Abstract
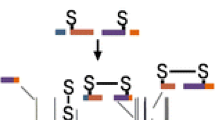
An efficient method for data processing and interpretation is needed to support and extend disulfide mass-mapping methodology based on partial reduction and cyanylation-induced cleavage to proteins containing more than four cystines. Here, the concept of “negative signature mass” is introduced as the novel feature of an algorithm designed to identify the disulfide structure of a cystinyl protein given an input of mass spectral data and an amino acid sequence. The “negative signature mass” process is different from the conventional approach in that it does not directly rule-in disulfide linkages, but rather eliminates linkages from a list of all possible theoretical linkages, with the goal of ruling out enough linkages so that only one disulfide structure can be constructed. The operating principles and the effectiveness of the algorithm are described in the context of analyzing ribonuclease A, a 124-residue protein containing eight cysteines in the form of four cystines (disulfides).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Smith, D. L.; Zhou, Z. R. Strategies for Locating Disulfide Bonds in Proteins. Methods Enzymol. 1990, 193, 374–389.
Zhou, Z. R.; Smith, D. L. Location of Disulfide Bonds in Antithrombin III. Biomed. Environ. Mass Spectrom. 1990, 19, 782–786.
Merewether, L. A.; Le, J.; Jones, M. D.; Lee, R.; Shimamoto, G.; Lu, H. S. Development of Disulfide Peptide Mapping and Determination of Disulfide Structure of Recombinant Human Osteoprotegerin Chimera Produced in Escherichia coli. Arch. Biochem. Biophys. 2000, 375, 101–110.
Wu, J.; Watson, J. T. A Novel Methodology for Assignment of Disulfide Bond Pairings in Proteins. Protein Sci. 1997, 6, 391–398.
Wu, J.; Watson, J. T. Assignment of Disulfide Bonds in Proteins by Chemical Cleavage and Peptide Mapping by Mass Spectrometry. Methods Mol. Biol. 2002, 194, 1–22.
Qi, J.; Wu, J.; Somkuti, G. A.; Watson, J. T. Determination of the Disulfide Structure of Sillucin, a Highly Knotted, Cysteine-Rich Peptide, by Cyanylation/Cleavage Mass Mapping. Biochemistry 2001, 40, 4531–4538.
Jacobson, G. R.; Schaffer, M. H.; Stark, G. R.; Vanaman, T. C. Specific Chemical Cleavage in High Yield at the Amino Peptide Bonds of Cysteine and Cystine Residues. J. Biol. Chem. 1973, 248, 6583–6591.
Gray, W. R. Disulfide Structures of Highly Bridged Peptides: A New Strategy for Analysis. Protein Sci. 1993, 2, 1732–1748.
Harary, F. Graph Theory. Addison-Wesley: New York, 1969 pp 8–23, 96.
Wu, J.; Gage, D. A.; Watson, J. T. A Strategy to Locate Cysteine Residues in Proteins by Specific Chemical Cleavage Followed by Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry. Anal. Biochem. 1996, 235, 161–174.
Gabow, H. N. An Efficient Implementation of Edmond’s Algorithm for Maximum Matching on Graphs. J. Assoc. Comput. Machin. 1976, 23, 221–234.
Wu, J.; Watson, J. T. Optimization of the Cleavage Reaction for Cyanylated Cysteinyl Proteins for Efficient and Simplified Mass Mapping. Anal. Biochem. 1998, 258, 268–276.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online July 31, 2003
Rights and permissions
About this article
Cite this article
Qi, J., Wu, W., Borges, C.R. et al. Automated data interpretation based on the concept of “negative signature mass” for mass-mapping disulfide structures of cystinyl proteins. J Am Soc Mass Spectrom 14, 1032–1038 (2003). https://doi.org/10.1016/S1044-0305(03)00145-4
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1016/S1044-0305(03)00145-4