Abstract
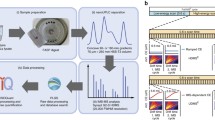
To enable the development of a tandem mass spectrometry (MS/MS) based methodology for selective protein identification and differential quantitative analysis, a novel derivatization strategy is proposed, based on the formation of a “fixed-charge” sulfonium ion on the side-chain of a methionine amino acid residue contained within a protein or peptide of interest. The gas-phase fragmentation behavior of these side chain fixed charge sulfonium ion containing peptides is observed to result in exclusive loss of the derivatized side chain and the formation of a single characteristic product ion, independently of charge state or amino acid composition. Thus, fixed charge containing peptide ions may be selectively identified from complex mixtures, for example, by selective neutral loss scan mode MS/MS methods. Further structural interrogation of identified peptide ions may be achieved by subjecting the characteristic MS/MS product ion to multistage MS/MS (MS3) in a quadrupole ion trap mass spectrometer, or by energy resolved “pseudo” MS3 in a triple quadrupole mass spectrometer. The general principles underlying this fixed charge derivatization approach are demonstrated here by MS/MS, MS3 and “pseudo” MS3 analysis of side chain fixed-charge sulfonium ion derivatives of peptides containing methionine formed by reaction with phenacylbromide. Incorporation of “light” and “heavy” isotopically encoded labels into the fixed-charge derivatives facilitates the application of this method to the quantitative analysis of differential protein expression, via measurement of the relative abundances of the neutral loss product ions generated by dissociation of the light and heavy labeled peptide ions. This approach, termed “selective extraction of labeled entities by charge derivatization and tandem mass spectrometry” (SELECT), thereby offers the potential for significantly improved sensitivity and selectivity for the identification and quantitative analysis of peptides or proteins containing selected structural features, without requirement for extensive fractionation or otherwise enrichment from a complex mixture prior to analysis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Blackstock, W. P.; Weir, M. P. Proteomics: Quantitative and physical mapping of cellular proteins. Trends Biotechnol. 1999, 17, 121–127.
Mann, M.; Hendrickson, R. C.; Pandey, A. Analysis of proteins and proteomes by mass spectrometry. Annu. Rev. Biochem. 2001, 70, 437–473.
Aebersold, R.; Goodlett, D. R. Mass spectrometry in proteomics. Chem. Rev. 2001, 101, 269–295.
Hunt, D. F.; Yates, J. R.; Shabanowitz, J.; Winston, S.; Hauer, C. R. Protein sequencing by tandem mass spectrometry. Proc. Natl. Acad. Sci. U.S.A. 1986, 83, 6233–6237.
Eng, J. K.; McCormack, A. L.; Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 1994, 5, 976–989.
Mann, M.; Wilm, M. Error-tolerant identification of peptides in sequence databases by peptide sequence tags. Anal. Chem. 1994, 66, 4390–4399.
Spahr, C. S.; Susin, S. A.; Bures, E. J.; Robinson, J. H.; Davis, M. T.; McGinley, M. D.; Kroemer, G.; Patterson, S. D. Simplification of complex peptide mixtures for proteomic analysis: Reversible biotinylation of cysteinyl peptides. Electrophoresis. 2000, 21, 1635–1650.
Washburn, M. P.; Wolters, D.; Yates, J. R. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 2001, 19, 242–247.
Spahr, C. S.; Davis, M. T.; McGinley, M. D.; Robinson, J. H.; Bures, E. J.; Beierle, J.; Mort, J.; Courchesne, P. L.; Chen, K.; Wahl, R. C.; Yu, W.; Luethy, R.; Patterson, S. D. Towards defining the urinary proteome using liquid chromatography-tandem mass spectrometry: I. Profiling an unfractionated tryptic digest. Proteomics 2001, 1, 93–107.
Davis, M. T.; Spahr, C. S.; McGinley, M. D.; Robinson, J. H.; Bures, E. J.; Beierle, J.; Mort, J.; Yu, W.; Luethy, R.; Patterson, S. D. Towards defining the urinary proteome using liquid chromatography-tandem mass spectrometry: II. Limitations of complex mixture analyses. Proteomics 2001, 1, 108–117.
Simpson, R. J.; Connelly, L. M.; Eddes, J. S.; Pereira, J. J.; Moritz, R. L.; Reid, G. E. Proteomic analysis of the human colon carcinoma cell line (LIM 1215): Development of a membrane protein database. Electrophoresis 2000, 21, 1707–1732.
