Abstract
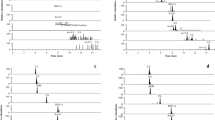
Short-chain fatty acids (SCFAs) are metabolites derived from gut microbiota and implicated in host homeostasis. Hence, the profiling SCFAs from biological samples plays an important role in revealing the interaction between gut microbiota and pathogens. Previous studies, liquid chromatography–tandem mass spectrometry (LC–MS/MS) combined with various derivatization strategies have been performed to obtain the SCFA profiles from biological samples. However, it is poor evidence to compare these derivatization regents and conditions. Thus, we present the evaluation of three major derivatization reagents, namely 3-nitrophenylhydrazine (3-NPH), O-benzylhydroxylamine (O-BHA), and 2-picolylamine (2-PA), for the analysis of eight SCFAs classified as C2–C5 isomers using LC–MS/MS. First, in a reversed-phase LC separation, 3-NPH showed good retention capacity. Although O-BHA derivatization showed higher sensitivity and good retention capacity than 2-PA, only 2-PA derivatization could successfully separate eight SCFAs. The matrix effects in human serum ranged 77.1–99.0% (RSD ≤ 3.4%, n = 6) for 3-NPH derivatives, 91.0–94.6% (RSD ≤ 5.4%, n = 6) for O-BHA derivatives, 81.6–99.5% (RSD ≤ 8.0%, n = 6) for 2-PA derivatives. These compared results showed each characteristic of 3-NPH, O-BHA, and 2-PA for SCFA derivatization based on LC–MS/MS approaches.
Graphical abstract





Similar content being viewed by others
Availability of data and materials
The datasets used and analyzed during the current study are available.
References
J.H. Cummings, E.W. Pomare, W.J. Branch, C.P. Naylor, G.T. Macfarlane, Gut 28, 1221–1227 (1987)
J. Hu, S. Lin, B. Zheng, P.C.K. Cheung, Crit. Rev. Food Sci. Nutr. 58, 1243–1249 (2018)
T. Ikeda, A. Nishida, M. Yamano, I. Kimura, Pharmacol. Ther. 239, 108273 (2022)
Y. Furusawa, Y. Obata, S. Fukuda, T.A. Endo, G. Nakato, D. Takahashi, Y. Nakanishi, C. Uetake, K. Kato, T. Kato, M. Takahashi, N.N. Fukuda, S. Murakami, E. Miyauchi, S. Hino, K. Atarashi, S. Onawa, Y. Fujimura, T. Lockett, J.M. Clarke, D.L. Topping, M. Tomita, S. Hori, O. Ohara, T. Morita, H. Koseki, J. Kikuchi, K. Honda, K. Hase, H. Ohno, Nature 504, 446–450 (2013)
E.E. Canfora, R.C.R. Meex, K. Venem, E.E. Blaak, Nat. Rev. Endocrinol. 15, 261–273 (2019)
S. Zhang, H. Wang, M.J. Zhu, Talanta 196, 249–254 (2019)
S. Scortichini, M.C. Boarelli, S. Silvi, D. Fiorini, J. Chromatogr. B 1143, 121972 (2020)
S. Saha, P. Day-Walsh, E. Shehata, P.A. Kroon, Molecules 26, 6444 (2021)
J. Han, K. Lin, C. Sequeira, C.H. Borchers, Anal. Chim. Acta 854, 86–94 (2015)
Y.Y. Jin, Z.Q. Shi, W.Q. Chang, L.X. Guo, J.L. Zhou, J.Q. Liu, L.F. Liu, G.Z. Xin, J. Pharm. Biomed. Anal. 161, 336–343 (2018)
G. Liebisch, J. Ecker, S. Roth, S. Schweizer, V. Öttl, H.F. Schött, H. Yoon, D. Haller, E. Holler, R. Burkhardt, S. Matysik, Biomolecules 9, 121 (2019)
H.Y. Liao, C.Y. Wang, C.H. Lee, H.L. Kao, W.K. Wu, C.H. Kuo, J. Proteome Res. 20, 3508–3518 (2021)
