Abstract
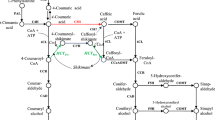
Two types of structurally distinct O-methyltransferases mediate the methylation of hydroxylated monomeric lignin precursors in angiosperms. Caffeate 3-O-methyltransferase (COMT; EC 2.1.1.68) methylates the free acids and caffeoyl CoA 3-O-methyltransferase (CCoAOMT; EC 2.1.1.104) methylates coenzyme A esters. Recently, we reported a novel hydroxycinnamic acid/hydroxycinnamoyl CoA ester O-methyltransferase (AEOMT) from loblolly pine differentiating xylem that was capable of methylating both acid and ester precursors with similar efficiency. In order to determine the possible existence and role of CCoAOMT in lignin biosynthesis in gymnosperms, a 1.3 kb CCoAOMT cDNA was isolated from loblolly pine that showed 79–82% amino acid sequence identity with many angiosperm CCoAOMTs. The recombinant CCoAOMT expressed in Escherichia coli exhibited a significant methylating activity with hydroxycinnamoyl CoA esters whereas activity with hydroxycinnamic acids was insignificant. Moreover, 3.2 times higher catalytic efficiency for methylating caffeoyl CoA over 5-hydroxyferuloyl CoA was observed which could serve as a driving force towards synthesis of guaiacyl lignin. The secondary xylem-specific expression of CCoAOMT was demonstrated using RNA blot analysis, western blot analysis, and O-methyltransferase enzyme assays. In addition, Southern blot analysis indicated that CCoAOMT may exist as a single-copy gene in loblolly pine genome. The transgenic tobacco plants carrying loblolly pine CCoAOMT promoter-GUS fusion localized the site of GUS activity at the secondary xylem tissues. These data suggest that CCoAOMT, in addition to AEOMT, plays an important role in the methylation pathway associated with lignin biosynthesis in loblolly pine.
Similar content being viewed by others
References
Bugos, R.C., Chiang, V.L. and Campbell, W.H. 1991. cDNA cloning, sequence analysis and seasonal expression of ligninbispecific caffeic acid 5-hydroxyferulic acid O-methyltransferase of aspen. Plant Mol. Biol. 17: 1203-1215.
Chiang, V.L. and Funaoka, M. 1990. The dissolution and condensation reactions in guaiacyl and syringyl units in residual lignin during Kraft delignification of sweetgum. Holzforschung 44: 147-155.
Chang, H.M. and Sarkanen, K.V. 1993. Species variation in lignin: effect of species on the rate of Kraft delignification. Tech. Ass. Pulp Paper Ind. 56: 132-134.
Christensen A.H., Sharrock, R.A. and Quail, P.H. 1992. Maize polyubiquitin genes. Plant Mol. Biol. 18: 675-689.
Feuillet, C., Lauvergeat, V., Deswarte, C., Pilate, G., Boudet, A. and Grima-Pettenati, J. 1995. Tissue-and cell-specific expression of a cinnamyl alcohol dehydrogenase promoter in transgenic poplar plants. Plant Mol. Biol. 27: 651-667.
Higuchi, T. 1985. Biosynthesis and Biodegradation of Wood Components. Academic Press, New York.
Higuchi, T., Shimada, M., Nakatsubo, F. and Tanahashi, M. 1977. Differences between biosynthesis of guaiacyl and syringyl lignins in woods. Wood Sci. Technol. 11: 153-167.
Horsch, R.B., Fry, J., Hoffmann, N., Neidermeyer, J., Rogers, S.G. and Fraley, R.T. 1988. Leaf disc transformation. In: S.B. Gelvin, R.A. Schilperoort and D.P.S. Verma (Eds.), Plant Molecular Biology Manual, Kluwer Academic Publishers, Dordrecht, Netherlands, pp. A5: 1-9.
Inoue, K., Sewalt, V.J.H., Ballance, G.M., Ni, W., Stuzer, C. and Dixon, R.A. 1998. Developmental expression and substrate specificities of alfalfa caffeic acid 3-O-methyltransferase and caffeoyl coenzyme A 3-O-methyltransferase in relation to lignification. Plant Physiol. 117: 761-770.
