Abstract
Potassium (K) is a major essential element in plant cells, and KUP/HAK/KT-type K+ transporters participate in the absorption of K+ into roots and in the long-distance transport to above-ground parts. In Arabidopsis thaliana, KUP9 is involved in the transport of K+ and Cs+ in roots. In this study, we investigated KUP9 function in relation to the K+ status of the plant. The expression of KUP9 was upregulated in older leaves on K+-depleted medium, compared to the expression of the other 12 KUP genes in the KUP/HAK/KT family in Arabidopsis. When grown on low K+ medium, the kup9 mutant had reduced chlorophyll content in seedlings and chlorosis in older rosette leaves. Tissue-specific expression of KUP9 determined by KUP9 promoter:GUS assay depended on the K+ status of the plants: In K+ sufficient medium, KUP9 was expressed in the leaf blade towards the leaf tip, whereas in K+ depleted medium expression was mainly found in the petioles. In accordance with this, K+ accumulated in the roots of kup9 plants. The short-term 43K+ tracer measurement showed that 43K was transferred at a lower rate in roots and shoots of kup9, compared to the wild type. These data show that KUP9 participates in the distribution of K+ in leaves and K+ absorption in roots under low K+ conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potassium, an abundant cation and the major osmolyte in plants, is essential for growth and adaption to various biotic and abiotic stresses. Severe K+ deficiency causes reduction in growth and biomass, disruption of stomatal control, leaf chlorosis and subsequent loss of photosynthesis efficiency. Either as direct consequences of K+ deficiency or due to lack of some K+ dependent enzyme activity, K+-deficient plants are highly vulnerable to drought, salinity, high light and anoxia (Wang et al. 2013; Hasanuzzaman et al. 2018). Uptake of this indispensable nutrient and its translocation throughout the plant is achieved by families of K+ channels and transporters. The Arabidopsis genome contains several K+ specific transporter families, KUP/HAK/KT transporters, Shaker K+ channels (tetrameric channels carrying a voltage sensor domain), cation/proton antiporters (CPAs), and TPK-type K+ channels (Ahn et al. 2004; Gambale and Uozumi 2006; Gierth and Mäser 2007; Sharma et al. 2013; Tsujii et al. 2020). Specifically, the Shaker K+ channel AKT1 and the KT/KUP/HAK family member HAK5 are considered to be the main contributors to K+ import from the soil into root cells (Rubio et al. 2008, 2010; Pyo et al. 2010; Nieves-Cordones et al. 2019).
These two transport systems differ in their K+ transport affinity; HAK5 dominates root K+ uptake under submillimolar concentrations of external K+, whereas AKT1 is the major contributor at higher concentrations. Additionally, HAK5 is essential for K+ retrieval in K+ deprived saline environments because under those conditions, AKT1 promotes K+ leakage from roots (Nieves-Cordones et al. 2010). The transport activity of both HAK5 and AKT1 is regulated via phosphorylation by Calcineurin B-like protein CBL1 and CBL-interacting protein kinase CIPK23 (Xu et al. 2006; Lee et al. 2007; Ragel et al. 2015). This regulation may be related to the modulation of K+ uptake and translocation mediated by AKT1, HAK5 and SKOR (Nieves-Cordones et al. 2019).
