Abstract
Arsenic (As) is a cancerogenic metalloid ubiquitously distributed in the environment, which can be easily accumulated in food crops like rice. Jasmonic acid (JA) and its derivatives play critical roles in plant growth and stress response. However, the role of endogenous JA in As accumulation and detoxification is still poorly understood. In this study, we found that JA biosynthesis enzymes Allene Oxide Synthases, OsAOS1 and OsAOS2, regulate As accumulation and As tolerance in rice. Evolutionary bioinformatic analysis indicated that AOS1 and AOS2 have evolved from streptophyte algae (e.g. the basal lineage Klebsormidium flaccidum) – sister clade of land plants. Compared to other two AOSs, OsAOS1 and OsAOS2 were highly expressed in all examined rice tissues and their transcripts were highly induced by As in root and shoot. Loss-of-function of OsAOS1 (osaos1–1) showed elevated As concentration in grains, which was likely attributed to the increased As translocation from root to shoot when the plants were subjected to arsenate [As(V)] but not arsenite [As (III)]. However, the mutation of OsAOS2 (osaos2–1) showed no such effect. Moreover, osaos1–1 and osaos2–1 increased the sensitivity of rice plants to both As(V) and As(III). Disrupted expression of genes involved in As accumulation and detoxification, such as OsPT4, OsNIP3;2, and OsOASTL-A1, was observed in both osaos1–1 and osaos2–1 mutant lines. In addition, a As(V)-induced significant decrease in Reactive Oxygen Species (ROS) production was observed in the root of osaos1–1 but not in osaos2–1. Taken together, our results indicate OsAOS1 modulates both As allocation and detoxification, which could be partially attributed to the altered gene expression profiling and ROS homeostasis in rice while OsAOS2 is important for As tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arsenic (As) is a natural component of our planet’s crust, which is widely distributed in the environment. The proportion of soil and water contaminated with As was progressively increasing due to anthropogenic activities (Deng et al. 2021; Khanna et al. 2022). For example, out of 112 soil samples in China, 24 (21.4%) had mean soil As concentrations above the World Health Organization (WHO) limit of 30 mg/kg; and a global prediction model shows arsenic exceeding 10 mg/L in most groundwater in Asia, the America and Africa (Antoniadis et al. 2019; Podgorski and Berg 2020). Excessive As is toxic to almost all animals and most plant species. For humans, long-term As intake can cause skin cancer and cancerous changes in internal organs such as the kidney, liver, and bladder (Clemens and Ma 2016). For plants, As toxicity has been observed to reduce photosynthetic pigments in leaves, inhibit plant height, shoot biomass, and reduce tiller number as well as substantial yield losses when five widely cultivated rice accessions were grown in the paddy soil with additional As (Azizur Rahman et al. 2007). The risk resulted from the increasing As concentrations in the soil-plant-human ecosystem is one of the major concerns for global food safety (Guan et al. 2021).
Rice (Oryza sativa) is a staple food for more than half of the world population. Arsenite [As(III)] and arsenate [As(V)] are two major forms of inorganic As in soil, which can be absorbed by rice roots through silicic acid transporters and phosphate transporters, respectively (Ma et al. 2008; Cao et al. 2017; Zhao et al. 2022). As is more easily accumulated in rice grains compared to the other cereals such as wheat (Triticum aestivum) and barley (Hordeum vulgare) partially due to the anoxic growth condition and the highly efficient pathway for silicon absorption (Su et al. 2010; Zhao et al. 2022). A large-scale investigation of a rice core collection consisting of 1763 germplasms revealed that As content in the grains varied from 0.217 to 2.610 mg/kg, with an average of 0.945 ± 0.312 mg/kg (Pinson et al. 2015). In addition, 5 ~ 22% of 471 high-yielding rice varieties also showed high grain As exceeding the threshold even when the plants were grown in the paddy soil with the As content under the national limit (Duan et al. 2017). It was estimated that about 60% of total inorganic As intake in the Chinese population was from the consumption of rice, and the ratio was much higher than the populations in Italy and the United States (Meharg et al. 2009). This indicated that rice is one of the most important sources of inorganic As intake through the food chain for the populations using rice as the staple food (Zhao et al. 2022). Therefore, developing rice lines with reduced grain As is one of the most plausible strategies for reducing human As intake, especially for billions of people in Asia consuming rice as their staple food.
Effective strategies for reducing grain As accumulation through biotechnology had been developed during last decades based on the discovering of the molecular mechanisms on As accumulation in rice (Deng et al. 2019; Deng et al. 2021; Yamaji and Ma 2021; Khanna et al. 2022; Zhao et al. 2022). For instance, overexpression of Nodulin 26-like Intrinsic Proteins (OsNIP1;1 and OsNIP3;3) in rice decreased As accumulation in the shoot and grain through disrupting the radial transport of As(III) in roots (Sun et al. 2018). Significantly reduced rice grain As was also succeeded through increasing the expression of C-type ATP-binding Cassette Transporter 1 (OsABCC1) in the root cortical cells together with the enhanced levels of enzymes for phytochelatins (PCs) synthesis (Deng et al. 2018), overexpression of PC Synthase 1 (OsPCS1) (Hayashi et al. 2017), seed-specific silencing of endogenous Multidrug and Toxic compound Extrusion 2 (OsMATE2) (Das et al. 2018), or ectopic expression of Arsenate Reductase PvACR3;1 from the As-hyperaccumulator, Pteris vittata (Chen et al. 2019) and ScACR3 from yeast (Saccharomyces cerevisiae) (Duan et al. 2012). Therefore, enhancing As vacuolar sequestration and chelation, increasing As efflux activity and decreasing As uptake are feasible to reduce As accumulation in rice grains for less human As intake through food chain.
Application of special materials such as fungus, fertilizers, and nanoparticles was also effective for conquering As toxicity and decreasing As accumulation in rice. For example, synergistic application of zinc oxide nanoparticles and salicylic acid in rice through modulation of the cellular redox status and antioxidant defense (Faizan et al. 2021). The colonization of endophytic fungus Serendipita indica and phosphorus synergistically recuperate arsenic induced stress in rice (Abbasi et al. 2021; Sehar et al. 2022), mainly through regulating secondary metabolism related enzymatic activity and root metabolic patterns (Sehar et al. 2023a, 2023b).
As a group of critical phytohormone functions in plant growth, development and resistance to environmental stresses (Zhang et al. 2023c), jasmonates [e.g. jasmonic acid (JA), jasmonoyl-l-isoleucine (JA-Ile), and methyl jasmonate (MeJA)] have been implicated in As accumulation and detoxification in plants (Verma et al. 2020; Chen et al. 2021; Li et al. 2022a, 2022b). For example, application of MeJA or JA alleviates As(III) (Mousavi et al. 2020; Verma et al. 2020; Li et al. 2022a, 2022b) and As(V) (Wang et al. 2018; Zhang et al. 2023b) toxicity and accumulation in rice, respectively. Similarly, the effect of MeJA on alleviating As(III) stress in oilseed (Brassica napus) was also observed (Farooq et al. 2018). The molecular mechanisms on JA pathway have been extensively studied (Wasternack and Feussner 2018; Hu et al. 2023). Jasmonates are synthesized from α-linolenic acid through the octadecanoid pathway and cytochrome P450 of the CYP74 family enzymes allene oxide synthases (AOSs) are the key enzymes for JA biosynthesis (Wasternack and Feussner 2018). Molecular evolutionary analysis of JA pathway showed that the genes associated with the JA pathway are found in most tested land plants (Chen et al. 2021). There are 4 members of AOS family in the rice genome, which were designated as OsAOS1–4 (Haga and Iino 2004). Purified recombinant OsAOS2 protein expressed in Escherichia coli successfully converted 13-hydroperoxylinolenic acid to allene oxid, an important step in the biosynthesis of JA (Ha et al. 2022). Due to the critical roles of JA in resistance to pathogen infection and insect herbivory (Mei et al. 2006; Ye et al. 2013), OsAOSs was widely known as in biotic stresses (Li et al. 2022b). However, the link between the core molecular components in JA biosynthesis and signaling and As tolerance are not well understood in rice.
Therefore, we hypothesized that OsAOSs are key determinates for As accumulation and detoxification in rice plants. We compared the expression pattern of OsAOSs among rice tissues and their expression in response to As(III) and As(V), and found that OsAOS1 and OsAOS2 showed the highest expression level in the examined tissues, and they were dramatically induced by As. We traced the evolutionary origin of AOS1 and AOS2 in green plants, and the mutant lines of OsAOS1 and OsAOS2 were employed for further evaluation of As accumulation and tolerance. Our results suggested that a single amino acid deletion of OsAOS1 increases As sensitivity of the osaos1–1 plants and higher As accumulation in rice grain.