Oda, Y.; Huang, K.; Cross, F. R.; Cowburn, D.; Chait, B. T. Accurate quantitation of protein expression and site-specific phosphorylation. Proc. Natl. Acad. Sci. U.S.A. 1999, 96, 6591–6596.
Pasa-Tolic, L.; Jensen, P. K.; Anderson, G. A.; Lipton, M. S.; Peden, K. K.; Martinovic, S.; Tolic, N.; Bruce, J. E.; Smith, R. D. High throughput proteome-wide precision measurements of protein expression using mass spectrometry. J. Am. Chem. Soc. 1999, 121, 7949–7950.
Ong, S.-E.; Blagoev, B.; Kratchmarova, I.; Bach-Kristensen, D.; Steen, H.; Pandey, A.; Mann, M. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteom. 2002, 1, 376–386.
Gygi, S. P.; Rist, B.; Gerber, S. A.; Turecek, F.; Gelb, M. H.; Aebersold, R. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 1999, 17, 994–999.
Smolka, M.; Zhou, H.; Aebersold, R. Quantitative protein profiling using two-dimensional gel electrophoresis, isotope-coded affinity tag labeling, and mass spectrometry. Mol. Cell. Proteom. 2002, 1, 19–29.
Shiio, Y.; Donohoe, S.; Yi, E. C.; Goodlett, D. R.; Aebersold, R.; Eisenman, R. N. Quantitative proteomic analysis of Myc oncoprotein function. EMBO J. 2002, 21, 5088–5096.
Parker, K. C.; Patterson, D.; Williamson, B.; Marchese, J.; Graber, A.; He, F.; Jacobson, A.; Juhasz, P.; Martin, S. Depth of Proteome Issues: A yeast isotope-coded affinity tag reagent study. Mol. Cell. Proteom. 2004, 3, 625–659.
Mirgorodskaya, O. A.; Kozmin, Y. P.; Titov, M. I.; Korner, R.; Sonksen, C. P.; Roepstorff, P. Quantitation of peptides and proteins by matrix-assisted laser desorption/ionization mass spectrometry using 18O-labeled internal standards. Rapid Commun. Mass Spectrom. 2000, 14, 1226–1232.
Stewart, I. I.; Thomson, T.; Figeys, D. 18O Labeling: A tool for proteomics. Rapid Commun. Mass Spectrom. 2001, 15, 2456–2465.
Yao, X.; Freas, A.; Ramirez, J.; Demirev, P. A.; Fenselau, C. Proteolytic 18O Labeling for comparative oroteomics: Model studies with two serotypes of adenovirus. Anal. Chem. 2001, 73, 2836–2842.
Muenchbach, M.; Quadroni, M.; Miotto, G.; James, P. Quantitation and facilitated de novo sequencing of proteins by isotopic N-terminal labeling of peptides with a fragmentation-directing moiety. Anal. Chem. 2000, 72, 4047–4057.
Sechi, S. A method to identify and simultaneously determine the relative quantities of proteins isolated by gel electrophoresis. Rapid Commun. Mass Spectrom. 2002, 16, 1416–1424.
Geng, M.; Ji, J.; Regnier, F. E. Signature-peptide approach to detecting proteins in complex mixtures. J. Chromatogr. A. 2000, 870, 295–313.
Chakraborty, A.; Regnier, F. E. Global internal standard technology for comparative proteomics. J. Chromatogr. A 2002, 949, 173–184.
Ren, D.; Penner, N. A.; Slentz, B. E.; Mirzaei, H.; Regnier, F. E. Evaluating immobilized metal affinity chromatography for the selection of histidine-containing peptides in comparative proteomics. J. Proteom. Res. 2003, 2, 321–329.
Goodlett, D. R.; Keller, A.; Watts, J. D.; Newitt, R.; Yi, E. C.; Purvine, S.; Eng, J.; von Haller, P.; Aebersold, R.; Kolker, E. Differential stable isotope labeling of peptides for quantitation and de novo sequence derivation. Rapid Commun. Mass Spectrom. 2001, 15, 1214–1221.
Gehanne, S.; Cecconi, D.; Carboni, L.; Righetti, P. G.; Domenici, E.; Hamdan, M. Quantitative analysis of two-dimensional gel-separated proteins using isotopically marked alkylating agents and matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun. Mass Spectrom. 2002, 16, 1692–1698.