Jefferson, R.A., Kavanagh, T.A. and Bevan, N.W. 1987. GUS fusion: _-glucuronidase as a sensitive and versatile gene fusion maker in higher plants. EMBO J. 6: 3901-3907.
Joshi, C.P. 1987a. Putative polyadenylation signals in nuclear genes of higher plants: a compilation and analysis. Nucl. Acids Res. 15: 9627-9640.
Joshi, C.P. 1987b. An inspection of the domain between putative TATA box and translation start site in seventy-nine plant genes. Nucl. Acids Res. 15: 6643-6653.
Joshi, C.P. and Chiang, V.L. 1988. Conserved sequence motifs in plant S-adenosyl-L-methionine dependent methyltransferases. Plant Mol. Biol. 37: 663-674.
Joshi, C.P., Zhou, H., Huang, X. and Chiang, V. 1997. Context sequences of translation initiation codon in plants. Plant Mol. Biol. 35: 993-1001.
Kloesgen, R.B., Gierl, A., Schwarz-Sommer, Z.S. and Saedler, H. 1986. Molecular analysis of the waxy locus of Zea mays. Mol. Gen. Genet. 203: 237-244.
Kuroda, H., Shimada, M. and Higuchi, T. 1975. Purification and properties of O-methyltransferase involved in the biosynthesis of gymnosperm lignin. Phytochemistry 14: 1759-1763.
Li, L., Popko, J.L., Zhang, X.H., Osakabe, K., Tsai, C.J., Joshi, C.P. and Chiang, V.L. 1997. A novel multifunctional O-methyltransferase implicated in a dual methylation pathway associated with lignin biosynthesis in loblolly pine. Proc. Natl. Acad. Sci. USA 94: 5461-5466.
Martz, F., Maury, S., Pincon, G. and Legrand, M. 1998. cDNA cloning, substrate specificity and expression study of tobacco caffeoyl-CoA 3-O-methyltransferase, a lignin biosynthetic enzyme. Plant Mol. Biol. 36: 427-437.
McElroy, D., Zhang, W., Cao, J. and Wu, R. 1990. Isolation of an efficient actin promoter for use in rice transformation. Plant Cell 2: 163-171.
Seguin, A., Laible, G., Leyva, A., Dixon, R.A. and Lamb, C. 1997. Characterization of a gene encoding a DNA-binding protein that interacts in vitro with vascular specific cis elements of the PAL promoter. Plant Mol. Biol. 35: 281-291.
Shimada, M., Fushiki, H. and Higuchi, T. 1972. Omethyltransferase activity from Japanese black pine. Phytochemistry 11: 2657-2662.
Wakamiya, I., Newton, R.J., Johnston, J.S. and Price, H.J. 1993. Genome sizes and environmental factors in the genus Pinus. Am. J. Bot. 80: 1235-1241.
Whetten, R.W., MacKay, J.J. and Sederoff, R.R. 1998. Recent advances in understanding lignin biosynthesis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 49: 585-609.
Ye, Z.-H. and Varner, J.E. 1995. Differential expression of two Omethyltransferases in lignin biosynthesis in Zinnia elegans. Plant Physiol. 108: 459-467.
Ye, Z.-H., Kneusel, R.E., Matern, U. and Varner, J.E. 1994. An alternative methylation pathway in lignin biosynthesis in Zinnia. Plant Cell 6: 1427-1439 (1994).
Zhang, X.H. and Chiang, V.L. 1997. Molecular cloning of 4-coumarate:coenzyme A ligase in loblolly pine and the roles of this enzyme in the biosynthesis of lignin in compression wood. Plant Physiol. 113: 65-74.
Zhong, R., Morrison, W.H., Negrel, J. and Ye, Z.-H. 1998. Dual methylation pathways in lignin biosynthesis. Plant Cell 10: 2033-2045.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Li, L., Osakabe, Y., Joshi, C.P. et al. Secondary xylem-specific expression of caffeoyl-coenzyme A 3-O-methyltransferase plays an important role in the methylation pathway associated with lignin biosynthesis in loblolly pine. Plant Mol Biol 40, 555–565 (1999). https://doi.org/10.1023/A:1006244325250
Issue Date:
DOI: https://doi.org/10.1023/A:1006244325250