Results
Allene oxide synthases AOS1 and AOS2 are evolutionarily conserved in green plants
We found that allene oxide synthases required for the biosynthesis of JA are widely presented in land plants, and the homologues of AOSs can be firmly traced to streptophyte algal species (Fig. 1 and S1) such as Klebsormidium flaccidum, and possibly from Chlorophyte algae such as Chlamydomonas reinhardtii and Volvox carteri (Chen et al. 2021). Using the sequence of OsAOS1 as the search query, lots of orthologs were obtained from the OneKP database (One Thousand Plant Transcriptomes Initiative 2019) and 188 orthologs were chosen for the further analysis. The phylogenetic tree of AOS1s from examined plant species showed that AOS1 orthologs could be divided into different clades according to the plant lineages, and the putative orthologues of OsAOS1 from Chromista species were independent of the other subgroups (Fig. 1A). Similar results were also observed when using OsAOS2 as a reference sequence (Fig. S1). It was revealed that PpAOS1 and PpAOS2 from the moss Physcomitrella patens (Stumpe et al. 2006), MpAOS1 and MpAOS2 from the liverwort Marchantia polymorpha, and KfAOS from K. flaccidum (Koeduka et al. 2015) exhibit high amino acid sequence similarity compared to those of angiosperms including rice (Fig. 1 and S1).
Evolutionary analysis of allene oxide synthase 1 (AOS1) homologues in land plants and algal species. A The phylogenetic tree of AOS1 homologues identified from the representative species from various linkages. The orthologues from Klebsormidium flaccidum, Oryza sativa, Arabidopsis thaliana, Marchantia polymorpha, Physcomitrella patens were highlighted with red asterisks. B Conserved motifs through multiple protein sequences alignment of AOS1 homologues from selected species. C Sequence alignment with the deduced protein fragments containing the mutation site of t of osaos1–1
Conserved motifs including Helix-I region [GXXX(F/L)], Helix-K region motif (also named as EXLR motif), and the Heme-binding domain (PXVXNKQCPG) of CYP74 members (Chehab et al. 2007; Zhou et al. 2019) were identified in OsAOS1 (Fig. 1B). Further sequence alignment analyses indicated high evolutionary conservation of these domains in the representative species of the major green plant lineages, while the similarity of these domains with those in the chlorophyate alga C. reinhardtii was much lower than those from land plants (Fig. 1B). Therefore, AOS1 and AOS2 are largely conserved among green plants in terms of molecular evolutionary aspects.
OsAOS1 and OsAOS2 are highly expressed and induced by As in rice tissues
There are 4 members of AOS family in the rice genome, which were designated as OsAOS1–4 (Haga and Iino 2004). According to the gene expression profiles in various organs of rice plants (cv. Nipponbare) grown in paddy soil until filling stage, OsAOS1 (Os03g0767000, LOC_Os03g55800) displayed constitutively high expression pattern in all examined tissues. The highest expression level of OsAOS1 was observed in the leaf sheath II, node II, node I, internode II, and root, while moderate transcripts were also detected in the leaf blade and spikelet. Low expression of OsAOS2 (Os03g0225900, LOC_Os03g12500) was observed in the nodes, internode, rachis and spikelet. OsAOS3 (Os02g0218700, LOC_Os02g12680) and OsAOS4 (Os02g0218800, LOC_Os02g12690) showed low expression levels except in roots (Fig. 2A).
The response of OsAOSs to As(V) and As(III) was further compared by using published transcriptomic datasets (Huang et al. 2012; Yu et al. 2012; Huang et al. 2019). Dramatically induced expression of OsAOS1–4 was observed in the roots of plants in response to 20, 25 or 80 μM As(III) for 6 h to 72 h, while transcripts of OsAOS3 and OsAOS4 in the shoots were not responsive to As(III) treatment (Fig. 2B). The enhanced expression of OsAOS1 in the root tip (0–2 cm of cv. Zhonghua 11) was observed with 3 μM As(V) treatment for 1 h but rapidly decreased at 6 h and 24 h of As(V) treatment. On the other hand, the transcripts of OsAOS2 were increased progressively along with the treatment time under 3 μM As(V) (Fig. 2C). Under the treatment of 25 μM As(V), significantly increased transcripts of OsAOS1 and OsAOS2 in roots were also detected, but the expression of OsAOS3 and OsAOS4 was hardly affected by As(V) (Fig. 2C). All these results indicated that two key enzymes for JA biosynthesis OsAOS1 and OsAOS2 respond to both As(III) and As(V) in rice, rendering further functional analysis.
Mutation of OsAOS1 increases grain As accumulation
To investigate the molecular function and physiological role of OsAOS1 and OsAOS2, the gene-edited lines generated through clustered regularly interspaced short palindromic repeats/CRISPR–associated nuclease 9 (CRISPR/Cas9) were obtained from the commercial rice genome-scale mutagenesis library (Lu et al. 2017). The target sites were in the exons of OsAOS1 and OsAOS2, respectively (Fig. S2A, S3A). Sequencing results identified a 3-bp deletion (c.156_158delGGA) and a 1-bp insertion plus a substitution from G by T (c.157G > T + c.158inT) in the offspring of T2 generation of OsAOS1 mutant lines, leading to an amino acid (p.Asp53del) deletion or frameshift-induced pre-stop (p.Asp53TyrfsTer152), respectively (Fig. S2B). The mutants were then designated as osaos1–1 and osaos1–2. Truncated OsAOS2 was observed in the mutant line osaos2–1 (p. Pro58AlafsTer424), which was resulted from a 1-bp insertion (c.171inC) in the coding region and corresponding frameshift (Fig. S3B). The deleted amino acid Asp(D)53 in osaos1–1 was conserved with the putative homologues of AOSs from Brachypodium distachyon, Physcomitrella patens, and Marchantia polymorpha but not in Arabidopsis thaliana or Hordeum vulgare (Fig. 1C), and the predicted 3D structure of this mutant was almost identical to that of the functional OsAOS1 (Fig. S2C). In addition, the mutation site of osaos2–1 occurred in a putative conserved domain among land plants (Fig. S1B), and the predicted 3D structure also confirmed the large fragment deletion (Fig. S3C).
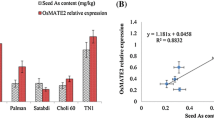
Due to the extremely low seed setting of osaos1–2, the homozygous mutant lines, osaos1–1 and osaos2–1 were selected for the further analysis. Five independent plants of each mutant line and the corresponding wild-type plants (WT, cv. Zhonghua 11) were grown to maturity in pots with paddy soil. There was little variation on the seed setting rate, grain weight between the mutant lines and the WT plants (Fig. 3A, B). Grain yield per plant of osaos1–1 and osaos2–1 was slightly lower (not significant P > 0.05) than that of WT (Fig. 3C). In comparison with WT, significant increase of As and manganese (Mn) concentrations were found in the brown rice grains of osaos1–1, while there was no significant difference in the concentrations of iron (Fe), and copper (Cu) between WT and osaos1–1 (Fig. 3D). Significant decrease of Zn concentration was observed in the brown rice of osaos1–1 and osaos2–1, while mutation of OsAOS2 hardly affected the accumulation of the other examined metals (Fig. 3D). The results indicated that the deletion of Asp53 in OsAOS1 hardly affect the yield but significantly increase As accumulation in rice grain.
Yield-related traits and mineral concentrations in the grain of osaos1–1, osaos2–1 and WT. Seed setting rate (A), grain weight (B) and grain yield per plant (C) of osaos1–1, osaos2–1 and WT grown in soil-contained pots until mature. D The concentrations of As, Cu, Fe, Mn and Zn in the brown rice of the mutant lines and WT. Values are means of three independent replicates ± SE (n = 4 for A-C, and n = 10 for D). Different letters indicate significant differences in each graph (p < 0.05)
Mutation of OsAOS1 increases As translocation from root to shoot
When the plants were treated with 2 and 5 μM As(V) for 6 d, no difference of As concentrations in the root of the plants was observed (Fig. 4A), however, significantly increased As concentration was found in the shoots of osaos1–1 compared to that of WT in 2 μM As(V) (Fig. 4B). Significantly lower total As was taken up by the roots and accumulated in the whole plants of osaos1–1 than that of WT (Fig. 4C), while the ratio of As translocated from root to shoot in osaos1–1 was significantly higher than that of WT (Fig. 4D). Xylem sap As concentration was higher (no statistical significance) in osaos1–1 and osaos2–1 than that of WT subjected to 2 μM As(V) for 30 min (Fig. 4F). However, mutation of OsAOS2 hardly affected As distribution between root and shoot (Fig. 4).