Shen, M.; Guo, L.; Wallace, A.; Fitzner, J.; Eisenman, J.; Jacobson, E.; Johnson, R. S. Isolation and isotope labeling of cysteine-and methionine-containing tryptic peptides: Application to the study of cell surface proteolysis. Mol. Cell. Proteom. 2003, 2, 315–324.
Kuyama, H.; Watanabe, M.; Toda, C.; Ando, E.; Tanaka, K.; Nishimura, O. An approach to quantitative proteome analysis by labeling tryptophan residues. Rapid Commun. Mass Spectrom. 2003, 17, 1642–1650.
Qiu, Y.; Sousa, E. A.; Hewick, R. M.; Wang, J. H. Acid-labile isotope-coded extractants: A class of reagents for quantitative mass spectrometric analysis of complex protein mixtures. Anal. Chem. 2002, 74, 4969–4979.
Julka, S.; Regnier, F. Quantification in proteomics through stable isotope coding: A review. J. Proteom. Res. 2004, 3, 350–363.
Hansen, K. C.; Schmitt-Ulms, G.; Chalkley, R.; Hirsch, J.; Baldwin, M. A.; Burlingame, A. L. Mass spectrometric analysis of protein mixtures at low levels using cleavable 13C-isotope-coded affinity tag and multidimensional chromatography. Mol. Cell. Proteom. 2003, 2, 299–314.
Zhou, H.; Ranish, J. A.; Watts, J. D.; Aebersold, R. Quantitative proteome analysis by solid-phase isotope tagging and mass spectrometry. Nat. Biotechnol. 2002, 20, 512–515.
Zhang, R.; Sioma, C. S.; Wang, S.; Regnier, F. E. Fractionation of isotopically labeled peptides in quantitative proteomics. Anal. Chem. 2001, 73, 5142–5149.
Zhang, R.; Sioma, C. S.; Thompson, R. A.; Xiong, L.; Regnier, F. E. Controlling deuterium isotope effects in comparative proteomics. Anal. Chem. 2002, 74, 3662–3669.
Krutchinsky, A. N.; Chait, B. T. On the nature of the chemical noise in MALDI mass spectra. J. Am. Soc. Mass Spectrom. 2002, 13, 129–134.
Washburn, M. P.; Ulaszek, R. R.; Yates, J. R. Reproducibility of quantitative proteomic analyses of complex biological mixtures by multidimensional protein identification technology. Anal. Chem. 2003, 75, 5054–5061.
Washburn, M. P.; Koller, A.; Oshiro, G.; Ulaszek, R. R.; Plouffe, D.; Deciu, C.; Winzeler, E.; Yates, J. R. Protein pathway and complex clustering of correlated mRNA and protein expression analyses inSaccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 2003, 100, 3107–3112.
Thompson, A.; Schaefer, J.; Kuhn, K.; Kiene, S.; Schwarz, J.; Schmidt, G.; Johnstone, R.; Neumann, T.; Hamon, C. Tandem Mass Tags: A novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 2003, 75, 1895–1904.
Schwartz, J. C.; Wade, A. P.; Enke, C. G.; Cooks, R. G. Systematic delineation of scan modes in multidimensional mass spectrometry. Anal. Chem. 1990, 62, 1809–1818.
Wilm, M.; Neubauer, G.; Mann, M. Parent ion scans of unseparated peptide mixtures. Anal. Chem. 1996, 68, 527–533.
Carr, S. A.; Huddleston, M. J.; Annan, R. S. Selective detection and sequencing of phosphopeptides at the femtomole level by mass spectrometry. Anal. Biochem. 1996, 239, 180–192.
Weckwerth, W.; Willmitzer, L.; Fiehn, O. Comparative quantification and identification of phosphoproteins using stable isotope labeling and liquid chromatography/mass spectrometry. Rapid Commun. Mass Spectrom. 2000, 14, 1677–1681.
Schlosser, A.; Pipkorn, R.; Bossemeyer, D.; Lehmann, W. D. Analysis of protein phosphorylation by a combination of elastase digestion and neutral loss tandem mass spectrometry. Anal. Chem. 2001, 73, 170–176.
Steen, H.; Kuster, B.; Fernandez, M.; Pandey, A.; Mann, M. Detection of tyrosine phosphorylated peptides by precursor ion scanning quadrupole TOF mass spectrometry in positive ion mode. Anal. Chem. 2001, 73, 1440–1448.
