Abstract
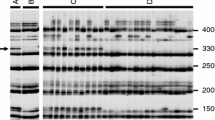
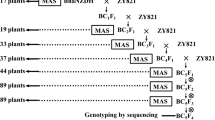
Bulked segregant analysis was used to identify RAPD markers in oilseed rape (Brassica napus L.) that were linked to a male fertility restorer gene for Ogura cytoplasmic male sterility. After screening for polymorphisms using 960 primers, 14 RAPD markers were mapped to a 25 cM region including the restorer locus, a mapping population of 242 F2 individuals being employed. The map was used to select 11 markers that were investigated for polymorphisms between the restorer donor line and 46 recipient lines. A set of four RAPD markers, one in coupling phase with the restorer allele and three with the non-restorer allele, which were informative in all 46 combinations, were used in marker assisted selection of plants homozygous for the restorer allele. A total of 906 homozygous restored plants were found among the 4605 BC1F2 plants analysed. Phenotypic data of a subset of the classified plants was compared with the RAPD data and the expected number of recombinants was calculated from the map data. A close correspondence between the expected and observed numbers of plants with a deviating phenotype was found. Thus, use of a set of dominant RAPD markers provides a way obtaining reliable data for marker-assisted selection.
Similar content being viewed by others
References
Ayliffe AM, Lawrence GJ, Ellis JG, Pryor AJ: Heteroduplex molecules formed between allelic sequences cause nonparental RAPD bands. Nucl Acids Res 22: 1632–1636 (1994).
Delourme R, Bouchereau A, Hubert N, Renard M, Landry BS: Identification of RAPD markers linked to a fertility restorer gene for the Ogura radish cytoplasmic male sterility of rapeseed (Brassica napus L.). Theor Appl Genet 88: 741–748 (1994).
Edwards MD, Page NJ: Evaluation of marker-assisted selection through computer simulation. Theor Appl Genet 88: 376–382 (1994).
Haley SD, Afanador L, Kelly JD: Selection for monogenic pest resistance traits with coupling-and repulsion-phase RAPD markers. Crop Sci 34: 1061–1066 (1994).
Halldén C, Nilsson N-O, Hjerdin A, Rading IM: A gel electrophoresis system adapted for large-scale molecular marker analysis. Anal Biochem 242: 145–147 (1996).
Haldén C, Hansen M, Nilsson N-O, Hjerdin A, Säll T: Competition as a source of errors in RAPD analysis. Theor Appl Genet 93: 1185–1192 (1996).
Hjerdin A, Säll T, Tuvesson S, Halldén C: RFLP markers in the genus Beta: characterization of DNA sequences from a Beta vulgaris library. Genetica 92: 91–99 (1994).
Hospital F, Chevalet C, Mulsant P: Using markers in gene introgression breeding programmes. Genetics 132: 1199–1210 (1992).
Johnson E, Miklas PN, Stavely JR, Martinez-Cruzado JC: Coupling-and repulsion phase RAPDs for marker-assisted selection of PI 181996 rust resistance in common bean. Theor Appl Genet 90: 659–664 (1995).
Kosambi DD: The estimation of map distances from recombination values. Ann Eugen 12: 172–175 (1944).
Michelmore RW, Paran I, Kesseli RV: Identification of markers linked to disease resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci 88: 9828–9832 (1991).
Miklas PN, Afanador L, Kelly JD: Recombination-facilitated RAPD marker-assisted selection for disease resistance in common bean. Crop Sci 36: 86–90 (1996).
Nilsson NO, Halldén C, Hansen M, Hjerdin A, Säll T: Comparing the distribution of RAPD and RFLP markers in a high density linkage map of sugar beet. Genome, in press (1997).
Ohmori T, Murata M, Motoyoshi: Molecular characterization of RAPD and SCAR markers linked to the Tm-1 locus in tomato. Theor Appl Genet 92: 151–156 (1996).
Paran I, Michelmore RW: Development of reliable PCR-based markers linked to downy mildew resistance genes in lettuce. Theor Appl Genet 85: 985–993 (1993).
Penner GA, Bush A, Wise R, Kim W, Domier L, Kasha K, Laroche A, Scoles G, Molnar SJ, Fedak G: Reproducibility of random amplified polymorphic DNA (RAPD) analysis among laboratories. PCR Meth Appl 2: 341–345 (1993).
Rafalski JA, Tingey SV: Genetic diagnostics in plant breeding: RAPDs, microsatellites and machines. Trends Genet 9: 275–280 (1993).
Reiter RS, Williams JGK, Feldmann KA, Rafalski JA, Tingey SV, Scolnik PA: Global and local genome mapping in Arabidopsis thaliana by using recombinant inbred lines and random amplified polymorphic DNAs. Proc Natl Acad Sci USA 89: 1477–1481 (1992).
Stam P: Construction of integrated genetic linkage maps by means of a new computer package, JOINMAP. Plant J 3: 739–744 (1993).
Weeden NF, Timmerman GM, Hemmat M, Kneen BE, Lodhi MA: Inheritance and reliability of RAPDmarkers. In: Hoisington D, McNab A (eds) Proceedings of the Symposium Applied RAPD Technology Plant Breeding, Minneapolis, pp. 12–17 (1992).
Welsh J, McClelland M: Fingerprinting genomes using PCR with arbitrary primers. Nucl Acids Res 18: 7213–7218 (1990).
Williams JGK, Kubelik AR, Livak KL, Rafalski JA, Tingey SV: DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl Acids Res 18: 6531–6535 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hansen, M., Halldén, C., Nilsson, NO. et al. Marker-assisted selection of restored male-fertile Brassica napus plants using a set of dominant RAPD markers. Molecular Breeding 3, 449–456 (1997). https://doi.org/10.1023/A:1009647417936
Issue Date:
DOI: https://doi.org/10.1023/A:1009647417936