Abstract
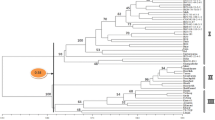
Forty-four accessions of cultivated peanut (Arachis hypogaea L.) representing sixbotanical varieties of two subspecies along with three accessions ofthe wild relative A. monticola Krapov et Rigoni were evaluated for their genetic relationships using theAFLP marker technology. Fifteen AFLP primer pairs (EcoRI/MseI) generated 28distinct polymorphic markers that were employed to develop uniqueprofiles of all accessions and to construct a phenogram. The resultsshowed that the botanical varieties aequatoriana and peruviana werecloser to subspecies hypogaea than subspeciesfastigiata Waldr. to which they belong, and the wildA. monticola was notdistinct from the cultivated A.hypogaea. Although the extent of geneticdiversity in peanut is low compared to many other crops, our studiesshow that by employing the AFLP approach, sufficient DNA variationcan be detected in the cultivated peanut germplasm to conductevolutionary studies.
Similar content being viewed by others
References
Ayala F.J. 1982. Population and Evolutionary Genetics. Benjamin/ Cummings Publishing Company, Inc, Menlo Park, CA.
Bassam B.J., Gaetano-Anolles G. and Gresshoff P.M. 1991. Fast and sensitive silver staining of DNA in polyacrylamide gels. Analytical Biochemistry 196: 80–83.
Chen J. and Dellaporta S. 1994. Urea-based plant DNA miniprep. In: Freeling M (ed.)Walbot V., The Maize Handbook. Springer-Verlag, New York, pp. 526–528.
Garcia G.M., Stalker H.T. and Kochert G. 1995. Introgression analysis of an interspecific hybrid population in peanuts (Arachis hypogaea L.) using RFLP and RAPD markers. Genome 38: 166–176.
Gepts P. 1993. The use of molecular and biochemical markers in crop evolution studies. Evolutionary Biology 27: 51–94.
Grieshammer U. and Wynne J.C. 1990. Isozyme variability in mature seeds of U. S. peanut cultivars and collections. Peanut Science 18: 72–75.
Halward T.M., Stalker H.T., LaRue E. and Kochert G. 1991. Genetic variation detectable with molecular markers among unadapted germplasm resources of cultivated peanut and related wild species. Genome 34: 1013–1020.
Halward T.M., Stalker H.T., LaRue E. and Kochert G. 1992. Use of single-primer DNA amplification in genetic studies of peanut (Arachis hypogaea L.). Plant Molecular Biology 18: 315–325.
He G.H., Prakash C.S., Jarret R.L., Tuzun S. and Qiu J. 1994. Comparison of gel matrices for resolving PCR-ampli.ed DNA fingerprint profiles. PCR Meth. Appl. 4: 50–51.
He G.H. and Prakash C.S. 1997. Identification of polymorphic DNA markers in cultivated peanut (Arachis hypogaea L.). Euphytica 97: 143–149.
Knauft D.A. and Gorbet D.W. 1989. Genetic diversity among peanut cultivars. Crop Sci. 29: 1417–1422.
Kochert G., Halward T., Branch W.D. and Simpson C.E. 1991. RFLP variability in peanut (Arachis hypogaea L.) cultivars and wild species. Theor. Appl. Genet. 81: 565–570.
Krapovickas A., Fernandez A. and Seeligman P. 1974. Recuperacion de la fertilidad en un hibrido interspeifico esteril de Arachis (Leguminosae). Bonplandia 3: 129–142.
Krapovickas A. and Gregory W.C. 1994. Taxonomia del genero Arachis (Leguminosae). Bonplandia 8: 1–186.
Lacks G.D. and Stalker H.T. 1993. Isozyme analyses of Arachis species and interspecific hybrids. Peanut Science 20: 76–81.
Lanham P.G., Fennell S., Moss J.P. and Powell W. 1992. Detection of polymorphic loci in Arachis germplasm using random amplified polymorphic DNAs. Genome 35: 885–889.
Lanham P.G., Forster B.P., McNicol P., Mossand J.P. and Powell W. 1994. Seed storage protein variation in Arachis species. Genome 37: 487–496.
Lu J. and Pickersgill B. 1993. Isozyme variation and species relationships in peanut and its wild relatives (Arachis L.-Leguminosae). Theor. Appl. Genet. 85: 550–560.
Paik-Ro O.G., Smith R.L. and Knauft D.A. 1992. Restriction fragment length polymorphism evaluation of six peanut species within the Arachis section. Theor. Appl. Genet. 84: 201–208.
Pickersgill B. 1986. Evolution of hierarchical variation patterns under domestication and their taxonomic treatment. In: Style S.T (ed.), Infraspecific Classification of Wild and Cultivated Plants. Clarendon Press, Oxford, pp. 191–209.
Rohlf F.J. 1992. Numerical Taxonomy and Multivariate Analysis System. Applied Biostatistics Inc., New York.
Shattuck-Eidens D.M., Bell R.N., Neuhausen S.L. and Helentjaris T. 1990. DNA sequence variation within maize and melon: Observation from polymerase chain reaction amplification and direct sequency. Genetics 126: 207–217.
Singh A.K. and Moss J.P. 1984. Utilization of wild relatives in genetic improvement of Arachis hypogaea L. 5: Genome analysis in section Arachis and its implications in gene transfer. Theor. Appl. Genet. 68: 355–365.
Singh A.K., Sivaramakrishnan S., Mengesha M.H. and Ramaiah C.D. 1991. Phylogenetic relations in section Arachis based on seed protein profile. Theor. Appl. Genet. 82: 593–597.
Singh A.K., Santosh Gurt and Jambunathan R. 1994. Phylogenetic relationships in the genus Arachis based on seed protein profiles. Euphytica 74: 219–225.
Vos P., Hogers R., Bleeker M., Reijans M., Van de Lee T., Hornes M. et al. 1995. AFLP: A new technique for DNA fingerprinting. Nucleic Acids Research 21: 4407–4414.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
He, G., Prakash, C. Evaluation of genetic relationships among botanical varieties of cultivated peanut (Arachis hypogaea L.) using AFLP markers. Genetic Resources and Crop Evolution 48, 347–352 (2001). https://doi.org/10.1023/A:1012019600318
Issue Date:
DOI: https://doi.org/10.1023/A:1012019600318