The functional roles of Shaker K+ channels have been discussed in detail, such as regulation of stomatal movement, sugar retrieval from the phloem, and pollen tube development (Mouline et al. 2002; Hosy et al. 2003; Dreyer et al. 2017). In contrast, the functional role of KUP/HAK/KT proteins seems to be limited to K+ import and translocation. Escherichia coli Kup was the first member of the KUP/HAK/KT family to be identified, followed by the homologous HAK which was isolated from the eukaryote Schwanniomyces occidentalis (Schleyer and Bakker 1993; María et al. 1995). These findings led to the isolation of homologs from plants (Quintero and Blatt 1997; Santa-maría et al. 1997; Fu and Luan 1998; Kim et al. 1998). Arabidopsis KUP1 and KUP2 were shown to import K+ into E. coli and Arabidopsis suspension cells (Kim et al. 1998). To study the transport activity of KT/KUP/HAK, a K+ uptake-deficient E. coli strain was used as a functional expression system, leading to confirmation of the K+ uptake activities of KUP 4, 5, 6, 7, 10 and 11(Uozumi 2001; Ahn et al. 2004). KUP2, whose expression is regulated by bHLH and WRKY transcription factors, participates in maintaining Na+/K+ homeostasis and confers salt tolerance in plants (Rajappa et al. 2020). Examination of a kup2 kup6 kup8 triple mutant in Arabidopsis suggests that all three KUPs act as K+ effluxers in roots. In addition, KUP6 expression was detected in guard cells and vascular tissues (Osakabe et al. 2013). KUP7 functions as a K+ uptake transporter in planta, based on its ability to transport K+ when expressed in yeast and on the sensitivity of the kup7 mutant to K+ deficiency (Han et al. 2016). Surprisingly, KUP5 has cytosolic adenylate cyclase activity, which is essential for its K+ transport activity (Al-Younis et al. 2018). Recent publications described the significance of KUP8 in the prevention of heavy metal accumulation, and the involvement of KUP4/TRH1 in root hair formation and gravitropism (Sanz-Fernández et al. 2021; Templalexis et al. 2021).
Additionally, KT/KUP/HAK is known to be one of the main transporter families that mediate Cs+ import from the soil into plants, together with members of the cation/H+ exchanger (CHX) family or the cyclic nucleotide gated channel (CNGC) family (White and Broadley 2000; Kanter et al. 2010). Bacterial Kup transporters are different from K+ channels in terms of their tertiary structure (Sato et al. 2014; Tascón et al. 2020). E. coli Kup is permeable to Rb+ and Cs+ in addition to K+ and these ions can support cell proliferation during K+ deficiency (Tanudjaja et al. 2017). In contrast, in plants, no beneficial effects of Cs+ uptake have been reported, even though Cs+ also belongs to the Group I alkali metals and has similar chemical properties as K+ (Nieves-Cordones et al. 2017; Rai et al. 2017). Rather, Cs+ accumulation can cause severe growth retardation due to nutritional competition with K+ and blockage of certain K+ channels by Cs+. A number of studies have implicated HAK5 as the major contributor to Cs+ uptake into plants (Qi et al. 2008; Adams et al. 2013, 2019a; Alemán et al. 2014; Genies et al. 2017). In contrast, AKT1 is blocked by external Cs+, which leads to leakage of K+ from the roots under low K+ conditions and subsequently growth inhibition due to K+ deficiency (Adams et al. 2019a).
Recent reports provide evidence for a role of KUP9 in K+ acquisition, auxin homeostasis, IAA metabolism in root tips and negative regulation of Cs+ accumulation (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). The K+ and Cs+ uptake ability of KUP9 was previously established by E. coli assay (Kobayashi et al. 2010). But while KUP9 expression is induced in response to low K+ condition, its subcellular localization is restricted to the ER (Genies et al. 2021). Thus, whether KUP9 directly mediates K+ import and Cs+ extrusion in vivo or affects these processes indirectly through another transport system in the plasma membrane remains unknown.
This study focused on the expression and function of Arabidopsis KUP9 during K+ deficiency and provides insight into additional physiological processes involved in regulation of KUP9 expression. We observed KUP9 expression in shoots and studied how loss of KUP9 activity in kup9 mutant leads to chlorosis in rosette leaves under K+-depleted conditions. We therefore measured the expression of KUP9 and K+ content and performed imaging of the short-term uptake of the radioisotope 43K+ in kup9 at different K+ concentrations. KUP9 contributes to the distribution of K+ in shoots as well as in roots under low K+ conditions.
Results
Upregulation of kup9 under low K+ conditions
While the expression of KUP/HAK/KT genes under sufficient K+ conditions has been previously determined in Arabidopsis (Ahn et al. 2004), changes in their expression upon K+ depletion have not been studied in detail. We isolated total RNA from younger and older leaves and from roots of Col-0 plants grown for 2 weeks on media containing either 2 mM K+ (sufficient) or 100 μM K+ (low). This study was conducted with media containing NH4+, these conditions inhibit some K+ transport systems including HAK5 (Santa-María et al. 2018). RT-qPCR was performed to determine gene expression of members of the KUP/HAK/KT family with or without K+ starvation (Fig. 1). As previously reported, HAK5 expression was strongly induced in older leaves and roots in response to K+ starvation (Ahn et al. 2004). Among the KUP members, KUP9 (and to a lesser degree KUP12) were upregulated in older leaves, while KUP2 and KUP5 expression was reduced in both leaves and roots.
