Abstract
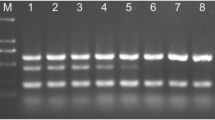
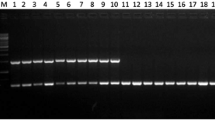
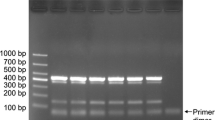
A multiplex Polymerase Chain Reaction (PCR) assay was developed to detect and quantify four fungal foliar pathogens in wheat. For Septoria tritici (leaf blotch) and Stagonospora nodorum (leaf and glume blotch), the β-tubulin gene was used as the target region. Diagnostic targets for Puccinia striiformis (stripe or yellow rust) and P. recondita (brown rust) were obtained from PCR products amplified with β-tubulin primer sequences. Final primer sets were designed and selected after being tested against several fungi, and against DNA of infected and healthy wheat leaves. For detection of the four pathogens, PCR products of different sizes were amplified simultaneously, whereas no products were generated from wheat DNA or other non-target fungi tested. The presence of each of the diseases was wheat tissue- and cultivar specific. Using real-time PCR measurements with the fluorescent dye SYBR Green I, PCR-amplified products could be quantified individually, by reference to a standard curve generated by adding known amounts of target DNA. Infection levels for each of the diseases were measured in the flag leaf of 19 cultivars at Growth Stage (GS) 60–64 in both 1998 and 1999. The infection levels for the cultivars were ranked, and showed, with a few exceptions, a good correlation with the NIAB Recommended List for winter wheat, which is based on visual assessment of symptoms. With PCR, the presence of the different pathogens was accurately diagnosed and quantification of pre-symptomatic infection levels was possible. Although sampling and DNA detection methods need further optimisation, the results show that multiplex PCR and quantitative real-time PCR assays can be used in resistance screening to measure the interaction between different pathogens and their hosts at different growth stages, and in specific tissues. This should enable an earlier identification of specific resistance mechanisms in both early-stage breeding material and field trials.
Similar content being viewed by others
References
Ahn SJ, Costa J and Emanuel JR (1996) PicoGreen quantification of DNA: effective evaluation of pre-or post-PCR. Nucleic Acids Research 24: 2623–2625
Anonymous (1997—2000) Cereal variety handbooks 1997—2000. NIAB recommended lists for winter wheat. National Institute of Agricultural Botany, Cambridge, UK.
Bayles RA (1997) Disease development — the interaction of variety resistance and pathogen. Aspects of Applied Biology 50: 249–254
Bayles RA and Stigwood PL (1999) Yellow rust of wheat. In: Clarkson JDS (ed.) UK cereal pathogen virulence survey 1999 Annual Report (pp 29–35) The United Kingdom Cereal Pathogen Virulence Survey Committee, Cambridge, UK.
Beck JJ and Ligon JM (1995) Polymerase chain reaction assays for the detection of Stagonospora nodorum and Septoria tritici in wheat. Phytopathology 85: 319–324
Bubert A, Hein I, Rauch M, Lehner A, Yoon B, Goebel W and Wagner M(1999) Detection and differentiation of Listeria spp. by a single reaction based on multiplex PCR. Applied and Environmental Microbiology 65: 4688–4692
Cook RJ, Hims MJ and Vaughan TB (1999) Effects of fungicide spray timing on winter wheat disease control. Plant Pathology 48: 33–50
Cook RJ, Polley RW and Thomas MR (1991) Disease-induced losses in winter wheat in England andWales 1985—1989. Crop Protection 10: 504–508
Doohan FM, Parry DW and Nicholson P (1999) Fusarium ear blight of wheat: the use of quantitative PCR and visual disease assessment in studies of disease control. Plant Pathology 48: 209–217
Eyal Z (1999) The septoria tritici and stagonospora nodorum blotch diseases of wheat. European Journal of Plant Pathology 105: 629–641
Foster SJ, Singh G, Fitt BDL and Ashby AM (1999) Development of PCR based diagnostic techniques for the two mating types of Pyrenopezia brassicae (light leaf spot) on winter oilseed rape (Brassica napus spp. oleifera) Physiological and Molecular Plant Pathology 55: 111–119
Fraaije BA, Lovell DJ, Rohel EAand Hollomon DW(1999) Rapid detection and diagnosis of Septoria tritici epidemics in wheat using a polymerase chain reaction/PicoGreen assay. Journal of Applied Microbiology 86: 701–708
Higuchi R, Fockler C, Dollinger G and Watson R (1993) Kinetic PCRanalysis: real-time monitoring ofDNAamplification reactions. Biotechnology 11: 1026–1030
Johnson R (1992) Reflections of a plant pathologist on breeding for disease resistance, with emphasis on yellowrust and eyespot of wheat. Plant Pathology 41: 239–254
Kema GHJ and Silfhout CH (1997) Genetic variation for virulence and resistance in the wheat-Mycosphaerella graminicola pathosystem III. Comparative seedling and adult plant experiment. Phytopathology 87: 266–272
Kendall SJ, Hollomon DW and Selley A(1998) Immunodiagnosis as an aid to the timing of fungicide sprays for the control of Mycosphaerella graminicola on winter wheat in the UK. In: Proceedings Brighton Crop Protection Conference — Pests and Diseases 1998, Vol 2. (pp 701–706) British Crop Protection Council, Farnham, UK
