Abstract
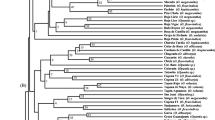
RAPD analysis was applied to reveal the genetic diversities of 4 speciesof subg. Lithocerasus within the genus Prunus using 40accessions representing the subgenera Prunophora, Amygdalus,Lithocerasus and Cerasus. The accession of subg. Lithocerasus are phenotypically similar to members of subgenus Cerasus but different with them in interspecific crossing tests and isozymeprofiles. Two major clusters, `Prunophora and Amygdalus group' and `Cerasus group' were constructed in the phenogram. We revealthat the examined 4 species of subg. Lithocerasus species; P.tomentosa Thunb., P. japonica Thunb., P. glandulosa Thunb.and P. besseyi Bailey, were genetically closer to the members ofsubgenera Prunophora and Amygdalus than to subg. Cerasus.
Similar content being viewed by others
References
Badenes, M.L. & D.E. Parfitt, 1995. Phylogenetic relationships of cultivated Prunus species from an analysis of chloroplast DNA variation. Theor Appl Genet 90: 1035-1041.
Chaparro, J.X., D.J. Werner, D. O' Malley & R.R. Sederoff, 1994. Targeted mapping and linkage analysis of morphological isozyme, and RAPD markers in peach. Theor Appl Genet 87: 805-815.
Dirlewanger, E., T. Pascal, C. Zuger & J. Kervella, 1996. Analysis of molecular markers associated with powdery mildew resistance genes in peach (Prunus persica (L.) Batsch) × Prunus davidiana Carr. hybrids. Theor Appl Genet 93: 909-919.
Doyle, J.J. & J.L. Doyle, 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phyto Bull 19: 11-15.
Garley, B., 1980. The sand cherry: its origin, improvement and nomenclature. Fruit Var J 34: 13-17.
Ingram, C., 1948. Ornamental cherries. London.
Kataoka, I., A. Sugiura & T. Tomana, 1988. Interspecific hybridization between Microcerasus and other Prunus spp. J Jpn Soc Hort Sci 56: 398-407.
Maniatis, T., T. Fritsch & E.F. Sambrook, 1989. Molecular cloning, Second Ed. 6.14-6.15. Cold Spring Harbor Laboratory Press, U.S.A.
Mowrey, B.D. & D.J. Werner, 1990. Phylogenetic relationships among species of Prunus as inferred by isozyme markers. Theor Appl Genet 80: 129-133.
Rehder, A., 1940. Prunus. In 'Manual of cultivated trees and shrubs hardly in North America. 2nd ed., Macmillan, New York, pp. 452-481.
Shimada, T., T. Haji, M. Yamaguchi, T. Takeda, K. Nomura & M. Yoshida, 1994. Classification of mume (Prunus mume Sieb. et Zucc.) by RAPD assay. J Jap Hort Sci 63: 543-551.
Shimada, T., H. Hayama, M. Yamaguchi, K. Nishimura & M. Yoshida, 1997. Phylogenetic studies on Prunus species by Southern blot analysis of cpDNA. Mem. Grad. School Sci. & Technol., Kobe Univ. 15A: 59-71
Sokal, R.R. & C.D. Michener, 1958. A statistical method for evaluating systematic relationships. Univ Kansas Sci Bull 38: 1409-1438.
Tanaka, Y., 1934. Studies on the graft compatibility in Rosaceae and their related species. Agr and Hort 9: 2592-2616 (in Japanese).
Uematsu, C., T. Sasakuma & Y. Ogihara, 1991. Phylogenetic relationships in the stone fruit group of Prunus as revealed by restriction fragment analysis of chloroplast DNA. Japan J Genet 66: 59-69.
Zhen-Xiang Lu, G.L. Reighard & W.V. Baird, 1996. Identification of peach rootstock cultivars by RAPD markers. Hort Sci 31: 127-129.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shimada, T., Hayama, H., Nishimura, K. et al. The genetic diversities of 4 species of subg. Lithocerasus (Prunus, Rosaceae) revealed by RAPD analysis. Euphytica 117, 85–90 (2001). https://doi.org/10.1023/A:1004193327542
Issue Date:
DOI: https://doi.org/10.1023/A:1004193327542