Relative expression of KUP/HAK/KT transporters. Plants were grown for 2 weeks in medium containing 2 mM or 100 μM K+. Plants were at the same stage when they were dissected into the four youngest leaves (YL), the remaining older leaves (OL) and roots. RT-qPCR was performed to determine the relative levels of the KUP/HAK/KT transcripts, using ubiquitin 10 (UBQ10) as the reference gene
Phenotype of kup9 under low K+ conditions
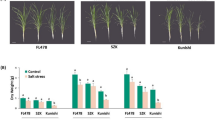
Next, we investigated the effects of low K+ conditions on the phenotypes of KT/KUP/HAK loss-of-function mutants. We obtained homozygous SALK lines for all members of the family, except kup6 and kup11 (no lines were available). All kup mutant lines were sown on agar plates containing 2 mM K+ or 100 μM K+, and after 2 weeks of growth the chlorophyll content was determined (Fig. 2). Under low K+ conditions, chlorophyll content was reduced in all lines including the wild type. However, both kup9 and kup12 showed a stronger decrease in chlorophyll content than the wild type, and kup9 was the most affected (Fig. 2A and B). In older, hydroponically grown plants, chlorosis was clearly visible at the edges of older rosette leaves of kup9 at K+ concentrations at and below 300 μM, although the overall size of the kup9 plants was similar to that of the wild type (Fig. 2C). In kup12 and hak5 plants, chlorosis at the leaf edges was also observed at or below 200 μM K+. These results highlight the contribution of KUP9 in older leaves under K+ deficiency.
Phenotypes of kup/hak/kt mutants under low K+ and K+ sufficient conditions. A Photograph of two-week-old plants grown on medium containing 2 mM (top) or 100 μM K+ (bottom). B Chlorophyll content of the shoot tissue of the same plants. Data are shown as mean ± S.D. of n = 4 independent plants. *P < 0.05, ns, not significant (P > 0.05); one-way ANOVA. Scale bar = 1 cm C Phenotype during reproductive growth with different amounts of KCl. Representative images of 63-day-old wild-type, kup9 kup12 and hak5 plants, grown for the last 30 days on hydroponic medium containing 0.1, 0.2 or 0.3 mM KCl
Change of KUP9 expression pattern in response to external K+ concentrations
To understand the reason for the low-K+ sensitivity of kup9, we examined the expression pattern of KUP9 using Col-0 plants transformed with a KUP9 promoter-GUS construct. Since KUP9 expression in young seedlings had been determined previously (Zhang et al. 2020; Genies et al. 2021), we performed GUS staining on plants grown for two weeks with either low or sufficient K+ (Fig. 3). With sufficient K+ supply, KUP9 expression was observed mainly in the maturation region of the primary root (except for very end of the root tip which was stained under both K+ conditions) and in leaves not including petioles. In roots, KUP9 was mainly expressed in the vasculature (Fig. 3M and m). In contrast, when plants were grown on low K+, the expression pattern of KUP9 shifted to lateral root tips and regions closer to the end of the primary root. The very end of the root tip which was stained under both K+ conditions (Zhang et al. 2020; Genies et al. 2021). In leaves, KUP9 expression shifted to petioles and regions close to them (Fig. 3K, L, k, and l). These results suggested a role of KUP9 in modulating K+ translocation in leaves and roots in response to K+ depletion.