Koenraadt H and Jones AL (1992) The use of allele-specific oligonucleotide probes to characterize resistance to benomyl in field strains of Venturia inaequalis. Phytopathology 82: 1354–1358
Livak KJ, Flood SJA, Marmero J, Giusti W and Deetz K (1995) Oligonucleotides with fluorescent dyes at opposite ends provide a quenched probe system useful for detecting PCR products and nucleic acid hybridization. PCR Methods and Applications 4: 357–362
Lovell DJ, Parker SR, Hunter T, Royle DJ and Coker RR (1997) Influence of crop growth and structure on the risk of epidemics by Mycosphaerella graminicola (Septoria tritici) in winter wheat. Plant Pathology 46: 126–138
McIntosh RA, Wellings CR and Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO, Australia, and Kluwer Academic Publishers, Dordrecht, The Netherlands
Messmer MM, Seyfarth R, Keller M, Schachermayr G, Winzeler M, Zanetti S, Feuillet C and Keller B (2000) Genetic analysis of durable leaf rust resistance in winter wheat. Theoretical and Applied Genetics 100: 419–431
Mitchell SE, Kresovich SK, Jester CA, Hernandez CJ and Szewc-McFadden AK (1997) Application of multiplex PCR and fluorescence-based, semi-automated allele sizing technology for genotyping plant genetic resources. Crop Science 37: 617–624
Nicholson P, Rezanoor HN, Simpson DR and Joyce D (1997) Differentiation and quantification of the cereal eyespot fungi Tapesia yallundae and Tapesia acuformis using a PCR assay. Plant Pathology 46: 842–856
Park RF, Jahoor A and Felsenstein FG (2000) Population structure of Puccinia recondita in Western Europe during 1995, as assessed by variability in pathogenicity and molecular markers. Journal of Phytopathology 148: 169–179
Parker SR, Perry J and Royle DJ (1995a) Reliable measurement of disease severity. Aspects of Applied Biology 43: 205–214
Parker SR, Shaw MW and Royle DJ (1995b) The reliability of visual estimates of disease Severity on cereal leaves. Plant Pathology 44: 856–864
Parker SR, Shaw MW and Royle DJ (1997) Measurements of spatial patterns of disease in winter wheat crops and the implications for sampling. Plant Pathology 46: 470–480
Peters BA, Loughman R and Di Prinzio P (1996) Leaf development in relation to infection by Stagonosporum nodorum and Septoria tritici in wheat. Australian Journal of Agricultural Research 47: 1169–1179
Polz MF and Cavanaugh CM (1998) Bias in template-to-product ratios in multitemplate PCR. Applied and Environmental Microbiology 64: 3724–3730
Sambrook J, Fritsch EF and Maniatis T (1989) Molecular cloning — a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Plainview, NY, USA
Sanger F, Nicklen S and Coulson AR (1977) DNA sequencing with chain termination inhibitors. Proceedings National Academy of Sciences USA 74: 5463–5467
Schneeberger C, Speiser P, Kury F and Zeillinger R (1995) Quantitative detection of reverse transcriptase-PCR products by means of a novel and sensitive DNA stain. PCR Methods and Applications 4: 234–238
Shin JH, Nolte FS, Holloway BP and Morrison CJ (1999) Rapid identification of up to three Candida species in a single reaction tube by 5′exonuclease assay using fluorescent DNA probes. Journal of Clinical Microbiology 37: 165–170
Thelwell N, Millington S, Solinas A, Booth J and Brown T (2000). Mode of action and application of Scorpion primers to mutation detection. Nucleic Acids Research 28: 3752–3761
Tottman DR (1987) The decimal code for the growth stages of cereals, with illustrations. Annals of Applied Biology 110: 441–454
Tyagi S and Kramer FR (1996) Molecular Beacons: probes that fluoresce upon hybridization. Nature Biotechnology 14: 303–308
Wainshilbaum SJ and Lipps PE (1991) Effect of temperature and growth stage of wheat on development of leaf and glume blotch caused by Septoria tritici and S. nodorum. Plant Disease 75: 993–998
Weller SA, Elphinstone JG, Smith NC, Boonham N and Stead DE (2000) Detection of Ralstonia solanacearum strains with a quantitative, multiplex, real-time, fluorogenic PCR (TaqMan) assay. Applied and Environmental Microbiology 66: 2853–2858
Wiese MV (1987) Compendium of wheat diseases, 2nd edn. APS press, St. Paul, USA
Willits DA and Sherwood JE (1999) Polymerase Chain Reaction detection of Ustilago hordei in leaves of susceptible and resistant barley varieties. Phytopathology 89: 212–217
Wittwer CT, Herrmann MG, Moss AA and Rasmussen RP (1997) Continuous fluorescence monitoring of rapid cycleDNAampli-fication. Biotechniques 22: 130–138
Zhangh AW, Hartman GL, Curio-Penny B, Pedersen WL and Becker KB (1999) Molecular detection of Diaporthe phaseolorum and Phomopsis longicolla from soybean seeds Phytopathology 89: 796–804.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fraaije, B., Lovell, D., Coelho, J. et al. PCR-based Assays to Assess Wheat Varietal Resistance to Blotch (Septoria Tritici and Stagonospora Nodorum) and Rust (Puccinia Striiformis and Puccinia Recondita) Diseases. European Journal of Plant Pathology 107, 905–917 (2001). https://doi.org/10.1023/A:1013119206261
Issue Date:
DOI: https://doi.org/10.1023/A:1013119206261