As accumulation and distribution in the seedlings of osaos1–1, osaos2–1 and WT at vegetative growth stage. As concentration in the root (A), shoot (B) of the plants harboring 3–4 leaves subjected to 2 or 5 μM As(V) for 6 days. Total As taken up by the plants (C) and the ratio of As translocated from root to shoot of each genotype (D) was calculated. E Increased plant height before and after the As treatments. F As in the xylem sap collected from the 30-day-old plants treated with 2 μM As(V) for 0.5 hour. Values are means of three independent replicates ± SE (n = 4). Different letters indicate significant differences in each graph (p < 0.05)
As concentrations were significantly decreased in the roots of osaos1–1 and osaos2–1 compared to that of WT when the plants were cultured with 2 or 5 μM As(III) for 6 d (Fig. S4A). The total As uptake by the mutant lines was also much lower than that of WT (Fig. S4C). However, there was no difference in shoot As concentration and As distribution ratio between root and shoot of the three genotypes (Fig. S4B, D). We then compared As efflux activity from root to external solution after the 15-d-old plants in short-term (4 h) As(V) treatment. After transferring to the nutrient solution without As for 6 h, As efflux percentage (%) was calculated according to the As accumulated in plant root, shoot and presented in the external solution. The results revealed that mutation of OsAOS1 or OsAOS2 did not affect As efflux activity (Fig. S5). Thus, our results suggested that increased grain As in osaos1–1 mutant is likely attributed to the elevated As translocation from root to shoot subjected to As(V).
OsAOS1 and OsAOS2 are the key components of As tolerance in rice
When the plants were subjected to As(V) treatment, the growth of WT and both aos mutant lines were significantly affected (Fig. S6). Compared to the WT, remarkably lower increased plant height in both osaos1–1 and osaos2–1 was observed after the treatment for 4 d in 2 μM As(V), while the difference could be observed in the treatment with 5 μM As(V) from 2 d (Fig. 4E). The root and shoot dry weights osaos1–1 were significantly lower than those of WT, but there was no difference between osaos2–1 and WT in terms of biomass (Fig. S6A, B). Interestingly, the root and shoot dry weights displayed no difference among osaos1–1, osaos2–1 and WT subjected to 2 or 5 μM As(III) for 6 d (Fig. S7). It’s implicated that osaos1–1 and osaos2–1 are more sensitive to As(V) than As(III).
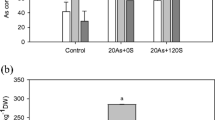
Compared to those of WT, curly and chlorosis young leaves, and incompact plant architecture were more clearly presented in osaos1–1 under the highest As(V) condition (Fig. 5A-H, a-c). In addition, significantly reduced root and shoot dry weights were observed in the mutant lines. Particularly at 2 μM As(V), root dry weight was decreased by 25% and 32% in osaos1–1 and osaos2–1 while shoot dry weight was reduced by 17% and 26% in osaos1–1 and oasos2–1, respectively (Fig. 5I, J).
Plant growth and biomass of osaos1–1, osaos2–1 and WT under As(V) treatment. Ten-day-old seedlings of the three genotypes subjected to 0 (A, E), 2 (B, F), 5 (C, G) or 10 μM As(V) (D, H) for 10 days. The aboveground tissues (A-D) and whole plants (E-H) were recorded, and the youngest leaves of the plants under 10 μM As(V) were zoomed in a-c. The dry weight of root (I) and shoot (J) of the plants were measured. Values are means of three independent replicates ± SE (n = 4). Different letters indicate significant differences in each graph (p < 0.05)
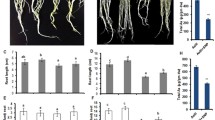
The root length and relative elongation of seminal roots of the three genotypes under As treatments showed no difference under control condition, but both were significantly reduced root length in osaos1–1 subjected to 15 μM As(III) or 2 μM As(V) for 2 d (Fig. 6A and B). The retarded root growth was also found when osaos2–1 was cultured in the presence of As(III) and As(V) for 24 h (Fig. 6B), but the inhibition of root elongation by As(III) was only statistically significant at 0–24 h (Fig. 6B).
Relative root elongation of osaos1–1, osaos2–1 and WT subjected to As. Two-day-old seedlings of the plants were treated with 15 μM As(III) or 2 μM As(V) for 2 days, the length of the seminal root was measured (A) and the relative root elongation rate was calculated (B). Values are means of three independent replicates ± SE (n = 10). Different letters indicate significant differences in each graph (p < 0.05)
Disrupted gene expression in the mutant lines of OsAOS1 and OsAOS2
To dissect the potential molecular mechanisms underlying the above observations, the expression levels of the critical genes involved As accumulation and detoxification mediated by OsAOS1 and OsAOS2 were determined. Among the genes for As(V) uptake including several Phosphate (P) transporters (OsPT1 OsPT4, and OsPT8), upregulated expression of OsPT8 was observed in the shoots of osaos2–1 and dramatic downregulation of OsPT4 was found in the root of both osaos1–1 and osaos2–1 in the presence of 2 μM As(V) for 6 d (Fig. 7A, B). Significant down-regulated expression of OsNIP3;2 was also observed in the root of the two mutant lines (Fig. 7C). Significant upregulation of Low Silicon 6 (OsLsi6) was induced by 2 μM As(V) in the shoot and 5 μM As(V) in the root of osaos1–1. However, expression of OsLsi1, OsLsi2, OsLsi3, OsABCC1, OsABCC7, OsHAC1;1, OsHAC1;2, and OsHAC4 remained unchanged among the genotypes in response to As(V) treatments (Fig. S8). Significant downregulation of OsOASTL1-A1 involved in As detoxification was identified in the root of both mutant lines subjected to 2 μM As(V) compared to that of WT (Fig. 7C). As a result, the altered expression of the genes involved in As accumulation and tolerance was obvious in the mutant lines of OsAOS1 and OsAOS2.
Expression of genes involved in As accumulation and detoxification in rice. After being treated with 2 or 5 μM As(V) for 6 days (Same condition as that in Fig. 4A-D), the root and shoot were harvested for RNA extraction and 1-st strand cDNA synthesis. The relative expression levels of the genes were compared through quantitative real-time PCR. Values are means of three independent replicates ± SE (n = 3). Different letters indicate significant differences in each graph (p < 0.05). Abbreviations: Phosphate (P) transporters (OsPT1,OsPT4, OsPT8), Nodulin 26-like Intrinsic Protein (OsNIP3;2), O-acetylserine (thiol) lyase (OsOASTL-A1)
Decreased reactive oxygen species (ROS) accumulation in the root of osaos1–1
ROS can be rapidly induced by 25–400 μM As in plants (Deng et al. 2020; Sharma et al. 2021), and MeJA alleviates As toxicity in rice partially through ROS scavenging (Lu et al. 2020). These shed some light on the potential involvement of OsAOS1 and OsAOS2 in As detoxification through ROS homeostasis. It was found that no significant difference in ROS fluorescence was observed in the root tip (0–5 mm) and basal root region (7–12 mm) of WT exposed to 2 μM As(V) for 1 and 6 h. There was no difference in ROS production with 1 hour As(V) treatment between osaos1–1 and WT (Fig. 8) However, the fluorescent signal intensity was dramatically decreased in both root tip and basal root region of osaos1–1 in response to 2 μM As(V) for 6 hours (Fig. 8). Interestingly, mutation of OsAOS2 hardly affected the accumulation of ROS in root in both control and As(V) treatment (Fig. 8). The result indicated that mutation of OsAOS1 but not OsAOS2 showed disrupted ROS homeostasis, which could be related to a higher root to shoot As translocation rate in osaos1–1 mutant.
ROS accumulation in the roots of osaos1–1, osaos2–1 and WT subjected to As(V). After being treated with 2 μM As(V) for 1 or 6 hours, the seminal root of the 3-day-old seedlings were stained with 10 μM CM-H2DCFDA for 1 hour and was washed for 3 times was employed for ROS detection. The representative figures were present in (A) and (B). The signal intensity of root tip (0–5 mm) (C) and basal root region (7–12 mm) (D) was calculated from 5 independent replicates. Values are means of three independent replicates ± SE (n = 5). Different letters indicate significant differences in each graph (p < 0.05). Scale bar = 100 μm
Discussion
OsAOS1 is involved in As accumulation in rice
Arsenic contamination in plant derived-food is a worldwide health concern (Zhao et al. 2022). Several pathways for phytohormone metabolism and signaling transduction have been identified in response to As stress (Hu et al. 2020; Chen et al. 2021; Deng et al. 2021; Sharma et al. 2021; Li et al. 2023; Zhang et al. 2023a). It was reported that the transcripts of key genes involved in JA biosynthesis and signaling are significantly regulated by As(III) and As(V) in rice, barley, and Arabidopsis (Huang et al. 2012; Yu et al. 2012; Srivastava et al. 2015; Zvobgo et al. 2018). Application of exogenous JA partially alleviated As(V)-induced rice root inhibition, and pretreated with JA decreased As accumulation in both shoot and root of rice seedlings subjected to As(V) (Wang et al. 2018; Ronzan et al. 2019; Mousavi et al. 2020; Li et al. 2022a, 2022b; Zhang et al. 2023b). These results implicated the involvement of JA in As accumulation and detoxification in plants. However, whether As translocation and tolerance were controlled by JA biosynthesis and signaling is poorly understood. Here, we demonstrated that OsAOS1, a putative enzyme for JA biosynthesis, limits grain As accumulation (Fig. 3D). In addition, the contribution of OsAOS1 to As accumulation in rice was preferentially more effective when subjecting plants to As(V) instead of As(III) (Fig. 4, S4).
