Abstract
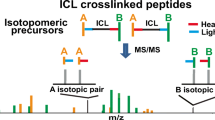
In a previous report (Young et al., Proc. Natl. Acad. Sci. U.S.A. 2000, 97, 5802–5806), we provided a proof-of-principle for fold recognition of proteins using a homobifunctional amine-specific chemical crosslinking reagent in combination with mass spectrometry analysis and homology modeling. In this current work, we propose a systematic nomenclature to describe the types of peptides that are generated after proteolysis of crosslinked proteins, their fragmentation by tandem mass spectrometry, and an automated algorithm for MS/MS spectral assignment called “MS2Assign.” Several examples are provided from crosslinked peptides and proteins including HIV-integrase, cytochrome c, ribonuclease A, myoglobin, cytidine 5-monophosphate N-acetylneuraminic acid synthetase, and the peptide thymopentin. Tandem mass spectra were obtained from various crosslinked peptides using post source decay MALDI-TOF and collision induced dissociation on a quadrupole-TOF instrument, along with their automated interpretation using MS2Assign. A variety of possible outcomes are described and categorized according to the number of modified lysines and/or peptide chains involved, as well as the presence of singly modified (dead-end) lysine residues. In addition, the proteolysis and chromatographic conditions necessary for optimized crosslinked peptide recovery are presented.
Article PDF
Similar content being viewed by others
References
Young, M. M.; Tang, N.; Hempel, J. C.; Oshiro, C. M.; Taylor, E. W.; Kuntz, I. D.; Gibson, B. W.; Dollinger, G. High Throughput Protein Fold Identification by Using Experimental Constraints Derived from Intramolecular Cross-Links and Mass Spectrometry. Proc. Natl. Acad. Sci. U.S.A. 2000, 97, 5802–5806.
Muller, D. R.; Schindler, P.; Towbin, H.; Wirth, U.; Voshol, H.; Hoving, S.; Steinmetz, M. O. Isotope-Tagged Cross-Linking Reagents. A New Tool in Mass Spectrometric Protein Interaction Analysis. Anal. Chem. 2001, 73, 1927–1934.
Rappsilber, J.; Siniossoglou, S.; Hurt, E. C.; Mann, M. A Generic Strategy to Analyze the Spatial Organization of Multi-Protein Complexes by Cross-Linking and Mass Spectrometry. Anal. Chem. 2000, 72, 267–275.
Shih, C. L.; Chen, M. J.; Linse, K.; Wang, K. Molecular Contacts between Nebulin and Actin: Cross-Linking of Nebulin Modules to the N-Terminus of Actin. Biochemistry 1997, 36, 1814–1825.
Sinz, A.; Wang, K. Mapping Protein Interfaces with a Fluorogenic Cross-Linker and Mass Spectrometry: Application to Nebulin-Calmodulin Complexes. Biochemistry 2001, 40, 7903–7913.
Guo, X.; Schilling, B.; Young, M. M.; Medzihradszky, M.; Kuntz, I. D.; Guy, R. K.; Gibson, B. W. Using Homobifunctional Crosslinking Reagents with Normal and N-15 Labeled Proteins for the Deteremination of Protein Tertiary Structure and Protein-Protein Interactions. Proceedings of the 50th ASMS Conference on Mass Spectrometry and Allied Topics; Orlando, FL, June, 2002
Taverner, T.; Hall, N. E.; O’Hair, R. A.; Simpson, R. J. Characterization of an Antagonist Interleukin-6 Dimer by Stable Isotope Labeling, Cross-Linking, and Mass Spectrometry. J. Biol. Chem. 2002, 277, 46487–46492.
Pearson, K. M.; Pannell, L. K.; Fales, H. M. Intramolecular Cross-Linking Experiments on Cytochrome C and Ribonuclease A Using an Isotope Multiplet Method. Rapid Commun. Mass Spectrom. 2002, 16, 149–159.
Chen, T.; Jaffe, J. D.; Church, G. M. Algorithms for Identifying Protein Cross-Links Via Tandem Mass Spectrometry. J. Comput. Biol. 2001, 8, 571–583.
Back, J. W.; Hartog, A. F.; Dekker, H. L.; Muijsers, A. O.; de Koning, L. J.; de Jong, L. A New Crosslinker for Mass Spectrometric Analysis of the Quaternary Structure of Protein Complexes. J. Am. Soc. Mass Spectrom. 2001, 12, 222–227.
