Abstract
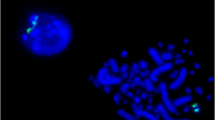
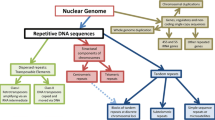
The majority of chicken repetitive sequence is nuclear-membrane-associated sequence (CNM), which resides in a large number of microchromosomes (chromosomes 11–39) and is absent from macrochromosomes 1–5, ZW, and some of the intermediate chromosomes 6–10. Two repetitive families, EcoRI/XhoI, are confined to the female-specific W chromosome. The core repeat units of the three families are 21 bp, containing (A)3–5 and (T)3–5 clusters separated by 5–7-bp sequences. In this article, we describe the isolation and initial characterization of a novel repeat family that is related to CNM/EcoRI/XhoI families. The novel family, designated as PIR, consists of multiple types of partially inverted repeat units of about 1.2, 1.4 and 1.6 kb. The PIR sequence is restricted to chicken chromosome 8, and accounts for about 3.8 mb, or 2500 copies of the 1.4-kb units, of the chicken genome. The evolution of PIR and related sequences is discussed.
Similar content being viewed by others
References
Auer H, Mayr B, Lambrou M, Schleger W (1987) An extended chicken karyotype, including the NOR chromosome. Cytogenet Cell Genet 45: 218–221.
Bedinger P, Munn M, Alberts BM (1989) Sequence-specific pausing during in vitro DNA replication on double DNA templates. JBiolChem 264: 16880–16886.
Boggs BA, Chinault AC (1994) Analysis of replication timing properties of human X-chromosomal loci by fluorescence in situ hybridization. Proc Natl Acad Sci USA 91: 6083–6087.
Epplen JT, Leipoldt M, Engel W, Schmidtke J (1978) DNA sequence organisation in avian genomes. Chromosoma 69: 307–321.
Hori T, Suzuki Y, Solovei I et al. (1996) Characterization of DNA sequences constituting the terminal heterochromatin of the chicken Z chromosome. Chromosome Res 4: 411–426.
John B, Mikilos GLG (1988) The Eukaryotic Genome in Development and Evolution. London: Allen and Unwin, pp 173–182.
Kalitsis P, Earle E, Vissel B, Shaffer LG, Choo KH (1993) A chromosome 13-specific human satellite I DNA subfamily with minor presence on chromosome 21: further studies on Robertsonian translocations. Genomics 16: 104–112.
Kodama H, Saitoh H, Tone M, Kuhara S, Sakaki Y, Mizuno S (1987) Nucleotide sequences and unusual electrophoretic behavior of the W chromosome-specific repeating DNA units of the domestic fowl, Gallusgallusdomesticus. Chromosoma 96: 18–25.
Ladjali-Mohammedi K, Bitgood JJ, Tixier-Boichard M, Ponce de Leon FA (1999) International System for Standardized Avian Karyotypes (ISSAK): standardized banded karyotypes of the domestic fowl (Gallus domesticus). Cytogenet Cell Genet 86: 271–276.
Lee C, Court DR, Cho C, Haslett JL, Lin CC (1997) Higher-order organization of subrepeats and the evolution of cervid satellite I DNA. J Mol Evol 44: 327–335.
Leung FC (1999) New applications of low-C0tDNA as a DNA fingerprint probe. Electrophoresis 20: 1762–1767.
Li Y, Lee C, Hsu TH, Li SY, Lin CC (2000) Direct visualization of the genomic distribution and organization of two cervid centromeric satellite DNA families. Cytogenet Cell Genet 89: 192–198.
Liao D, Pavelitz T, Kidd JR, Kidd KK, Weiner AM (1997) Concerted evolution of the tandemly repeated genes encoding human U2 snRNA (the RNU2 locus) involves rapid intrachromosomal homogenization and rare interchromosomal gene conversion. EMBO J 16: 588–598.
Lundqvist E, Johansson I, Ingelman-Sundberg M (1999) Genetic mechanisms for duplication and multiduplication of the human CYP2D6 gene and methods for detection of duplicated CYP2D6 genes. Gene 226: 327–338.