The physiological processes and some critical proteins involved in As accumulation in rice grain and As distribution among various tissues have been revealed recently (Deng et al. 2020; Deng et al. 2021; Tang and Zhao 2021; Yamaji and Ma 2021; Zhao et al. 2022). Briefly, the uptake and translocation from root to shoot of arsenous acid [As(OH)3] is achieved by the effective cooperation of two silicon (Si) transporters, OsNIP2;1 (OsLsi1, low silicon rice 1), and OsLsi2 (Ma et al. 2008). The homologues of OsLsi1, OsNIP3;2, is involved in As(III) uptake by lateral roots (Chen et al. 2017). The aquaporins including OsLsi1, OsNIP1;1 and OsNIP1;3 were also responsible for the efflux of As(III) (Zhao et al. 2010; Sun et al. 2018), which contribute to As accumulation. The tonoplast-localized C-type ATP-binding cassette transporter OsABCC1 plays critical roles in limiting As accumulation in rice grain and in As detoxification through sequestration of As-phytochelatins (As-PCs) into vacuole (Song et al. 2014; Deng et al. 2018), while OsABCC7 is involved in the root-to-shoot translocation of As in rice (Tang et al. 2019). Phosphate (P) transporters OsPT1, OsPT4 and OsPT8 also function in the uptake of As(V) and accumulation of As, and the contribution of OsPT4 in the inorganic As accumulation in rice grain was estimated as 20 ~ 44% (Kamiya et al. 2013; Wang et al. 2016; Cao et al. 2017), which implicated that As(V) could be considered as a major source of As contamination for rice grown in anoxic condition. Furthermore, the As(V) absorbed by rice roots can be readily transformed to As(III) in the presence of arsenate reductases including OsHAC1;1, OsHAC1;2 and OsHAC4, which also contribute to the accumulation and detoxification of As in rice plants (Shi et al. 2016; Xu et al. 2017).
Here, we found mutation of OsAOS1 significantly increased As content in the brown rice grains of the plants grown in soils with additional As (Fig. 3D), which was probably through increasing As translocation from root to shoot but not via As uptake and efflux (Fig. 4 and Fig. S5). Interestingly, the increased As translocation from root to shoot was only observed in osaos1–1 exposed to As(V) but not to As(III) (Fig. 4 and Fig. S4). The results suggested that the contribution of OsAOS1 to As accumulation was likely dependent on the pathway of phosphate (P) but not silicic acid accumulation, which was consistent with the altered expression of OsPT1, and OsPT4 in osaos1–1 (Fig. 7). While the expression of genes including OsLsi1, OsLsi2, OsLsi3 and OsLsi6 responsible for Si uptake, root-to-shoot translocation and distribution was hardly affected except the increased transcription of OsLsi6 in osaos1–1 (Fig. S8). The expression of OsHAC1;1, OsHAC1;2, and OsHAC4 was hardly affected by the mutation of OsAOS1 or OsAOS2 (Fig. S8), implicating that the deoxidize of As(V) to As(III) in rice root cells was likely not affected by JA pathway.
The role of OsAOS1 in alleviating As toxicity in rice
To adapt to the environmental stresses, plants have evolved various strategies to prevent themselves from the damage. JA and its derivatives represent a group of phytohormones, eliciting various effects on the tolerance of plant to both biotic and abiotic stresses (Yu et al. 2019). Pretreatment with JA or MeJA alleviates As toxicity in rice, especially in root growth (Wang et al. 2018; Ronzan et al. 2019; Mousavi et al. 2020). Consistently, enhanced sensitivity including inhibited root elongation and withered young leaves was observed in osaos1–1 and osaos2–1 subjected to both As(III) and As(V) compared to that of WT (Figs. 5, 6 and Fig. S6), indicating that OsAOS1 and OsAOS2 may confer to As detoxification.
The alleviated effects mediated by JA was widely observed in plants subjected to cadmium (Cd), salt, drought and other abiotic stresses (Yu et al. 2019; Kamiya et al. 2013; Wang et al. 2020). Consistent with the rapidly induced expression of OsAOS1 and OsAOS2 in rice (Fig. 1B, C) (Huang et al. 2012; Yu et al. 2012), the expression of genes promoting endogenous JA synthesis was also upregulated by Cd treatment in Arabidopsis (Lei et al. 2020). Exogenous application of MeJA partially alleviated Cd-generated chlorosis of new leaves (Lei et al. 2020). Consistently, mutation of OsAOS1 or OsAOS2 enhanced the curling and chlorosis of young leaves (Fig. 5). Increased shoot As and reduced root As concentrations were observed in the rice mutant coleoptile photomorphogenesis2 (cpm2), the loss-of-function of Allene Oxide Cyclase (AOC) required for JA synthesis (Ronzan et al. 2019), which was very similar to our finding in osaos1–1 (Fig. 4). Furthermore, the phenotype of retarded root system of cpm2 exposed to As is similar to that of osaos1–1 and osaos2–1 (Figs. 5 and 6) (Ronzan et al. 2019). Knockout of AtAOS increased Cd concentration in both roots and shoots, and confered to the increased sensitivity to Cd, which could be recovered by exogenous MeJA (Lei et al. 2020). The expression of genes responsible for Cd uptake and long-distance transport such as AtIRT1, AtHMA2 and AtHMA4 was greatly upregulated in ataos mutant but decreased in the presence of MeJA (Lei et al. 2020), however, only the translocation of As from root to shoot but not the uptake and efflux of As was not affected in osaos1–1 (Fig. 4, S5).
ROS homeostasis in plant cells also confers the detoxification of heavy metals (Huang et al. 2012; Smirnoff and Arnaud 2019; Nazir et al. 2020). Application of exogenous ROS also decreased the accumulation of heavy metals in root through reducing root uptake and facilitating root-to-shoot translocation of heavy metals (Cheng et al. 2023; Deng et al. 2023). It’s implicated that the ameliorating effects of JA partially rely on ROS scavenging (Chen et al. 2021). In this study, dramatically decreased ROS was detected in the mutant lines of OsAOS1 but not OsAOS2 in response to As(V) (Fig. 8), which might be one of the possible strategies for As detoxification mediated by OsAOS1. The reason for the increased As sensitivity of osaos2–1 is probably attributed to the reduced expression level of OsABCC1 and OsOASTL-A1 (Fig. 7, S8).
Synergistic interplay of one of the major ROS - hydrogen peroxide (H2O2) with JA was found in plants. For example, application of MeJA induced accumulation of H2O2 in the detached leaves of rice (Hung et al. 2006), while H2O2 could act as a signaling molecule for the MeJA-triggered antioxidant defense in wheat (Nazir et al. 2020). H2O2 at nanomolar levels is helpful in the maintenance of cellular homeostasis in crops, while elevated levels of H2O2 trigger oxidative burst and lead to cell death (Smirnoff and Arnaud 2019). Therefore, the production, intracellular transport and scavenging of ROS should be tightly regulated. In this study, ROS accumulation was inhibited in the root of osaos1–1 subjected to As(V) compared to that of WT (Fig. 8), which is probably attributed to the increased As translocation from root to shoot (Fig. 4).
In summary, our results indicate that OsAOS2 is important for As tolerance, while OsAOS1 confers both As allocation and detoxification, which could be partially attributed to the indirect effects of altered gene expression profiling and ROS homeostasis (Fig. 9).
OsAOS1 and OsAOS2 contribute to differential accumulation and detoxification of As in rice. OsAOS1 (black routes) and OsAOS2 (green routes) are the key enzymes for the biosynthesis of JA in rice. OsAOS1 limited As accumulation in rice grain and As detoxification, which is partially attributed to the indirectly altered expression of genes such as OsPT1, OsPT4, OsNIP3;2, OsLsi6 and OsOASTL-A1. ROS homeostasis likely contributes to the regulation of As transport and tolerance mediated by OsAOS1 but not OsAOS2. Increased As sensitivity in the loss-of-function lines of OsAOS2 is largely depended on the changed expression of OsPT8, OsNIP3;2, OsLsi6 and OsOASTL-A1
Materials and methods
Plant materials and growth conditions
The mutant lines of OsAOS1 and OsAOS2 generated through CRISPR/Cas9 were obtained from the commercial rice genome-scale mutagenesis library, Biogle Genome Editing Center (Lu et al. 2017). The homozygous mutant lines, osaos1–1 and osaos2–1 were isolated through the sequencing results of the amplified DNA fragments containing the target sites of each candidate genes. Primers used in the study are listed in Table S1.
The two mutant lines together with the corresponding wild-type rice Zhonghua 11 (ZH11, Oryza sativa ssp. Japonica) were cultured in hydroponic condition or paddy soil for the phenotypic analyses. Briefly, rice seeds of each genotype were soaked in tap water at 37 °C for 2–3 d for germination, the seedlings were transferred to a net floating on a 0.5 mM CaCl2 solution for 3 days. After about 3–4 d, the plants were moved to the pots (13 L) containing one-half-strength Kimura B solution (Deng et al. 2013) for 6 d, and then the plants were transferred to smaller pots (4.2 L) for the further treatment. The nutrition solution was renewed every 2 d. The hydroponic solution was replaced by one-half-strength Kimura B solution without phosphorus (P) for 2 d and then followed with As-contained low-phosphorus (1 μM P) solution when the plants were treated with As(V) treatment. For treatment of plants with As(III), the required As concentrations were added directly to the one-half-strength Kimura B solution. The concentration of phosphorus in the normal one-half-strength Kimura B solution is 90 μM.