Back, J. W.; Sanz, M. A.; De Jong, L.; De Koning, L. J.; Nijtmans, L. G.; De Koster, C. G.; Grivell, L. A.; Van Der Spek, H.; Muijsers, A. O. A Structure for the Yeast Prohibitin Complex: Structure Prediction and Evidence from Chemical Crosslinking and Mass Spectrometry. Protein Sci. 2002, 11, 2471–2478.
Trester-Zedlitz, M.; Kamada, K.; Burley, S. K.; Fenyo, D.; Chait, B. T.; Muir, T. W. A Modular Cross-Linking Approach for Exploring Protein Interactions. J. Am. Chem. Soc. 2003, 125, 2516–2525.
Roepstorff, P.; Fohlman, J. Proposal for a Common Nomenclature for Sequence Ions in Mass Spectra of Peptides. Biomed. Mass Spectrom. 1984, 11, 601.
Biemann, K. Nomenclature for Peptide Fragment Ions (Positive Ions). Appendix 5. Methods Enzymol. 1990, 193, 886–887.
Tullius, M. V.; Munson, R. S.; Wang, J.; Gibson, B. W. Purification, Cloning, and Expression of a Cytidine 5′-Monophosphate N-Acetylneuraminic Acid Synthetase from Haemophilus ducreyi. J. Biol. Chem. 1996, 271, 15373–15380.
Medzihradszky, K. F. Methods Enzymol., in press
Vestal, M. L.; Juhasz, P.; Martin, S. A. Delayed Extraction Matrix-Assisted Laser Desorption Time-of-Flight Mass Spectrometry. Rapid Commun. Mass Spectrom. 1995, 9, 1044–1050.
Spengler, B.; Kirsch, D.; Kaufmann, R.; Jaeger, E. Peptide Sequencing by Matrix-Assisted Laser-Desorption Mass Spectrometry. Rapid Commun. Mass Spectrom. 1992, 6, 105–108.
Ngoka, L. C. M.; Gross, M. L. A Nomenclature System for Labeling Cyclic Peptide Fragments. J. Am. Soc. Mass Spectrom. 1999, 10, 360–363.
Yalcin, T.; Csizmadia, I. G.; Peterson, M. R.; Harrison, A. G. The Structure and Fragmentation of b n (n>3) Ions in Peptide Spectra. J. Am. Mass Spectrom. 1996, 7, 233–242.
Harrison, A. G.; Csizmadia, I. G.; Tang, T. H. Structure and Fragmentation of B2 Ions in Peptide Mass Spectra. J. Am. Soc. Mass Spectrom. 2000, 11, 427–436.
Hines, N. M.; Falick, A. M.; Burlingame, A. L.; Gibson, B. W. Pattern-Based Algorithm for Peptide Sequencing from Tandem High Energy Collision-Induced Dissociation Mass Spectra. J. Am. Soc. Mass Spectrom. 1992, 3, 326–336.
Falick, A. M.; Hines, W. M.; Medzihradszky, K. F.; Baldwin, M. A.; Gibson, B. W. Low-Mass Ions Produced from Peptides by High-Energy Collision-Induced Dissociation in Tandem Mass Spectrometry. J. Am. Soc. Mass Spectrom. 1993, 4, 882–893.
Kim, J. Y.; Kim, K. W.; Kwon, H. J.; Lee, D. W.; Yoo, J. S. Probing Lysine Acetylation with a Modification-Specific Marker Ion Using High-Performance Liquid Chromatography/Electrospray-Mass Spectrometry with Collision-Induced Dissociation. Anal. Chem. 2002, 74, 5443–5449.
Green, N. S.; Reisler, E.; Houk, K. N. Quantitative Evaluation of the Length of Homobifunctional Protein Cross-Linking Reagents Used as Molecular Rulers. Protein Sci. 2001, 10, 1293–1304.
Eng, J. K.; McCormack, A. L.; Yates, J. R., III. An Approach to Correlate Tandem Mass Spectral Data of Peptides with Amino Acid Sequences in a Protein Database. J. Am. Soc. Mass Spectrom. 1994, 5, 976–989.
Field, H. I.; Fenyo, D.; Beavis, R. C. Radars, a Bioinformatics Solution that Automates Proteome Mass Spectral Analysis, Optimizes Protein Identification, and Archives Data in a Relational Database. Proteomics 2002, 2, 36–47.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online June 25, 2003
Rights and permissions
About this article
Cite this article
Schilling, B., Row, R.H., Gibson, B.W. et al. MS2Assign, automated assignment and nomenclature of tandem mass spectra of chemically crosslinked peptides. J Am Soc Mass Spectrom 14, 834–850 (2003). https://doi.org/10.1016/S1044-0305(03)00327-1
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1016/S1044-0305(03)00327-1