Molloy, M. P.; Andrews, P. C. Phosphopeptide derivatization signatures to identify serine and threonine phosphorylated peptides by mass spectrometry. Anal. Chem. 2001, 73, 5387–5394.
Li, W.; Boykins, R. A.; Backlund, P. S.; Wang, G.; aChen, H.-C. Identification of phosphoserine and phosphothreonine as cysteic acid and β-methylcysteic acid residues in peptides by tandem mass spectrometric sequencing. Anal. Chem. 2002, 74, 5701–5710.
Steen, H.; Mann, M. A new derivatization strategy for the analysis of phosphopeptides by precursor ion scanning in positive ion mode. J. Am. Soc. Mass Spectrom. 2002, 13, 996–1003.
Back, J. W.; Hartog, A. F.; Dekker, H. L.; Muijsers, A. O.; de Koning, L. J.; de Jong, L. A new crosslinker for mass spectrometric analysis of the quaternary structure of protein complexes. J. Am. Soc. Mass Spectrom. 2001, 12, 222–227.
Dongre, A. R.; Jones, J. L.; Somogyi, A.; Wysocki, V. H. Influence of peptide composition, gas-phase basicity, and chemical modification on fragmentation efficiency: Evidence for the mobile proton model. J. Am. Chem. Soc. 1996, 118, 8365–8374.
O’Hair, R. A. J. The role of nucleophile-electrophile interactions in the unimolecular and bimolecular gas-phase ion chemistry of peptides and related systems. J. Mass Spectrom. 2000, 35, 1377–1381.
Schlosser, A.; Lehmann, W. D. Five-membered ring formation in unimolecular reactions of peptides: A key structural element controlling low-energy collision-induced dissociation of peptides. J. Mass Spectrom. 2000, 35, 1382–1390.
Wysocki, V. H.; Tsaprailis, G.; Smith, L. L.; Breci, L. A. Mobile and localized protons: A framework for understanding peptide dissociation. J. Mass Spectrom. 2000, 35, 1399–1406.
Kapp, E. A.; Schütz, F.; Reid, G. E.; Eddes, J. S.; Moritz, R. L.; O’Hair, R. A. J.; Speed, T. P.; Simpson, R. J. Mining a tandem mass spectrometry database to determine the trends and global factors influencing peptide fragmentation. Anal. Chem. 2003, 75, 6251–6264.
Reid, G. E.; Roberts, K. D.; Kapp, E. A.; Simpson, R. J. Statistical and mechanistic approaches to understanding the gas-phase fragmentation behavior of methionine sulfoxide containing peptides. J. Proteom. Res. 2004, 3, 751–759.
Knapp, D. R. Chemical derivatization for mass spectrometry. Methods Enzymol. 1990, 193, 314–329.
Anderegg, R. J. Derivatization in mass spectrometry: Strategies for controlling fragmentation. Mass Spectrom. Rev. 1988, 7, 395–424.
Roth, K. D. W.; Huang, Z.-H.; Sadagopan, N.; Watson, J. T. Charge derivatization of peptides for analysis by mass spectrometry. Mass Spectrom. Rev. 1998, 17, 255–274.
Sadagopan, N.; Watson, J. T. Mass spectrometric evidence for mechanisms of fragmentation of charge-derivatized peptides. J. Am. Soc. Mass. Spectrom. 2001, 12, 399–409.
Spengler, B.; Luetzenkirchen, F.; Metzger, S.; Chaurand, P.; Kaufmann, R.; Jeffery, W.; Bartlet-Jones, M.; Pappin, D. J. C. Peptide sequencing of charged derivatives by post-source decay MALDI mass spectrometry. Int. J. Mass Spectrom. Ion Processes 1997, 169/170, 127–140.
Keough, T.; Youngquist, R. S.; Lacey, M. P. Sulfonic acid derivatives for peptide sequencing by MALDI MS. Anal. Chem. 2003, 75, A156-A165.
Frechet, J. M. J.; Farrall, M. J.; Nuyens, L. J. Polymeric reagents: II. Synthesis and applications of crosslinked poly(vinylpyridinium hydrobromide perbromide) resins. J. Macromol. Sci. Chem. 1977, A11, A507-A514.