For hydroponic culture, the plants were grown in a growth chamber at 25–30 °C under long-day condition (16-h/8-h day/night photoperiod). The concentrations of As(III) and As(V) employed for the treatments were decided by the growth stage, the organs of the plants which we mainly focus on, as well as their varied toxicity. Three concentrations of phosphorus (P) were employed under different conditions in this study, the plants without As(V) treatments were cultured with half-strength Kimura B solution (90 μM P), before As(V) treatment, the plants were pre-cultured with half-strength Kimura B solution without P, and if the treatment with As(V) last for more than two days, the final P concentration in the solution was 1 μM.
For soil culture, the plants were grown in the paddy soil-contained pots (30 L) until fully mature under normal condition since May to August of 2022, which were conducted in the west campus of Yangtze University located in Jingzhou, Hubei Province, China. The hydro-soluble As concentration in the soil was about 2 mg/kg DW.
Bioinformatics analysis of OsAOSs
The phylogenetic analysis of AOS members was performed according to our previous reports (Deng et al. 2022; Jiang et al. 2023). Briefly, the homologues of AOS from representative plant and algal species were identified through BLASTP searches from the OneKP database (https://db.cngb.org/blast/blast/blastp/) using OsAOS1 (OsAOS2, Supplementary Fig. 1) as the reference, the candidate sequences that satisfied the criteria, E value < 10−10 and query coverage > 50% were chosen for the further analysis. The phylogenetic tree was generated and displayed by CIPRES Science Gateway V 3.3 (https://www.phylo.org) and iTOL 6.0 visualization (https://itol.embl.de/), respectively. Furthermore, the multiple alignments of AOS1 homologues were conducted using MAFFT with the default setting (https://mafft.cbrc.jp/alignment/server/), and the 3D structure was predicted via SWISS-MODEL (https://swissmodel.expasy.org/interactive).
Gene expression pattern analysis
To investigate the expression pattern of OsAOS genes in various organs, the rice plants (cv. Nipponbare) grown in paddy soil until grain filling stage, three independent samples of root, node II, Node I, internode II, internode I, leaf blade II, leaf blade I, rachis, and spikelet were sampled for RNA extraction and transcriptomic analysis (Personal Biotechnology Co., Shanghai, China). To examine the response of OsAOS genes to As(III) and 25 μM As(V), the datasheets derived from the previous studies were employed (Huang et al. 2012; Yu et al. 2012; Huang et al. 2019). To understand the expression pattern of OsAOS genes under As(V) conditions, the 0–2 cm length from the tip of seminal roots of WT (ZH11, 2 d old) seedlings subjected to 3 μM As(V) for 1, 6 and 24 hours were sliced for RNA-sequencing (Personal Biotechnology Co., Shanghai, China). The mean value of three FPKM (Fragments Per Kilobase of exon model per Million mapped fragments) of each gene (Fig. 1A), or the fold change compared to those under control conditions (Fig. 1B, C) were calculated for the generation of heat map, which were displayed by using TBtools (Chen et al. 2020).
Phenotypic analysis at vegetative and reproductive stages
Two concentrations (2 and 5 μM) of As(III) [NaAsO2] and As(V) [Na2HAsO4] respectively were added to the hydroponic solutions for the determination of As accumulation 3 to 4-leaf stages in the tissues osaos1–1, osaos2–1 and WT plants. The nutrient solution was renewed every 2 d. The growth parameters of each plant including plant height were recorded every 2 d. After 6 d of As treatments, the leaves and the roots were harvested after being washed with pre-chilled 0.5 mM CaCl2 solution for 3 times. The fresh weight (FW, mg) of each sample was measured and then dried at 65 °C for 3 d.
For As tolerance investigation, 10-d-old seedlings of the three genotypes were treated with 2, 5, and 10 μM As(V)-contained one-half-strength Kimura B solution with 1 μM P for 10 d, and then the samples were harvested and their dry weight were measured. Four and eight biological replicates were used for dry weight and plant height, respectively. The experiments were repeated at least twice.
Five plants of each genotype (osaos1–1, osaos2–1, and WT) were grown in soil in a 30-L pot with 5 replicates for each genotype. At harvested, the seed setting ratio were calculated according to our previous study by using 8.5% NaCl solution for selection (Deng et al. 2013). The grain yield per plant and the weight per 1000 seeds were recorded. Brown rice sets from each plant were randomly selected for further digestion and metal measurement.
Determination of metals in plant tissues and xylem sap
For xylem sap collection, 30-day-old seedlings of osaos1–1, osaos2–1, and WT were employed. The hydroponic solution was based on one-half-strength Kimura B solution without P for 2 d, and then 2 μM As(V) was added into the pots and mixed well with glass rod, xylem sap was collected after the plants were treated for 30 mins with a pipette for 0.5 h (Deng et al. 2013; Tang et al. 2019). Five independent samples from each genotype and treatment were collected and then diluted with 2% HNO3 for metal concentration determination by inductively coupled plasma-mass spectrometry (ICP-MS, NexION 1000; PerkinElmer). The dried shoot, root, basal region, and brown rice without husk was digested with concentrated HNO3 at a temperature up to 140 °C. The metal concentrations in the digested solution were determined by Inductively Coupled Plasma-Mass Spectrometry (ICP-MS) after dilution with MQ water (Cheng et al. 2023; Ning et al. 2023).
Measurement of As efflux activity
For As efflux activity test, the germinated seeds of each genotype (osaos1–1, osaos2–1, and WT) were grown in one-half-strength Kimura B solution until 12-d-old, and then the plants were pre-treated one-half-strength Kimura B without P for 2 days. The plants were moved in to the 5 μM As(V)-contained one-half-strength Kimura B solution (−P) for 4 hours, the roots were washed with 0.5 mM CaCl2 and the plants were transferred into 110 mL black bottles with one-half-strength Kimura B solution (−P) for efflux. After 6 h, 1 mL solution was collected for As concentration measurement, the roots and shoots were harvested after being washed with pre-cold 0.5 mM CaCl2 solution for 3 times. One plant for one black bottle, and 5 replicates were set for each genotype in this experiment. As efflux ratio was calculated as the following formula: As in solution/[As in solution + As in root + As in shoot] × 100%.
Relative root elongation under As
To compare the As tolerance of osaos1–1, osaos2–1, and WT, 2-day-old seedlings were exposed to a 0.5 mM CaCl2 solution (pH 5.6) containing 15 μM As(III) or 2 μM As(V) for 2 d. The root length of each seedling was measured with a ruler before and after the treatments for 24 and 48 h, and the relative root elongation (root elongation with As/root elongation without As × 100%) was calculated. Fifteen replicates were conducted for each treatment, and the experiments were repeated at least twice.
RNA extraction and quantitative real-time PCR
The roots and shoots of osaos1–1, osaos2–1, and WT subjected to 2 or 5 μM As(V) for 6 d were harvested for RNA extraction, which is following the user manual of TRIpure Reagent (Aidlab). The first strand of cDNA synthesized using the TRUEScript RT Kit (+ gDNA Eraser) reverse transcription kit (Aidlab) was employed as the template. Quantitative real-time PCR (qPCR) with 3 biological replicates was performed according to our previous report (Jiang et al. 2023). Ct values were normalized to the corresponding endogenous control gene (OsActin1, LOC_Os03g50885), the relative expression levels of the genes involved in As uptake, translocation, and detoxification and JA-responsive marker genes were calculated using the ΔΔ Ct method (Huang et al. 2016). The primers used in this study are shown in Supplemental Table S1.
Reactive oxygen species detection
After germination for 3 d on the floating net with the supplement of 0.5 mM CaCl2, the seedlings of osaos1–1, osaos2–1, and WT were treated with 2 μM As(V). To detect the accumulation of ROS, the roots of the plants subjected to As(V) for 1 and 6 h were stained with 10 μM ROS indicator CM-H2DCFDA (C6827, Thermo Fisher Scientific) in PBS solution for 1 h under dark condition. The green fluorescent signal in the root region was observed after being washed with PBS for 3 times and the photos were taken with a fluorescent microscope. The plants without As(V) treatment were employed as controls. The intensity of the signal was captured and calculated with the software ImageJ (version 1.8.0, National Institutes of Health, USA) (Cheng et al. 2023). Five seedlings were used for each treatment.
Data analysis
Data analysis was conducted by using SPSS software, with the statistical significance analysis performed using one-way analysis of variance (ANOVA) or the least significant difference (LSD) test for mean comparison (at the 5% level).
Availability of data and materials
The author responsible for distribution of materials integral to the findings presented in this article is Fenglin Deng (dfl@yangtzeu.edu.cn).
