Abstract
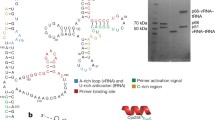
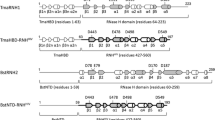
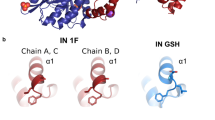
The natural form of the human immunodeficiency virus type one reverse transcriptase (HIV‐1 RT) found in virion particles is a heterodimer composed of the p66 and p51 subunits. The catalytic activity resides in the larger subunit in the heterodimeric (p66/p51) enzyme while in the monomeric form it is inactive. In contrast, Murine leukemia virus RT (MuLV RT) is functionally active in the monomeric form. In the primary amino acid sequence alignment of MuLV RT and HIV‐1 RT, we have identified three specific regions in MuLV RT, that were missing in HIV‐1 RT. In a separate study, we have shown that a chimeric RT construct comprising of the polymerase domain of HIV‐1 RT and RNase-H domain of MuLV RT is functionally active as monomer [20]. In this communication, we demonstrate that insertion of a peptide (corresponding to amino acid residues 480–506) from the connection subdomain of MuLV RT into the connection subdomain of HIV‐1 RT (between residues 429 and 430) results in a functionally active monomeric chimeric RT. Furthermore, this chimeric enzyme does not dimerize with exogenously added p51 subunit of HIV‐1 RT. Functional analysis of the chimeric RT revealed template specific variations in its catalytic activity. The chimeric enzyme catalyzes DNA synthesis on both heteropolymeric DNA and homopolymeric RNA (poly rA) template but curiously lacks reverse transcriptase ability on heteropolymeric RNA template. Similar to MuLV RT, the polymerase activity of the chimeric enzyme is not affected by acetonitrile, a reagent which dissociates dimeric HIV‐1 RT into inactive monomers. These results together with a proposed 3‐D molecular model of the chimeric enzyme suggests that the insertion of the missing region may induce a change in the spatial position of RNase H domain such that it is functionally active in monomeric conformation.
Similar content being viewed by others
References
Gilboa E, Mitra SW, Goff S, Baltimore DA: Detailed model of reverse transcription and tests of crucial aspects. Cell 18: 93–100, 1979
Levine JG, Hatfield D, Oroszlan S, Rein A: In: A.M. Skalka, S.P. Goff (eds). Reverse Transcriptase. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1993
Molling K, Bolognesi DP, Bauer H, Busen W, Plassmann HW, Hausen P: Association of viral reverse transcriptase with an enzyme degrading the RNA moiety of RNA-DNA hybrids. Nature - New Biol 234: 240–243, 1971
Varmus H, Swanstrom R: In: R. Weiss, N. Teich, H. Varmus, J. Coffin (eds). RNA Tumor Virus. Cold Spring harbor Laboratory Press, Cold Spring Harbor, NY, 1985, pp 75–134
Keller W, Crouch R: Degradation of DNA RNA hybrids by ribonuclease H and DNA polymerases of cellular and viral origin. Proc Natl Acad Sci USA 69: 3360–3364, 1972
Gerard GF, Grandgenett DP: Purification and characterization of the DNA polymerase and RNase-H activities in Moloney murine sarcomaleukemia virus. J Virol 15: 785–797, 1975
diMarzo-Veronese F, Copeland TD, DeVico AL, Rahman R, Oroszlan S, Gallo RC, Sarngadharan MG: Characterization of highly immunogenic p66/p51 as the reverse transcriptase of HTLV-III/LAV. Science 231: 1289–1291, 1986
Davies JF, Hostomska Z, Hostomsky Z, Jordan SR, Matthews DA: Crystal structure of the ribonuclease H domain of HIV-1 reverse transcriptase. Science 252: 88–95, 1991
North TW, Cronn RC, Remington KM, Tandberg RT, Judd RC: Characterization of reverse transcriptase from feline immunodeficiency Virus. J Biol Chem 265: 5121–5128, 1990
Le Grice SFJ, Ette R, Mills J, Mous J: Comparison of the human immunodeficiency virus type 1 and 2 proteases by hybrid gene construction and trans-complementation. J Biol Chem 264: 14902–14908, 1989
Mous J, Heimer EP, Le Grice SFJ: Processing protease and reverse transcriptase from human immunodeficiency virus type I polyprotein in Escherichia coli. J Virol 62: 1433–1436, 1988
Varmus H, Swantrom R: In: R. Weiss, N. Teich, H. Varmus, J. Coffin (eds). RNA Tumor Virus. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1984, pp 369–512
Wondrak EM, Lower J, Kurth R: Functional purification and enzymatic characterization of the RNA-dependent DNA polymerase of the human immunodeficiency virus. J Gen Virol 67: 2791–2797, 1986
Lightfoote M, Coligan JE, Folks TM, Fauci AS, Martin M A, Venkatesan S: Structural characterization of reverse transcriptase and endonuclease polypeptides of the acquired immunodeficiency syndrome retrovirus. J Virol 60: 771–775, 1986
Hostomsky Z, Hostomska Z, Fu TB, Taylor J: Reverse transcriptase of human immunodeficiency virus type 1: Functionality of subunits of the heterodimer in DNA synthesis. J Virol 66: 3179–3182, 1992
LeGrice SFJ, Naas T, Wohlgensinger B, Schatz O: Subunit-selective mutagenesis indicates minimal polymerase activity in heterodimer-associated p51 HIV-1 reverse transcriptase. EMBO J 10: 3905–3911, 1991
