Abstract
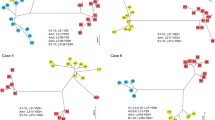
Interferon treatment of chronic hepatitis C virus infection is successful in a minority of patients. The sequence of the interferon sensitivity determining region (ISDR) of the NS5A protein may determine the outcome of therapy in patients infected with HCV genotype 1. To determine whether IFN treatment caused selection of ISDR quasispecies and whether sequences bearing the putative IFN-resistance motif (HCV-J sequence) were selected, we examined amino acid changes in the ISDR in patients with HCV of different genotypes with and without therapy. We found that the ISDR sequence was highly variable and variability was greatest in patients with HCV of genotype 1. IFN treatment was found to exert a selection pressure on ISDR quasispecies, but the putative interferon-resistant variant was not enriched in patients of any genotype. Hence factors other than the sequence of the ISDR region played a role in the IFN resistance of these patients.
Similar content being viewed by others
References
WHO Group: Global surveillance and control of hepatitis C. J. Viral Hep 6:35–47, 1999
Poynard T: Management of chronic hepatitis C. J Viral Hep 5:000, 1998
Simmonds P. Variability of hepatitis C virus. Hepatology 21(2):570–583, 1995.
Simmonds P, Alberti A, Alter HJ, Bonino F, Bradley DW, Brechot C, Brouwer JT, Chan SW, Chayama K, Chen DS, et al: A proposed system for the nomenclature of hepatitis C viral genotypes. Hepatology 19(5):1321–1324, 1994 (letter)
Martell M, Esteban JI, Quer J, Genesca J, Weiner A, Esteban R, Guardia J, Gomez J. Hepatitis C virus (HCV) circulates as a population of different but closely related genomes: Quasispecies nature of HCV genome distribution. J Virol 66(5):3225–3229, 1992
Chayama K, Tsubota A, Arase Y, Saitoh S, Ikeda K, Matsumoto T, Hashimoto M, Kobayashi M, Kanda M, Morinaga T, et al. Genotype, slow decrease in virus titer during interferon treatment and high degree of sequence variability of hypervariable region are indicative of poor response to interferon treatment in patients with chronic hepatitis type C. J Hepatol 23(6):648–653, 1995
Enomoto N, Sato C, Asahina Y, et al. Comparison of full length sequences of interferon sensitive and resistant hepatitis C virus 1b: Sensitivity to interferon is conferred by amino acid substitutions in the NS5A region. J Clin Invest 96:224–230, 1995
Zeuzem S, Lee JH, Roth WK: Mutations in the nonstructural 5A gene of European hepatitis C virus isolates and response to interferon alfa [see comments]. Hepatology 25(3):740–744, 1997
Herion D, Hoofnagle JH: The interferon sensitivity determining region: all hepatitis C virus isolates are not the same. [editorial; comment]. Hepatology 25(3):769–771, 1997
Saiz JC, Lopez-Labrador FX, Ampurdanes S, Dopazo J, Forns X, Sanchez-Tapias JM, Rodes J. The prognostic relevance of the nonstructural 5A gene interferon sensitivity determining region is different in infections with genotype 1b and 3a isolates of hepatitis C virus. J Infect Dis 177(4):839–847, 1998
Murakami T, Enomoto N, Kurosaki M, Izumi N, Marumo F, Sato C: Mutations in nonstructural protein 5A gene and response to interferon in hepatitis C virus genotype 2 infection. Hepatology 30(4):1045–1053, 1999
Gale M Jr, Blakely CM, Kwieciszewski B, Tan SL, Dossett M, Tang NM, Korth MJ, Polyak SJ, Gretch DR, Katze MG: Control of PKR protein kinase by hepatitis C virus nonstructural 5A protein: Molecular mechanisms of kinase regulation. Mol Cell Biol 18(9):5208–5218, 1998
Gale MJ Jr, Korth MJ, Tang NM, Tan SL, Hopkins DA, Dever TE, Polyak SJ, Gretch DR, Katze MG: Evidence that hepatitis C virus resistance to interferon is mediated through repression of the PKR protein kinase by the nonstructural 5A protein. Virology 230(2):217–227, 1997
Polyak SJ, Paschal DM, McArdle S, Moradpour D, Gretch DR: Characterization of the effects of hepatitis C virus non-structural 5A protein expression in human cell lines and on interferon-sensitive virus replication. Hepatology 29(4):1262–1271, 1999
Song J, Fujii M, Wang F, Itoh M, Hotta H: The NS5A protein of hepatitis C virus partially inhibits the antiviral activity of interferon. J Gen Virol 80(Pt 4):879–886, 1999
Paterson M, Laxton C, Thomas HC, Ackrill AM, Foster GR: Hepatitis C virus NS5A protein inhibits interferon antiviral activity, but the effects do not correlate with clinical response. Gastroenterology 117:1187–1197, 1999
Pawlotsky JM, Germanidis G, Neumann AU, Pellerin M, Frainais PO, Dhumeaux D: Interferon resistance of hepatitis C virus genotype 1b: Relationship to nonstructural 5A gene quasispecies mutations. J Virol 72(4):2795–2805, 1998
Polyak SJ, McArdle S, Liu SL, Sullivan, DG, Chung, M, Hofgartner, WT Carithers RL Jr, McMahon BJ, Mullins JI, Corey L, and Gretch DR: Evolution of hepatitis C virus quasispecies in hypervariable region 1 and the putative interferon sensitivity-determining region during interferon therapy and natural infection. J. Virol 72:4288–4296, 1998
McHutchison JG, Gordon SC, Schiff ER, Shiffman ML, Lee WM, Rustgi VK, Goodman ZD, Ling MH, Cort S, Albrecht JK: Interferon alfa-2b alone or in combination with ribavirin as initial treatment for chronic hepatitis C. Hepatitis Interventional Therapy Group [see comments]. N Engl J Med 339(21):1485–1492, 1998
el-Zayadi A, Simmonds P, Dabbous H, Prescott L, Selim O, Ahdy A: Response to interferon-alpha of Egyptian patients infected with hepatitis C virus genotype 4. J Viral Hep 3(5):261–264, 1996
Domingo E, Biological significance of viral quasispecies. Viral Hep Rev 2:247–261, 1996
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Paterson, M., Laxton, C., Goldin, R.D. et al. Selection of HCV NS5A Quasispecies During IFN Therapy in Patients with Chronic HCV. Dig Dis Sci 46, 1399–1408 (2001). https://doi.org/10.1023/A:1010623417345
Issue Date:
DOI: https://doi.org/10.1023/A:1010623417345