Liu, T.-H. The role of sulfur in proteins. In The Proteins, Vol. III; Neurath, H.; Hill, R. L.; Boeder, C.-L., Eds.; Academic: New York, 1977; Chap III, pp 239–402.
Stirling, C. J. M. The Chemistry of Functional Groups: The Chemistry of the Sulfonium Group, Pt. 1. John Wiley and Sons: Chichester, UK, 1981.
Lundblad, R. L. Techniques in protein modification. In The Modification of Methionine; CRC Press: Boca Raton, Florida, 1995; Chap. VIII.
Gundlach, H. G.; Moore, S.; Stein, W. H. The reaction of iodoacetate with methionine. J. Biol. Chem. 1959, 234, 1761–1764.
Rogers, G. A.; Shaltiel, N.; Boyer, P. D. Facile alkylation of methionine by benzyl bromide and demonstration of fumarase inactivation accompanied by alkylation of a methionine residue. J. Biol. Chem. 1976, 251, 5711–5717.
Toennies, G.; Kolb, J. J. Methionine studies: VII. Sulfonium derivatives. J. Am. Chem. Soc. 1945, 67, 849–851.
Lavine, T. F.; Floyd, N. F.; Cammaroti, M. S. The formation of sulfonium salts from alcohols and methionine in sulfuric acid. J. Biol. Chem. 1954, 207, 107–117.
Lawson, W. B.; Schramm, H. J. Modification of a methionine residue near the active site of chymotrypsin. J. Am. Chem. Soc. 1962, 84, 2017–2018.
Gross, E. Cyanogen bromide reaction with peptides. Methods Enzymol. 1967, 11, 238–255.
Tang, J. R.; Hartley, B. S. A diagonal electrophoretic method for selective purification of methionine peptides. Biochem. J. 1967, 102, 593–599.
Gairi, M.; Lloyd-Williams, P.; Albericio, F.; Giralt, E. Severe side-reaction in the acidolytic cleavage of a C-terminal Met-containing peptide from the solid support: Formation of the homoserine lactone peptide. Tetrahedron Lett. 1994, 35, 175–178.
Taylor, K. L.; Pohl, J.; Kinkade, J. M. Unique autolytic cleavage of human myeloperoxidase: Implications for the involvement of active site Met409. J. Biol. Chem. 1992, 267, 25282–25288.
Kyte, J.; Degen, J.; Harkins, R. N. Purification of peptides that contain methionine residues. Methods Enzymol. 1983, 91, 367–377.
Weinberger, S. R.; Viner, R. I.; Ho, P. Tagless extraction-retentate chromatography: A new global protein digestion strategy for monitoring differential protein expression. Electrophoresis 2002, 23, 3182–3192.
Chassaing, G.; Lavielle, S.; Marquet, A. Transformation of methionine into S-tert-butylhomocysteine. Application to a methionine-containing peptide: Substance P. J. Org. Chem. 1983, 48, 1757–1760.
O’Hair, R. A. J.; Reid, G. E. Neighboring group versus cis-elimination mechanisms for side chain loss from protonated methionine, methionine sulfoxide, and their peptides gas-phase ion chemistry of biomolecules. Eur. Mass Spectrom 1999, 5, 325–334.
Reid, G. E.; Simpson, R. J.; O’Hair, R. A. J. Leaving group and gas phase neighboring group effects in the side chain losses from protonated serine and its derivatives. J. Am. Soc. Mass Spectrom. 2000, 11, 1047–1060.
Foti, S.; Saletti, R.; Marletta, D. Partial methanolysis and fast atom bombardment mass spectrometry of peptides containing sulfurated amino acids. Org. Mass Spectrom. 1991, 26, 903–907.
Lapko, V. N.; Smith, D. L.; Smith, J. B. Identification of an artifact in the mass spectrometry of proteins derivatized with iodoacetamide. J. Mass Spectrom. 2000, 35, 572–575.
Stirling, C. J. M. Sulfonium salts in organic chemistry of sulfur; Oae, S. Ed.; Plenum Press: New York, 1977; Chap. IX, pp 473–525.
Capozzi, G.; Modena, G. Reactive sulfonium salts. Stud. Org. Chem. 1985, 19, 246–298.
March, J. Advanced Organic Chemistry, 4th ed.; Wiley: New York, 1992; p 343.
Halvorsen, S. The reactivity of 2-bromo-1-phenylethanone (phenacyl bromide) toward nucleophilic species. J. Chem. Soc. Chem. Commun. 1978, 327–328.