Abbreviations
- AOS:
-
Allene oxide synthase
- As(III):
-
Arsenite
- As(V):
-
Arsenate
- As:
-
Arsenic
- CM-H2DCFDA:
-
5-(and-6)-chloromethyl-2′, 7′-dichlorodihydrofluorescein diacetate, acetyl ester
- CRISPR/Cas9:
-
Clustered regularly interspaced short palindromic repeats–associated nuclease 9
- ICP-MS:
-
Inductively coupled plasma-mass spectrometry
- JA:
-
Jasmonic acid
- JA-Ile:
-
Jasmonoyl-l-isoleucine
- MeJA:
-
Methyl jasmonate
References
Abbasi S, Lamb D, Rahman MA et al (2021) Response of phosphorus sensitive plants to arsenate. Environ Technol Innov 24:102008, ISSN 2352-1864. https://doi.org/10.1016/j.eti.2021.102008
Antoniadis V, Shaheen SM, Levizou E, Shahid M, Niazi NK, Vithanage M, Ok YS, Bolan N, Rinklebe J (2019) A critical prospective analysis of the potential toxicity of trace element regulation limits in soils worldwide: are they protective concerning health risk assessment? - a review. Environ Int 127:819–847. https://doi.org/10.1016/j.envint.2019.03.039
Azizur Rahman M, Hasegawa H, Mahfuzur Rahman M, Nazrul Islam M, Majid Miah MA, Tasmen A (2007) Effect of arsenic on photosynthesis, growth and yield of five widely cultivated rice (Oryza sativa L.) varieties in Bangladesh. Chemosphere 67:1072–1079. https://doi.org/10.1016/j.chemosphere.2006.11.061
Cao Y, Sun D, Ai H, Mei H, Liu X, Sun S, Xu G, Liu Y, Chen Y, Ma LQ (2017) Knocking out OsPT4 gene decreases arsenate uptake by rice plants and inorganic arsenic accumulation in rice grains. Environ Sci Technol 51:12131–12138. https://doi.org/10.1021/acs.est.7b03028
Chehab EW, Perea JV, Gopalan B, Theg S, Dehesh K (2007) Oxylipin pathway in rice and Arabidopsis. J Integr Plant Biol 49:43–51. https://doi.org/10.1111/j.1744-7909.2006.00405.x
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen X, Jiang W, Tong T, Chen G, Zeng F, Jang S, Gao W, Li Z, Mak M, Deng F, Chen ZH (2021) Molecular interaction and evolution of jasmonate signaling with transport and detoxification of heavy metals and metalloids in plants. Front Plant Sci 12:665842. https://doi.org/10.3389/fpls.2021.665842
Chen Y, Hua CY, Chen JX, Rathinasabapathi B, Cao Y, Ma LQ (2019) Expressing arsenite antiporter PvACR3;1 in rice (Oryza sativa L.) decreases inorganic arsenic content in rice grains. Environ Sci Technol 53:10062–10069. https://doi.org/10.1021/acs.est.9b02418
Chen Y, Sun SK, Tang Z, Liu G, Moore KL, Maathuis FJM, Miller AJ, McGrath SP, Zhao FJ (2017) The Nodulin 26-like intrinsic membrane protein OsNIP3;2 is involved in arsenite uptake by lateral roots in rice. J Exp Bot 68:3007–3016. https://doi.org/10.1093/jxb/erx165
Cheng J, Zhang S, Yi Y, Qin Y, Chen ZH, Deng F, Zeng F (2023) Hydrogen peroxide reduces root cadmium uptake but facilitates root-to-shoot cadmium translocation in rice through modulating cadmium transporters. Plant Physiol Biochem 200:107754. https://doi.org/10.1016/j.plaphy.2023.107754
Clemens S, Ma JF (2016) Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu Rev Plant Biol 67:489–512. https://doi.org/10.1146/annurev-arplant-043015-112301
Das N, Bhattacharya S, Bhattacharyya S, Maiti MK (2018) Expression of rice MATE family transporter OsMATE2 modulates arsenic accumulation in tobacco and rice. Plant Mol Biol 98:101–120. https://doi.org/10.1007/s11103-018-0766-1
Deng F, Liu X, Chen Y, Rathinasabapathi B, Rensing C, Chen J, Bi J, Xiang P, Ma LQ (2020) Aquaporins mediated arsenite transport in plants: molecular mechanisms and applications in crop improvement. Crit Rev Environ Sci Technol 50:1613–1639. https://doi.org/10.1080/10643389.2019.1662704
Deng F, Yamaji N, Ma JF, Lee SK, Jeon JS, Martinoia E et al (2018) Engineering rice with lower grain arsenic. Plant Biotechnol J 16:1691–1699. https://doi.org/10.1111/pbi.12905
Deng F, Yamaji N, Xia J, Ma JF (2013) A member of the heavy metal P-type ATPase OsHMA5 is involved in xylem loading of copper in rice. Plant Physiol 163:1353–1362. https://doi.org/10.1104/pp.113.226225
Deng F, Yu M, Martinoia E, Song WY (2019) Ideal cereals with lower arsenic and cadmium by accurately enhancing vacuolar sequestration capacity. Front Genet 10:322. https://doi.org/10.3389/fgene.2019.00322
Deng F, Zeng F, Chen G, Feng X, Riaz A, Wu X et al (2021) Metalloid hazards: from plant molecular evolution to mitigation strategies. J Hazard Mater 409:124495. https://doi.org/10.1016/j.jhazmat.2020.124495
Deng F, Zeng F, Shen Q, Abbas A, Cheng J, Jiang W et al (2022) Molecular evolution and functional modification of plant miRNAs with CRISPR. Trends Plant Sci 27:890–907. https://doi.org/10.1016/j.tplants.2022.01.009
Deng M, Wang S, Huang H, Ye D, Zhang X, Wang Y et al (2023) Hydrogen peroxide mediates cadmium accumulation in the root of a high cadmium-accumulating rice (Oryza sativa L.) line. J Hazard Mater 448:130969. https://doi.org/10.1016/j.jhazmat.2023.130969
Duan G, Kamiya T, Ishikawa S, Arao T, Fujiwara T (2012) Expressing ScACR3 in rice enhanced arsenite efflux and reduced arsenic accumulation in rice grains. Plant Cell Physiol 53:154–163. https://doi.org/10.1093/pcp/pcr161
Duan G, Shao G, Tang Z, Chen H, Wang B, Tang Z et al (2017) Genotypic and environmental variations in grain cadmium and arsenic concentrations among a panel of high yielding rice cultivars. Rice (N Y) 10:9. https://doi.org/10.1186/s12284-017-0149-2
Faizan M, Sehar S, Rajput VD, Faraz A, Afzal S, Minkina T, Sushkova S, Adil MF, Yu F, Alatar AA, Akhter F, Faisal M (2021) Modulation of cellular redox status and antioxidant defense system after synergistic application of zinc oxide nanoparticles and salicylic acid in rice (Oryza sativa) plant under arsenic stress. Plants (Basel) 10(11):2254. https://doi.org/10.3390/plants10112254
Farooq MA, Zhang K, Islam F, Wang J, Athar HUR, Nawaz A et al (2018) Physiological and iTRAQ-based quantitative proteomics analysis of methyl jasmonate-induced tolerance in Brassica napus under arsenic stress. Proteomics 18:e1700290. https://doi.org/10.1002/pmic.201700290
Guan DX, Dai ZH, Sun HJ, Ma LQ (2021) Arsenic and selenium in the plant-soil-human ecosystem: CREST publications during 2018–2021. Crit Rev Environ Sci Technol 1-6. https://doi.org/10.1080/10643389.2021.2010836
Ha SB, Lee BC, Lee DE, Kuk YI, Lee AY, Han O, Back K (2022) Molecular characterization of the gene encoding rice allene oxide synthase and its expression. Biosci Biotechnol Biochem 66(12):2719–2722. https://doi.org/10.1271/bbb.66.2719
Haga K, Iino M (2004) Phytochrome-mediated transcriptional up-regulation of ALLENE OXIDE SYNTHASE in rice seedlings. Plant Cell Physiol 45:119–128. https://doi.org/10.1093/pcp/pch025
Hayashi S, Kuramata M, Abe T, Takagi H, Ozawa K, Ishikawa S (2017) Phytochelatin synthase OsPCS1 plays a crucial role in reducing arsenic levels in rice grains. Plant J 91:840–848. https://doi.org/10.1111/tpj.13612
Hu B, Deng F, Chen G, Chen X, Gao W, Long L, Xia J, Chen ZH (2020) Evolution of abscisic acid signaling for stress responses to toxic metals and metalloids. Front Plant Sci 11:909. https://doi.org/10.3389/fpls.2020.00909
Hu S, Yu K, Yan J, Shan X, Xie D (2023) Jasmonate perception: ligand-receptor interaction, regulation, and evolution. Mol Plant 16:23–42. https://doi.org/10.1016/j.molp.2022.08.011
Huang TL, Nguyen QT, Fu SF, Lin CY, Chen YC, Huang HJ (2012) Transcriptomic changes and signalling pathways induced by arsenic stress in rice roots. Plant Mol Biol 80:587–608. https://doi.org/10.1007/s11103-012-9969-z