GUS reporter assay of KUP9 promoter activity in Arabidopsis. GUS staining of two-week-old seedlings containing a KUP9 promoter-GUS construct grown in medium containing 2 mM K+ (A-M) or l00 μM K+ (a-m). Whole plant without leaves (A, a), 5th or 6th leaf (B, C, b and c), 3rd or 4th leaf (D, E, d and e), 1st or 2nd leaf (F, G, f and g), petioles (H, h), lateral and primary roots (I, i), and root tip (J, j). Cross-sections of petioles (K, L, k and l) and roots (M and m). L and l are enlarged from K and k. Scale bars represent 1 mm in (A-G and a-g), 0.5 mm in (H-J and h-j), 100 μm in (K, and k) and 20 μm in (L and M, l and m)
Distribution of K+ in kup9 and hak5 plants
To investigate the role of KUP9 in root uptake and translocation of K+, we first compared K+, Ca2+ and Mg2+ content of young leaves, old leaves, petioles and stems, and roots of kup9 and wild type (Fig. 4). As a K+ uptake deficient control, hak5 was used. Under K+-sufficient conditions (2 mM), there were no differences in any of the tissues for all three ions, K+, Ca2+, and Mg2+. In contrast, under low K+ conditions (100 μM), K+ concentrations in the roots of kup9 were elevated, suggesting that KUP9 contributed to K+ translocation from the roots to the shoots. This was supported by the increased expression of KUP9 in petioles under low K+ conditions (Fig. 3). Next, we examined whether there were differences in the short-term K+ uptake rate and root-to-shoot translocation speed between wild-type, kup9 and hak5 plants by monitoring the uptake of short-lived radio isotopic potassium, 43K (Fig. 5). Significant differences were observed between the wild type and kup9 and hak5 plants that had been grown under low K+ conditions but not between wild type and mutants grown with sufficient K+ supply (Fig. 5). In both shoots and roots, K+-deficient kup9 showed less 43K accumulation than the wild type. Similar results were obtained for hak5 shoots and roots. These data suggest that under K+ limited conditions KUP9 supports K+ uptake in the roots as well as K+ translocation to the shoots.
K+ accumulation in kup9 and hak5. Plants were grown on synthetic agar medium containing 2 mM K+ or l00 μM K+ for 2 weeks. K+, Ca2+ and Mg2+ contents of young leaves (four youngest leaves), old leaves (remaining leaves), petioles and stems, and roots of wild-type, kup9 and hak5 plants were measured by ICP-OES. Data are shown as mean ± S.D. of n = 8 independent samples (four plants in each sample). P < 0.05, ns, not significant (P > 0.05); one-way ANOVA
Distribution of 43K in kup9 and hak5. A Representative autoradiography of the excised shoots (S), hypocotyls (H) and roots (R) of wild-type, kup9 and hak5 plants after uptake of 43K. Plants were grown on synthetic agar medium containing 2 mM K+ (left) or l00 μM K+ (right) for 2 weeks before being incubated for 1 h in hydroponic medium containing 43K. B Quantification 43K-uptake into shoots and roots of the plants shown in panel (A). Data are shown as mean ± S.D. of n = 5 independent plants. P < 0.05, ns, not significant (P > 0.05); one-way ANOVA. Scale bars represent 1 cm
Discussion
This study revealed the contribution of KUP9 to K+ distribution in leaves and roots in response to K+ concentration changes. The expression level of KUP9 increased in older leaves under low K+ conditions (Fig. 1) and the expression pattern of KUP9 changed in leaves and roots in response to the external K+ concentrations (Fig. 3). This is consistent with the observed chlorosis in older rosette leaves in kup9 under low K+ conditions (Figs. 2 and 3). K+ accumulated in kup9 roots (Fig. 4) and the short-term K+ uptake rate into tissues of kup9 was reduced, similar to the reduction observed for hak5 (Fig. 5). Previous reports had focused on the function and role of KUP9 as a K+ and Cs+ transporter in roots (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). Our study suggests that KUP9 was involved in K+ translocation from roots to shoots and K+ uptake in roots (Figs. 4 and 5).
