon Ty3 in Experimental Populations of Saccharomyces cerevisiae, Genetics, 1996, vol. 143, no. 4, pp. 1567-1577.
Varmus, H. and Brown, P., Retroviruses, in Mobile DNA, Berg, D.E. and Howe, M.M., Eds., Washington, DC: American Society for Microbiology, 1989, pp. 53-108.
Brunet, F. et al., Do Deletions of the Mos1-like Elements Occur Randomly in the Drosophilidae Family?, J. Mol. Evol., 2002, vol. 54, pp. 227-234.
Rubin, E. and Levy, A.A., Abortive Gap Repair: Underlying Mechanism for Ds Element Formation, Mol. Cell. Biol., 1997, vol. 17, no. 11, pp. 6294-6302.
Mammano, F. et al., Role of the Major Homology Region of Human Immunodeficiency Virus Type 1 in Virion Morphogenesis, J. Virol., 1994, vol. 68, pp. 4927-4936.
Pardue, M.-L. et al., Evolutionary Links between Telomeres and Transposable Elements, Genetica, 1997, vol. 100, pp. 73-84.
Covey, S.N., Amino Acid Sequence Homology in gag Region of Reverse Transcribing Elements and the Coat Protein Gene of Cauliflower Mosaic Virus, Nucleic Acids Res., 1986, vol. 14, pp. 623-633.
Yang, J., Malik, H.S., and Eickbush, T.H., Identification of the Endonuclease Domain Encoded by R2 and Other Site-specific, Non-long Terminal Repeat Retrotransposable Elements, Proc. Natl. Acad. Sci. USA, 1999, vol. 96, no. 14, pp. 7847-7852.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Capy, P., Maisonhaute, C. Acquisition/Loss of Modules: the Construction Set of Transposable Elements. Russian Journal of Genetics 38, 594–601 (2002). https://doi.org/10.1023/A:1016027530962
Issue Date:
DOI: https://doi.org/10.1023/A:1016027530962