Huang XY, Deng F, Yamaji N, Pinson SRM, Fujii-Kashino M, Danku J, Douglas A, Guerinot ML, Salt DE, Ma JF (2016) A heavy metal P-type ATPase OsHMA4 prevents copper accumulation in rice grain. Nat Commun 7:12138. https://doi.org/10.1038/ncomms12138
Huang Y, Chen H, Reinfelder JR, Liang X, Sun C, Liu C, Li F, Yi J (2019) A transcriptomic (RNA-seq) analysis of genes responsive to both cadmium and arsenic stress in rice root. Sci Total Environ 666:445–460. https://doi.org/10.1016/j.scitotenv.2019.02.281
Hung KT, Hsu YT, Kao CH (2006) Hydrogen peroxide is involved in methyl jasmonate-induced senescence of rice leaves. Physiol Plant 127:293–303. https://doi.org/10.1111/j.1399-3054.2006.00662.x
Jiang W, Tong T, Li W, Huang Z, Chen G, Zeng F, Riaz A, Amoanimaa-Dede H, Pan R, Zhang W, Deng F, Chen ZH (2023) Molecular evolution of plant 14-3-3 proteins and function of Hv14-3-3A in stomatal regulation and drought tolerance. Plant Cell Physiol 63:1857–1872. https://doi.org/10.1093/pcp/pcac034
Kamiya T, Islam R, Duan G, Uraguchi S, Fujiwara T (2013) Phosphate deficiency signaling pathway is a target of arsenate and phosphate transporter OsPT1 is involved in as accumulation in shoots of rice. Soil Sci Plant Nutr 59:580–590. https://doi.org/10.1080/00380768.2013.804390
Khanna K, Kohli SK, Kumar P, Ohri P, Bhardwaj R, Alam P, Ahmad P (2022) Arsenic as hazardous pollutant: perspectives on engineering remediation tools. Sci Total Environ 838:155870. https://doi.org/10.1016/j.scitotenv.2022.155870
Koeduka T, Ishizaki K, Mwenda CM, Hori K, Sasaki-Sekimoto Y, Ohta H, Kohchi T, Matsui K (2015) Biochemical characterization of allene oxide synthases from the liverwort Marchantia polymorpha and green microalgae Klebsormidium flaccidum provides insight into the evolutionary divergence of the plant CYP74 family. Planta 242:1175–1186. https://doi.org/10.1007/s00425-015-2355-8
Lei GJ, Sun L, Sun Y, Zhu XF, Li GX, Zheng SJ (2020) Jasmonic acid alleviates cadmium toxicity in Arabidopsis via suppression of cadmium uptake and translocation. J Integr Plant Biol 62:218–227. https://doi.org/10.1111/jipb.12801
Li H, Xu X, Han K, Wang Z, Ma W, Lin Y, Hua H (2022b) Isolation and functional analysis of OsAOS1 promoter for resistance to Nilaparvata lugens Stål infestation in rice. J Cell Physiol 237(3):1833–1844. https://doi.org/10.1002/jcp.30653
Li L, Zheng Q, Jiang W, Xiao N, Zeng F, Chen G, Mak M, Chen ZH, Deng F (2023) Molecular regulation and evolution of cytokinin signaling in plant abiotic stresses. Plant Cell Physiol 63:1787–1805. https://doi.org/10.1093/pcp/pcac071
Li Y, Huang YZ, Bao QL, Huang YC, Zhang SN (2022a) Effects of exogenous jasmonic acid on arsenic accumulation and response to stress in roots of rice seedlings. Environ Sci 43:4831–4838. https://doi.org/10.13227/j.hjkx.202111296
Lu X, Liu S, Zhi S, Chen J, Ye G (2020) Comparative transcriptome profile analysis of rice varieties with different tolerance to zinc deficiency. Plant Biol (Stuttg). https://doi.org/10.1111/plb.13227
Lu Y, Ye X, Guo R, Huang J, Wang W, Tang J, Tan L, Zhu JK, Chu C, Qian Y (2017) Genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system. Mol Plant 10:1242–1245. https://doi.org/10.1016/j.molp.2017.06.007
Ma JF, Yamaji N, Mitani N, Xu XY, Su YH, McGrath SP, Zhao FJ (2008) Transporters of arsenite in rice and their role in arsenic accumulation in rice grain. Proc Natl Acad Sci U S A 105:9931–9935. https://doi.org/10.1073/pnas.0802361105
Meharg AA, Williams PN, Adomako E, Lawgali YY, Deacon C, Villada A, Cambell RC, Sun G, Zhu YG, Feldmann J, Raab A, Zhao FJ, Islam R, Hossain S, Yanai J (2009) Geographical variation in total and inorganic arsenic content of polished (white) rice. Environ Sci Technol 43:1612–1617. https://doi.org/10.1021/es802612a
Mei C, Qi M, Sheng G, Yang Y (2006) Inducible overexpression of a rice allene oxide synthase gene increases the endogenous jasmonic acid level, PR gene expression, and host resistance to fungal infection. Mol Plant-Microbe Interact 19:1127–1237. https://doi.org/10.1094/MPMI-19-1127
Mousavi SR, Niknejad Y, Fallah H, Tari DB (2020) Methyl jasmonate alleviates arsenic toxicity in rice. Plant Cell Rep 39:1041–1060. https://doi.org/10.1007/s00299-020-02547-7
Nazir F, Fariduddin Q, Khan TA (2020) Hydrogen peroxide as a signalling molecule in plants and its crosstalk with other plant growth regulators under heavy metal stress. Chemosphere 252:126486. https://doi.org/10.1016/j.chemosphere.2020.126486
Ning M, Liu SJ, Deng F, Huang L, Li H, Che J, Yamaji N, Hu F, Lei GJ (2023) A vacuolar transporter plays important roles in zinc and cadmium accumulation in rice grain. New Phytol 239:1919–1934. https://doi.org/10.1111/nph.19070
One Thousand Plant Transcriptomes Initiative (2019) One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574:679–685. https://doi.org/10.1038/s41586-019-1693-2
Pinson SRM, Tarpley L, Yan W, Yeater K, Lahner B, Yakubova E, Huang XY, Zhang M , Guerinot ML , Salt DE (2015) Worldwide genetic diversity for mineral element concentrations in rice grain. Crop Sci 55:294–311. https://doi.org/10.2135/cropsci2013.10.0656
Podgorski J, Berg M (2020) Global threat of arsenic in groundwater. Science 368:845–850. https://doi.org/10.1126/science.aba1510
Ronzan M, Piacentini D, Fattorini L, Federica DR, Caboni E, Eiche E, Ziegler J, Hause B, Riemann M, Betti C, Altamura MM, Falasca G (2019) Auxin-jasmonate crosstalk in Oryza sativa L. root system formation after cadmium and/or arsenic exposure. Environ Exp Bot 165:59–69. https://doi.org/10.1016/j.envexpbot.2019.05.013
Sehar S, Adil MF, Askri SMH, Feng Q, Wei D, Sahito FS, Shamsi IH (2023b) Pan-transcriptomic profiling demarcates serendipita indica-phosphorus mediated tolerance mechanisms in rice exposed to arsenic toxicity. Rice (N Y) 16(1):28. https://doi.org/10.1186/s12284-023-00645-0
Sehar S, Adil MF, Ma Z, Karim MF, Faizan M, Zaidi SSA, Siddiqui MH, Alamri S, Zhou F, Shamsi IH (2023a) Phosphorus and Serendipita indica synergism augments arsenic stress tolerance in rice by regulating secondary metabolism related enzymatic activity and root metabolic patterns. Ecotoxicol Environ Saf 256:114866. https://doi.org/10.1016/j.ecoenv.2023.114866
Sehar S, Feng Q, Adil MF, Sahito FS, Ibrahim Z, Baloch DM, Ullah N, Ouyang Y, Guo Y, Shamsi IH (2022) Tandem application of endophytic fungus Serendipita indica and phosphorus synergistically recuperate arsenic induced stress in rice. Front Plant Sci 13:982668. https://doi.org/10.3389/fpls.2022.982668
Sharma SS, Kumar V, Dietz KJ (2021) Emerging trends in metalloid-dependent signaling in plants. Trends Plant Sci 26:452–471. https://doi.org/10.1016/j.tplants.2020.11.003
Shi S, Wang T, Chen Z, Tang Z, Wu Z, Salt DE, Chao DY, Zhao FJ (2016) OsHAC1;1 and OsHAC1;2 function as arsenate reductases and regulate arsenic accumulation. Plant Physiol 172:1708–1719. https://doi.org/10.1104/pp.16.01332
Smirnoff N, Arnaud D (2019) Hydrogen peroxide metabolism and functions in plants. New Phytol 221:1197–1214. https://doi.org/10.1111/nph.15488
Song WY, Yamaki T, Yamaji N, Ko D, Jung KH, Fujii-Kashino M, An G, Martinoia E, Lee Y, Ma JF (2014) A rice ABC transporter, OsABCC1, reduces arsenic accumulation in the grain. Proc Natl Acad Sci U S A 111:15699–15704. https://doi.org/10.1073/pnas.1414968111
Srivastava S, Srivastava AK, Sablok G, Deshpande TU, Suprasanna P (2015) Transcriptomics profiling of Indian mustard (Brassica juncea) under arsenate stress identifies key candidate genes and regulatory pathways. Front Plant Sci 6:646. https://doi.org/10.3389/fpls.2015.00646