Harris D, Lee R, Misra HS, Pandey PK, Pandey VN: The p51 subunit of human immunodeficiency virus type 1 reverse transcriptase is essential in loading the p66 subunit on the template primer. Biochemist y 37: 5903–5908, 1998
Kohlstaedt LA, Wang J, Friedman JM, Rice PA, Steitz TA: Crystal structural at 3.5A resolution of HIV-1 RT complexed with an inhibitor. Science 256: 1783–1790, 1992
Wang J, Smerdon SJ, Jager J, Kohlstaedt LA, Rice PA, Friedman JM, Steitz TA: Structural basis of asymmetry in the human immunodeficiency virus type 1 reverse transcriptase heterodimer. Proc Natl Acad Sci USA 91: 7242–7246, 1994
Misra HS, Pandey PK, Pandey VN: An enzymatically active chimeric HIV-1 reverse transcriptase (RT) with the RNase-H domain of murine leukemia virus RT exists as a monomer. J Biol Chem 273: 9785–9789, 1998
Maxam A, Gilbert W: Sequencing end-labeled DNA with base-specific chemical cleavages. Meth Enzymol 65: 499–560, 1980
Arts EJ, Li X, Gu Z, Kleiman L, Parniak M, Wainberg MA: Comparison of deoxyoligonucleotide and t-RNAlys-3 as primers in an endogenous human immunodeficiency virus-1 in vitro reverse transcription/template-switching reaction. Biol Chem 269: 14672–14680, 1994
Pandey VN, Kaushik N, Rege N, Sarafianos SG, Yadav PNS, Modak MJ: Role of methionine 184 of human immunodeficiency virus typereverse transcriptase in the polymerase function and fidelity of DNA synthesis. Biochemistry 35: 2168–2179, 1996
Kaushik N, Harris D, Rege N, Modak MJ, Yadav PNS, Pandey VN: Role of glutamine-151 of human immunodeficiency virus type-1 reverse transcriptase in RNA-directed DNA synthesis. Biochemistry 36: 14430–14438, 1997
Chowdhury K, Kaushik N, Pandey VN, Modak MJ: Elucidation of the role of arg 110 of murine leukemia virus reverse transcriptase in the catalytic mechanism: Biochemical characterization of mutant enzymes. Biochemistry 35: 16610–16620, 1996
Saiki RK, Scharf S, Faloona F, Mullis KB, Horn GT, Erlich HA, Arnheim N: Enzymatic amplification of b-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230: 1350–1354, 1985
Roth M, Tanese N, Goff S P: Purification and characterization of murine retroviral reverse transcriptase expressed in Escherichia coli. J Biol Chem 260: 9326–9335, 1985
Lee R, Kaushik N, Modak MJ, Vinayak R, Pandey VN: Polyamide nucleic acid targeted to the primer binding site of the HIV-1 RNA genome blocks in vitro HIV-1 reverse transcription. Biochemistry 37: 900–910, 1998
Hsieh JC, Zinnen S, Modrich P: Kinetic mechanism of the DNA-dependent DNA polymerase activity of human immunodeficiency virus-1 reverse transcriptase. J Biol Chem 268: 24607–24613, 1993
Kaushik N, Rege N, Sarafianos SG, Yadav PNS, Modak MJ, Pandey VN: Biochemical analysis of catalytically crucial aspartate mutants of human immunodeficiency virus type 1 reverse transcriptase. Biochemistry 35: 11536–11546, 1996
Modak MJ, Marcus SL: Purification and properties of Rauscher leukemia virus DNA polymerase and selective inhibition of mammalian viral reverse transcriptase by inorganic phosphate. J Biol Chem 252: 11–19, 1977
Telesnitsky A, Goff SP: RNase H domain mutations affect the interaction between Moloney murine leukemia virus reverse transcriptase and its primer-template. Proc Natl Acad Sci USA 90: 1276–1280, 1993
Harris D, Kaushik N, Pandey PK, Yadav PNS, Pandey VN: Functional analysis of amino acid residues constituting the dNTP binding pocket of HIV-1 reverse transcriptase. J Biol Chem 273: 33624–33634, 1998
Divita G, Restle T, Goody RS: Characterization of the dimerization process of HIV-1 reverse transcriptase heterodimer using intrinsic protein fluorescence. FEBS Lett 324: 153–158, 1993
Jonckheere H, Taymans JM, Balzarini J, Velazquez S, Camarasa MJ, Desmyter J, De Clercq E, Anne J: Resistance of HIV-1 reverse transcriptase against [2', 5'-bis-O-(tert-butyldimethylsilyl)-3'spiro-5"-(4"-amino-1"-oxathiole-2",2"-dioxide)] (TSAO) derivatives is determined by the mutation Glu 138 → Lys on the p51 subunit. J Biol Chem 269: 25255–25258, 1994
Sharma SK, Basu A, Fan N, Evans DB: Engineering of the humanimmunodeficiency-virus-type-1 (HIV-1) reverse transcriptase gene to prevent dimerization of the expressed chimeric protein: purification and characterization of a monomeric HIV-1 reverse transcriptase. Biotech Appl Biochem 19: 155–167, 1994
Huang H, Chopra R, Verdine GL, Harrison S: Structure of a covalently trapped catalytic complex of HIV-1RT: Implication for drug resistance. Science 282: 1669–1675, 1998
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Pandey, P.K., Kaushik, N., Talele, T.T. et al. Insertion of a peptide from MuLV RT into the connection subdomain of HIV-1 RT results in a functionally active chimeric enzyme in monomeric conformation. Mol Cell Biochem 225, 135–144 (2001). https://doi.org/10.1023/A:1012278308154
Issue Date:
DOI: https://doi.org/10.1023/A:1012278308154