Abstract
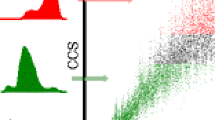
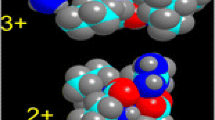
Molecular dynamics (MD) is an essential tool for correlating collision cross-section data determined by ion mobility spectrometry (IMS) with candidate (calculated) structures. Conventional methods used for ion structure determination rely on comparing the measured cross-sections with the calculated collision cross-section for the lowest energy structure(s) taken from a large pool of candidate structures generated through multiple tiers of simulated annealing. We are developing methods to evaluate candidate structures from an ensemble of many conformations rather than the lowest energy structure. Here, we describe computational simulations and clustering methods to assign backbone conformations for singly-protonated ions of the model peptide (NH2-Met-Ile-Phe-Ala-Gly-Ile-Lys-COOH) formed by both MALDI and ESI, and compare the structures of MIFAGIK derivatives to test the ‘sensitivity’ of the cluster analysis method. Cluster analysis suggests that [MIFAGIK + H]+ ions formed by MALDI have a predominantly turn structure even though the low-energy ions prefer partial helical conformers. Although the ions formed by ESI have collision cross-sections that are different from those formed by MALDI, the results of cluster analysis indicate that the ions backbone structures are similar. Chemical modifications (N-acetyl, methylester as well as addition of Boc or Fmoc groups) to MIFAGIK alter the distribution of various conformers; the most dramatic changes are observed for the [M + Na]+ ion, which show a strong preference for random coil conformers owing to the strong solvation by the backbone amide groups.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ruotolo, B. T.; Giles, K.; Campuzano, I.; Sandercock, A. M.; Bateman, R. H.; Robinson, C. V. Evidence for Macromolecular Protein Rings in the Absence of Bulk Water. Science 2005, 310, 1658–1661.
Kaddis, C. S.; Lomeli, S. H.; Yin, S.; Berhane, B.; Apostol, M. I.; Kickhoefer, V. A.; Rome, L. H.; Loo, J. A. Sizing Large Proteins and Protein Complexes by Electrospray Ionization Mass Spectrometry and Ion Mobility. J. Am. Soc. Mass Spectrom. 2007, 18, 1206–1216.
Kaddis, C. S.; Loo, J. A. Native protein MS and Ion Mobility: Large Flying Proteins with ESI. Anal. Chem. 2007, 79, 1778–1784.
Loo, J. A.; Kaddis, C. S. Direct characterization of protein complexes by electrospray ionization mass spectrometry and ion mobility analysis; John Wiley & Sons, Inc.: Hoboken, NJ, 2007, pp. 1–23.
Ruotolo, B. T.; Hyung, S.-J.; Robinson, P. M.; Giles, K.; Bateman, R. H.; Robinson, C. V. Ion mobility-mass spectrometry reveals long-lived, unfolded intermediates in the dissociation of protein complexes. Angew. Chem. Int. Ed. 2007, 46, 8001–8004.
Ruotolo, B. T.; Benesch, J. L. P.; Sandercock, A. M.; Hyung, S.-J.; Robinson, C. V. Ion mobility-mass spectrometry analysis of large protein complexes. Nat. Protocols 2008, 3, 1139–1152.
Uetrecht, C.; Versluis, C.; Watts, N. R.; Wingfield, P. T.; Steven, A. C.; Heck, A. J. R. Stability and shape of hepatitis B virus capsids in vacuo. Angew. Chem. Int. Ed. 2008, 47, 6247–6251.
Barrera, N. P.; Di Bartolo, N.; Booth, P. J.; Robinson, C. V. Micelles Protect Membrane Complexes from Solution to Vacuum. Science 2008, 321, 243–246.
Clemmer, D. E.; Hudgins, R. R.; Jarrold, M. F. Naked Protein Conformations: Cytochrome c in the Gas Phase. J. Am. Chem. Soc. 1995, 117, 10141–10142.
Shelimov, K. B.; Jarrold, M. F. Conformations, Unfolding, and Refolding of Apomyoglobin in Vacuum: An Activation Barrier for Gas-Phase Protein Folding. J. Am. Chem. Soc. 1997, 119, 2987–2994.
Hudgins, R. R.; Ratner, M. A.; Jarrold, M. F. Design of Helices hat are Stable in Vacuo. J. Am. Chem. Soc. 1998, 120, 12974–12975.
Ruotolo, B. T.; Verbeck, G. F.; Thomson, L. M.; Gillig, K. J.; Russell, D. H. Observation of conserved solution-phase secondary structure in gas-phase tryptic peptides. J. Am. Chem. Soc. 2002, 124, 4214–4215.
Ruotolo, B. T.; Russell, D. H. Gas-phase conformations of proteolytically derived protein fragments: Influence of solvent on peptide conformation. J. Phys. Chem. B 2004, 108, 15321–15331.
McLean, J. A.; Ruotolo, B. T.; Gillig, K. J.; Russell, D. H. Ion mobility-mass spectrometry: A new paradigm for proteomics. Int. J. Mass Spectrom. 2005, 240, 301–315.
Tao, L.; McLean, J. R.; McLean, J. A.; Russell, D. H. A Collision Cross-Section Database of Singly-Charged Peptide Ions. J. Am. Soc. Mass Spectrom. 2007, 18, 1232–1238.
Ruotolo, B. T.; Verbeck, G. F.; Thomson, L. M.; Woods, A. S.; Gillig, K. J.; Russell, D. H. Distinguishing between phosphorylated and nonphosphorylated peptides with ion mobility-mass spectrometry. J. Proteome Res. 2002, 1, 303–306.