Stumpe M, Bode J, Göbel C, Wichard T, Schaaf A, Frank W, Frank M, Reski R, Pohnert G, Feussner I (2006) Biosynthesis of C9-aldehydes in the moss Physcomitrella patens. Biochim Biophys Acta 1761:301–312. https://doi.org/10.1016/j.bbalip.2006.03.008
Su Y, McGrath S, Zhao F (2010) Rice is more efficient in arsenite uptake and translocation than wheat and barley. Plant Soil 328:27–34. https://doi.org/10.1007/s11104-009-0074-2
Sun SK, Chen Y, Che J, Konishi N, Tang Z, Miller AJ, Ma JF, Zhao FJ (2018) Decreasing arsenic accumulation in rice by overexpressing OsNIP1;1 and OsNIP3;3 through disrupting arsenite radial transport in roots. New Phytol 219:641–653. https://doi.org/10.1111/nph.15190
Tang Z, Chen Y, Miller AJ, Zhao FJ (2019) The C-type ATP-binding cassette transporter OsABCC7 is involved in the root-to-shoot translocation of arsenic in rice. Plant Cell Physiol 60:1525–1535. https://doi.org/10.1093/pcp/pcz054
Tang Z, Zhao F (2021) The roles of membrane transporters in arsenic uptake, translocation and detoxification in plants. Crit Rev Environ Sci Technol 51:2449–2484. https://doi.org/10.1080/10643389.2020.1795053
Verma G, Srivastava D, Narayan S, Shirke PA, Chakrabarty D (2020) Exogenous application of methyl jasmonate alleviates arsenic toxicity by modulating its uptake and translocation in rice (Oryza sativa L.). Ecotoxicol Environ Saf 201:110735. https://doi.org/10.1016/j.ecoenv.2020.110735
Wang J, Song L, Gong X, Xu J, Li M (2020) Functions of jasmonic acid in plant regulation and response to abiotic stress. Int J Mol Sci 21. https://doi.org/10.3390/ijms21041446
Wang P, Xu X, Tang Z, Zhang W, Huang XY, Zhao FJ (2018) OsWRKY28 regulates phosphate and arsenate accumulation, root system architecture and fertility in rice. Front Plant Sci 9:1330. https://doi.org/10.3389/fpls.2018.01330
Wang P, Zhang W, Mao C, Xu G, Zhao FJ (2016) The role of OsPT8 in arsenate uptake and varietal difference in arsenate tolerance in rice. J Exp Bot 67:6051–6059. https://doi.org/10.1093/jxb/erw362
Wasternack C, Feussner I (2018) The oxylipin pathways: biochemistry and function. Annu Rev Plant Biol 69:363–386. https://doi.org/10.1146/annurev-arplant-042817-040440
Xu J, Shi S, Wang L, Tang Z, Lv T, Zhu X, Ding X, Wang Y, Zhao FJ, Wu Z (2017) OsHAC4 is critical for arsenate tolerance and regulates arsenic accumulation in rice. New Phytol 215:1090–1101. https://doi.org/10.1111/nph.14572
Yamaji N, Ma JF (2021) Metalloid transporters and their regulation in plants. Plant Physiol. https://doi.org/10.1093/plphys/kiab326
Ye M, Song Y, Long J, Wang R, Baerson SR, Pan Z, Zhu-Salzman K, Xie J, Cai K, Luo S, Zeng R (2013) Priming of jasmonate-mediated antiherbivore defense responses in rice by silicon. Proc Natl Acad Sci USA 110(38):E3631–E3639. https://doi.org/10.1073/pnas.1305848110
Yu LJ, Luo YF, Liao B, Xie LJ, Chen L, Xiao S, Li JT, Hu SN, Shu WS (2012) Comparative transcriptome analysis of transporters, phytohormone and lipid metabolism pathways in response to arsenic stress in rice (Oryza sativa). New Phytol 195:97–112. https://doi.org/10.1111/j.1469-8137.2012.04154.x
Yu X, Zhang W, Zhang Y, Zhang X, Lang D, Zhang X (2019) The roles of methyl jasmonate to stress in plants. Funct Plant Biol 46:197–212. https://doi.org/10.1071/fp18106
Zhang J, Wysocki R, Li F, Yu M, Martinoia E, Song WY (2023a) Role of ubiquitination in arsenic tolerance in plants. Trends Plant Sci 28:880–892. https://doi.org/10.1016/j.tplants.2023.03.008
Zhang S, Bao Q, Huang Y, Han N (2023b) Exogenous plant hormones alleviate as stress by regulating antioxidant defense system in Oryza sativa L. Environ Sci Pollut Res Int 30:6454–6465. https://doi.org/10.1007/s11356-022-22627-3
Zhang Y, Berman A, Shani E (2023c) Plant hormone transport and localization: signaling molecules on the move. Annu Rev Plant Biol 74:453–479. https://doi.org/10.1146/annurev-arplant-070722-015329
Zhao F, Tang Z, Song J, Huang X, Wang P (2022) Toxic metals and metalloids: uptake, transport, detoxification, phytoremediation and crop improvement for safer food. Mol Plant 15:27–44. https://doi.org/10.1016/j.molp.2021.09.016
Zhao FJ, Ago Y, Mitani N, Li RY, Su YH, Yamaji N, McGrath SP, Ma JF (2010) The role of the rice aquaporin Lsi1 in arsenite efflux from roots. New Phytol 186:392–399. https://doi.org/10.1111/j.1469-8137.2010.03192.x
Zhou Y, Guang YL, Li JW, Wang F, Ahammed GJ, Yang YX (2019) The CYP74 gene family in watermelon: genome-wide identification and expression profiling under hormonal stress and root-knot nematode infection. Agronomy-Basel 9. https://doi.org/10.3390/agronomy9120872
Zvobgo G, Sagonda T, Lwalaba JLW, Mapodzeke JM, Muhammad N, Chen G, Shamsi IH, Zhang G (2018) Transcriptomic comparison of two barley genotypes differing in arsenic tolerance exposed to arsenate and phosphate treatments. Plant Physiol Biochem 130:589–603. https://doi.org/10.1016/j.plaphy.2018.08.006
Acknowledgements
This work was supported by MARA Key Laboratory of Sustainable Crop Production in the Middle Reaches of the Yangtze River, College of Agriculture, Yangtze University, China, and Hawkesbury Institute for the Environment, Western Sydney University, Australia.
Funding
This research was financially supported by the National Natural Science Foundation of China (32170276, 32001456, 32370285), Major International (Regional) Joint Research Project from NSFC-ASRT (32061143044), the Hubei Hongshan Laboratory (2021hskf004), and Yangtze University. ZH-C was funded by Australian Research Council (FT210100366), Grains Research & Development Corporation (WSU2303-001RTX), and Horticulture Innovation Australia (LP18000).
Author information
Authors and Affiliations
Contributions
FD conceived the study. XF, HT, and XC performed all the experiments, analyzed the results with the help of FZ, GC, FD, Z-HC, and YQ. FD wrote the manuscript with support from XF, HT, XC, Z-HC, YQ. Z-HC, YQ and FD conducted the final editing of the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The presented work has been performed in line with principles of the Committee on Publication Ethics (COPE).
Consent for publication
All authors agree to publish.
Competing interests
The authors declare that they have no competing interests.
Additional information
Handling Editor: Dr. Feibo Wu.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Supplemental Table S1.
Primers used in this study.
Additional file 2: Supplementary Fig. 1.
Evolutionary analysis of AOS2 homologues in land plants and algal species. Supplementary Fig. 2. CRISPR/Cas9-induced mutations of OsAOS1. Supplementary Fig. 3. CRISPR/Cas9-induced mutations of OsAOS2. Supplementary Fig. 4. As accumulation and distribution in the seedlings of osaos1–1, osaos2–1, and WT subjected to As(III) for 6 days. Supplementary Fig. 5. As efflux activity of osaos1–1, osaos2–1 and WT. Supplementary Fig. 6. Biomass of osaos1–1, osaos2–1 and WT subjected to 2 or 5 μM As(V) for 6 days. Supplementary Fig. 7. Plant growth and biomass of osaos1–1, osaos2–1 and WT subjected to 2 or 5 μM As(III) for 6 days.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fan, X., Tang, H., Chen, X. et al. Allene oxide synthase 1 contributes to limiting grain arsenic accumulation and seedling detoxification in rice. Stress Biology 3, 52 (2023). https://doi.org/10.1007/s44154-023-00136-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-023-00136-8