Matzke MA, Varga F, Berger H, et al (1990) A 41–42 bp tandemly repeated sequence isolated from nuclear envelopes of chicken erythrocytes is located predominantly on microchromosomes. Chromosoma 99: 131–137.
Matzke AJM, Varga F, Gruendler P et al. (1992) Characterization of a new repetitive sequences that is enriched on microchromosomes of turkey. Chromosoma 102: 9–14.
Parra I, Windle B (1993) High resolution visual mapping of stretched DNA by fluorescence hybridization. Nat Genet 5: 17–21.
Pinder DJ, Blake CE, Leach DR (1997) DIR: a novel DNA rearrangement associated with the inverted repeats. Nucleic Acids Res 25: 523–529.
Pluta AF, Mackay AM, Ainsztein AM, Goldberg IG, Earnshaw WC (1995) The centromere: hub of chromosomal activities. Science 270: 1591–1594.
Rogel-Gaillard C, Hayes H, Coullin P, Chardon P, Vaiman M (1997) Swine centromeric DNA repeats revealed by primed in situ (PRINS) labeling. Cytogenet Cell Genet 79: 79–84
Saitoh Y, Mizuno S (1992) Distribution of XhoI and EcoRI family repetitive DNA sequences into separate domains in the chicken W chromosome. Chromosoma 101: 474–477.
Saitoh H, Harata M, Mizuno S (1989) Presence of female-specific bent-repetitive DNA sequences in the genomes of turkey and pheasant and their interactions with W-protein of chicken. Chromosoma 98: 250–258.
Saitoh Y, Saitoh H, Ohtomo K, Mizuno S (1991) Occupancy of the majority of DNA in the chicken W chromosome by bent-repetitive sequences. Chromosoma 101: 32–40.
Senger G, Jones TA, Fidlerova H et al. (1994) Released chromatin: linearized DNA for high resolution fluorescence in situ hybridization. Hum Mol Genet 3: 1275–1280.
Solovei I, Gaginskaya E, Hutchison N, Macgregor H (1993) Avian sex chromosomes in the lampbrush form: the ZW lampbrush bivalents from six species of bird. Chromosome Res 1: 153–166.
Solovei I, Ogawa A, Naito M, Mizuno S, Macgregor H (1998) Specific chromomeres on the chicken W lampbrush chromosome contain specific repetitive DNA sequence families. Chromosome Res 6: 323–327
Sundquist WI, Klug A (1989) Telomeric DNA dimerizes by formation of guanine tetrads between hairpin loops. Nature 342: 825–829.
Tatusova TA, Madden TL (1999) BLAST 2 Sequences, a new tool for comparing protein and nucleotide sequences. FEMS Microbiol Lett 177: 187–188.
Tone M, Nakano N, Takao E, Narisawa S, Mizuno S (1982) Demonstration of W chromosome-specific repetitive DNA sequences in the domestic fowl, Gallus g. domesticus. Chromosoma 86: 551–569.
Valero MC, de Luis O, Cruces J, Perez Jurado LA (2000) Fine-scale comparative mapping of the human 7q11.23 region and the orthologous region on mouse chromosome 5G: the low-copy repeats that flank the Williams–Beuren syndrome deletion arose at breakpoint sites of an evolutionary inversion(s). Genomics 69: 1–13.
van de Rijke FM, Florijn RJ, Tanke HJ, Raap AK (2000) DNA fiber–FISH staining mechanism. J Histochem Cytochem 48: 743–745.
Viguera E, Canceill D, Ehrlich SD (2001) Replication slippage involves DNA polymerase pausing and dissociation. EMBO J 20: 2587–2595.
Weitzmann MN, Woodford KJ, Usdin K (1997) DNA secondary structures and the evolution of hypervariable tandem arrays. J Biol Chem 272: 9517–9523.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wang, X., Li, J. & Leung, F.C. Partially Inverted Tandem Repeat Isolated from Pericentric Region of Chicken Chromosome 8. Chromosome Res 10, 73–82 (2002). https://doi.org/10.1023/A:1014226412339
Issue Date:
DOI: https://doi.org/10.1023/A:1014226412339