Yoh, S. D.; Lee, O. S. A multiple Hammett study of the nucleophilic substitution reaction of a-carbonyl derivatives. Tetrahedron Lett. 1988, 29, 4431–4434.
Scheitjter, A.; Aviram, I. Reversal of the alkylation of the methionine residues of cytochrome c. FEBS Lett. 1972, 21, 293–296.
Toennies, G.; Kolb, J. J. Methionine Studies: VIII. Regeneration of Sulfides from Sulfonium Derivatives. J. Am. Chem. Soc. 1945, 67, 1141–1144.
Naider, F.; Bohak, Z. Regeneration of methionyl residues from their sulfonium salts in peptides and proteins. Biochemistry 1972, 11, 3208–3211.
Cantoni, G. L. Onium compounds and their biological significance. Comp. Biochem. 1960, 1, 181–241.
Trost, B. M.; Melvin, L. S. Organic Chemistry, Vol. 31: Sulfur Ylides: Emerging Synthetic Intermediates. Academic Press: New York, 1975.
O’Hair, R. A. J.; Freitas, M. A.; Gronert, S.; Schmidt, J. A. R.; Williams, T. D. Concerning the regioselectivity of gas-phase reactions of glycine with electrophiles: The cases of the dimethylchlorinium ion and the methoxymethyl cation. J. Org. Chem. 1995, 60, 1990–1998.
Mudd, W. A.; Klee, P. D. Enthalpy changes accompanying the transfer of a methyl group from S-adenosylmethionine and other sulfonium compounds to homocysteine. Biochemistry 1966, 5, 1653–1660.
Deakyne, C. A.; Knuth, D. M.; Meot-Ner, M.; Breneman, C. M.; Liebman, J. F. Experimental and theoretical study of the energetics of trialkylsulfonium ions. J. Mol. Struct. 1999, 485/486, 33–41.
Markham, G. D.; Bock, C. W. Structural and thermodynamic properties of sulfonium ions: An ab initio molecular orbital study. J. Phys. Chem. 1993, 97, 5562–5569.
Katritzky, A. R.; Shipkova, P. A.; Watson, C. H.; Eyler, J. R.; Kevill, D. N. Gas-phase investigations of sulfonium salts by electrospray FT-ICR/MS. Int. J. Mass Spectrom 1997, 165/166, 577–583.
Buckley, N.; Maltby, D.; Burlingame, A. L.; Oppenheimer, N. J. Reactions of charged substrates. 4: The gas-phase dissociation of (4-substituted benzyl)dimethylsulfoniums and -pyridiniums. J. Org. Chem. 1996, 61, 2753–2762.
Mestdagh, H.; Morin, N.; Rolando, C. Sulfonium ions: Collision-activated dissociation using fast atom bombardment ionization and a triple quadrupole mass spectromter. Org. Mass Spectrom. 1986, 21, 321–327.
Mestdagh, H.; Morin, N.; Rolando, C. Comparison of low energy CAD spectra of sulfonium, ammonium, and phosphonium cations: Propene elimination from allyl-substituted oniums. Org. Mass Spectrom. 1988, 23, 246–251.
Gerber, S. A.; Rush, J.; Stemman, O.; Kirschner, M. W.; Gygi, S. P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. U.S.A. 2003, 100, 6940–6945.
Gevaert, K.; Van Damme, J.; Goethals, M.; Thomas, G. R.; Hoorelbeke, B.; Demol, H.; Martens, L.; Puype, M.; Staes, A.; Vandekerckhove, J. Chromatographic isolation of methionine-containing peptides for gel-free proteome analysis: Identification of more than 800. Escherichia coli proteins. Mol. Cell. Proteom. 2002, 1, 896–903.
Ren, D.; Julka, S.; Inerowicz, H. D.; Regnier, F. E. Enrichment of cysteine-containing peptides from tryptic digests using a quaternary amine tag. Anal. Chem. 2004, 76, 4522–4530.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online May 31, 2005
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Reid, G.E., Roberts, K.D., Simpson, R.J. et al. Selective identification and quantitative analysis of methionine containing peptides by charge derivatization and tandem mass spectrometry. J Am Soc Mass Spectrom 16, 1131–1150 (2005). https://doi.org/10.1016/j.jasms.2005.03.015
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2005.03.015