Ruotolo, B. T.; Gillig, K. J.; Woods, A. S.; Egan, T. F.; Ugarov, M. V.; Schultz, J. A.; Russell, D. H. Analysis of phosphorylated peptides by ion mobility-mass spectrometry. Anal. Chem. 2004, 76, 6727–6733.
McLean, J. R.; McLean, J. A.; Wu, Z.; Becker, C.; Pérez, L. M.; Pace, C. N.; Scholtz, J. M.; Russell, D. H. Factors that Influence Helical Preferences for Singly-Charged Gas-Phase Peptide Ions: The Effects of Multiple Charge-Carrying Sites. J. Am. Chem. Soc., submitted.
Wilson, S. R.; Cui, W. Applications of simulated annealing to peptides. Biopolymers 1990, 29, 225–235.
Fernandez-Lima, F. A.; Wei, H.; Gao, Y. Q.; Russell, D. H. On the structure elucidation using IMS and Molecular Dynamics. J. Phys. Chem. A, submitted.
Stearns, J. A.; Boyarkin, O. V.; Rizzo, T. R. Spectroscopic signatures of gas-phase helices: Ac-Phe-(Ala)5-Lys-H+ and Ac-Phe-(Ala)10-Lys-H+. J. Am. Chem. Soc. 2007, 129, 13820–13821.
Stearns, J. A.; Guidi, M.; Boyarkin, O. V.; Rizzo, T. R. Conformation-specific infrared and ultraviolet spectroscopy of tyrosine-based protonated dipeptides. J. Chem. Phys. 2007, 127, 154322.
Stearns, J. A.; Mercier, S.; Seaiby, C.; Guidi, M.; Boyarkin, O. V.; Rizzo, T. R. Conformation-specific spectroscopy and photodissociation of cold, protonated tyrosine and phenylalanine. J. Am. Chem. Soc. 2007, 129, 11814–11820.
Damsbo, M.; Kinnear, B. S.; Hartings, M. R.; Ruhoff, P. T.; Jarrold, M. F.; Ratner, M. A. Application of evolutionary algorithm methods to polypeptide folding: Comparison with experimental results for unsolvated Ac-(Ala-Gly-Gly)5-LysH+. Natl. Acad. Sci. U.S.A. 2004, 101, 7215–7222.
Carpino, L. A.; Han, G. Y. 9-Fluorenylmethoxycarbonyl function, a new base-sensitive amino-protecting group 1970, 92, 5748–5749.
Reid, G.; Simpson, R.; O’Hair, R. J. A mass spectrometric and ab initio study of the pathways for dehydration of simple glycine and cysteine-containing peptide [M + H]+ ions. J. Am. Soc. Mass Spectrom. 1998, 9, 945–956.
Gillig, K. J.; Ruotolo, B. T.; Stone, E. G.; Russell, D. H.; Fuhrer, K.; Gonin, M.; Schultz, J. A. Coupling high-pressure MALDI with ion mobility/orthogonal time-of-flight mass spectrometry. Anal. Chem. 2000, 72, 3965–3971.
Stone, E.; Gillig, K. J.; Ruotolo, B.; Fuhrer, K.; Gonin, M.; Schultz, A.; Russell, D. H. Surface-induced dissociation on a MALDI-ion mobility-orthogonal time-of-flight mass spectrometer: Sequencing peptides from an “in-solution” protein digest. Anal. Chem. 2001, 73, 2233–2238.
Mason, E. A.; McDaniel, E. W. Transport Properties of Ions in Gases; Wiley: New York, 1988, pp. 1–29.
Sawyer, H. A.; Marini, J. T.; Stone, E. G.; Ruotolo, B. T.; Gillig, K. J.; Russell, D. H. The structure of gas-phase bradykinin fragment 1-5 (RPPGF) ions: an ion mobility spectrometry and H/D exchange ion-molecule reaction chemistry study. J. Am. Soc. Mass Spectrom. 2005, 16, 893–905.
Shvartsburg, A. A.; Jarrold, M. F. An exact hard-spheres scattering model for the mobilities of polyatomic ions. Chem. Phys. Lett. 1996, 261, 86–91.
Dahl, D. B. Model-Based Clustering for Expression Data via a Dirichlet Process Mixture Model; Cambridge University Press: Cambridge, 2006, 201–218.
Binder, D. A. Bayesian Cluster Analysis. Biometrika 1978, 65, 31–38.
Hubert, L.; Arabie, P. Comparing Partitions. J. Classification 1985, 2, 193–218.
Valentine, S. J.; Counterman, A. E.; Clemmer, D. E. A database of 660 peptide ion cross sections: Use of intrinsic size parameters for bona fide predictions of cross sections. J. Am. Soc. Mass Spectro. 1999, 10, 1188–1211.
Lee, S.-W.; Kim, H. S.; Beauchamp, J. L. Salt Bridge Chemistry Applied to Gas-Phase Peptide Sequencing: Selective Fragmentation of Sodiated Gas-Phase Peptide Ions Adjacent to Aspartic Acid Residues. J. Am. Chem. Soc. 1998, 120, 3188–3195.
Ganesh, S.; Jayakumar, R. Role of N-t-Boc group in helix initiation in a novel tetrapeptide. J. Peptide Res. 2002, 59, 249–256.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online May 5, 2009
Rights and permissions
About this article
Cite this article
Tao, L., Dahl, D.B., Pérez, L.M. et al. The contributions of molecular framework to IMS collision cross-sections of gas-phase peptide ions. J Am Soc Mass Spectrom 20, 1593–1602 (2009). https://doi.org/10.1016/j.jasms.2009.04.018
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2009.04.018