M. Dei Cas, R. Paroni, A. Saccardo, E. Casagni, S. Arnoldi, V. Gambaro, M. Saresella, C. Mario, F. La Rosa, I. Marventano, F. Piancone, G. Roda, J. Chromatogr. B Chromatogr. B 1154, 121982 (2020)
L. Chen, X. Sun, A.S. Khalsa, M.T. Bailey, K. Kelleher, C. Spees, J. Zhu, J. Pharm. Biomed. Anal. 200, 114066 (2021)
X. Wang, H. Ruan, Z. Zong, F. Mao, Y. Wang, Y. Jiao, L. Xu, T. Yang, W. Li, X. Liu, J. Chromatogr. B 1180, 122895 (2021)
M.A. Valdivia-Garcia, K.E. Chappell, S. Camuzeaux, L. Olmo-García, V.H. van der Sluis, S.T. Radhakrishnan, H. Stephens, S. Bouri, L.M. de Campos Braz, H.T. Williams, M.R. Lewis, G. Frost, J.V. Li, J. Pharm. Biomed. Anal. 221, 115060 (2022)
M. Zeng, H. Cao, J. Chromatogr. B 1083, 137–145 (2018)
R. Nagatomo, Y. Okada, M. Ichimura, K. Tsuneyama, K. Inoue, Anal. Sci. 34, 1031–1036 (2018)
R. Nagatomo, H. Kaneko, S. Kamatsuki, M. Ichimura-Shimizu, N. Ishimaru, K. Tsuneyama, K. Inoue, J. Chromatogr. B 1210, 123432 (2022)
Y. Kanemitsu, E. Mishima, M. Maekawa, Y. Matsumoto, D. Saigusa, H. Yamaguchi, J. Ogura, H. Tsukamoto, Y. Tomioka, T. Abe, N. Mano, Sci. Rep. 9, 19075 (2019)
J.C. Chan, D.Y. Kioh, G.C. Yap, B.W. Lee, E.C. Chan, J. Pharm. Biomed. Anal. 138, 43–53 (2017)
R. Jiang, Y. Jiao, P. Zhang, Y. Liu, X. Wang, Y. Huang, Z. Zhang, F. Xu, Anal. Chem. 89, 12223–12230 (2017)
H.E. Song, H.Y. Lee, S.J. Kim, S.H. Back, H.J. Yoo, Metabolites 9, 173 (2019)
J. Zheng, S.J. Zheng, W.J. Cai, L. Yu, B.F. Yuan, Y.Q. Feng, Anal. Chim. Acta 1070, 51–59 (2019)
Z. Chen, Z. Gao, Y. Wu, R. Shrestha, H. Imai, N. Uemura, K.I. Hirano, H. Chiba, S.P. Hui, J. Chromatogr. B 1126–1127, 121771 (2019)
S.R. Ma, Q. Tong, Z.X. Zhao, L. Cong, J.B. Yu, J. Fu, P. Han, L.B. Pan, R. Gu, R. Peng, Z.W. Zhang, Y. Wang, J.D. Jiang, Anal. Bioanal. Chem. 411, 3191–3207 (2019)
H. Fu, Q.L. Zhang, X.W. Huang, Z.H. Ma, X.L. Zheng, S.L. Li, H.N. Duan, X.C. Sun, F.F. Lin, L.J. Zhao, G.S. Teng, J. Liu, Rapid Commun. Mass Spectrom. 34, e8730 (2020)
J. Wei, L. Xiang, X. Li, Y. Song, C. Yang, F. Ji, A.C.K. Chung, K. Li, Z. Lin, Z. Cai, Anal. Chim. Acta 1100, 66–74 (2020)
M.J. McKay, M. Castaneda, S. Catania, K.A. Charles, E. Shanahan, S.J. Clarke, A. Engel, P. Varelis, M.P. Molloy, J. Chromatogr. B 1217, 123618 (2023)
C. Vagaggini, A. Brai, D. Bonente, J. Lombardi, F. Poggialini, C. Pasqualini, V. Barone, C. Nicoletti, E. Bertelli, E. Dreassi, J. Pharm. Biomed. Anal. 235, 115599 (2023)
Acknowledgements
This work was supported by JSPS KAKENHI Grant No. 22K06557.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nagatomo, R., Ichikawa, A., Kaneko, H. et al. Comparison of 3-nitrophenylhydrazine, O-benzyl hydroxylamine, and 2-picolylamine derivatizations for analysis of short-chain fatty acids through liquid chromatography coupled with tandem mass spectrometry. ANAL. SCI. 40, 843–851 (2024). https://doi.org/10.1007/s44211-023-00474-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44211-023-00474-7