KUP/HAK/KT transporters are distinct from K+ channels and Trk/Ktr/HKT K+ transporters, which share a similar ion conductance profile at their pore regions (Sato et al. 2014). Disruption of Kup in E. coli inhibits cell growth, which indicates that K+ uptake is the primary role of E. coli Kup (Tanudjaja et al. 2017). In addition, E. coli Kup transports Rb+ and Cs+, which allows the cells to compensate for K+ starvation (Tanudjaja et al. 2017). Arabidopsis KUP9 also transports Cs+, similar to other KUP/HAK/KT transporters in Arabidopsis (Qi et al. 2008; Kobayashi et al. 2010; Adams et al. 2013). (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021) Moreover, KUP9 also functions as an auxin transporter in the ER membrane in Arabidopsis (Zhang et al. 2020). Our study showed no significant differences in K+ content between roots, young and old leaves between the wild type and kup9, on sufficient K+ media (Fig. 4), which matches previously reported results obtained under similar conditions (Zhang et al. 2020; Genies et al. 2021). In the roots, a significant increase in K+ content was found in kup9. This result suggests that KUP9 is important for K+ translocation from root to shoot (Fig. 4). In contrast, short-term 43K absorption was lower in kup9 than in the wild type (Fig. 5). A similar reduction was found for the 86Rb uptake rate in hak5 (Gierth et al. 2005). The increase in KUP9 expression in response to K+ deficiency also resembled that of HAK5 in roots.
The results of the KUP9 promoter-GUS assays (Fig. 3) indicated that the localization of KUP9 expression in roots and leaves changed in response to the overall K+ status of the plant. K+-deficiency induced expression of KUP9 in lateral roots and in regions proximal to the tip of the main root, which might improve K+ absorption from the substrate (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). In leaves, KUP9 expression shifted from the leaf blade to the petiole and areas close to it. Consistent with this, chlorosis occurred at the leaf edges, indicating that those areas were K+-deficient (Fig. 3). Leaf chlorosis is a typical symptom of severe K+ deficiency. Under K+-deficient conditions plants typically redirect the transport of K+ from fully expanded leaves or other parts of the shoot towards younger leaves to ensure the supply of these growing tissues (Osmolovskaya et al. 2020). The observed changes in KUP9 expression in response to K+ deficiency are consistent with the hypothesis that KUP9 plays a role in K+ translocation in the shoot. The kup9 mutant was more sensitive to low K+-conditions than hak5 (Fig. 2B and C). Expression of HAK5 was most highly induced in roots under low K+ conditions (100 μM) (Fig. 1), while KUP9 expression under low K+ conditions was induced in leaves, which indicates that KUP9 had functions beyond root K+ uptake (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021).
KUP9 is located in the ER membrane in roots (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). How does K+ move from roots to shoots through KUP9? Recently, KUP9 has been implicated in Cs+ efflux from root cells to the apoplast (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). This makes sense if KUP9 is present in the plasma membrane. It is possible that KUP9 changes its intracellular localization depending on the tissue and environmental conditions. Such dual localization exists for KUP4, which localizes to the ER as well as the plasma membrane in certain areas of the root (Rigas et al. 2013; Templalexis et al. 2021). One way to achieve this would be by posttranslational modification. For example, interaction of the Shaker K+ channel AKT2 with a pair of proteins in the cytosol, calcineurin B-like protein 4 (CBL4) and CBL-interacting protein kinase 6 (CIPK6), promotes the translocation of AKT2 from ER to plasma membrane (Held et al. 2011). This recruitment of AKT2 is dependent on dual lipid modification of CBL4, a common feature among Ca2+ dependent phosphorylation modules (Saito et al. 2018). An example of regulation of KUP/HAK/KT transporters by Ca2+ dependent phosphorylation is the activation of HAK5-mediated K+ uptake via phosphorylation by CIPK23 interacting with CBL1, 8, 9, and 10 (Ragel et al. 2015). It is therefore reasonable to assume that KUP9 could also be regulated by a similar Ca2+ dependent phosphorylation system. However, the subcellular localization of KUP9 in shoots has not been determined, and further study will be required to understand the regulation of KUP9 trafficking in leaves.
Data accumulated by several groups have shown that KUP9 transports K+ from the ER to the cytoplasm and also promotes Cs+ efflux in roots (Adams et al. 2019b; Zhang et al. 2020; Genies et al. 2021). Our study adds new information by showing an involvement of KUP9 in K+ transport to the shoots. Together these findings suggest that KUP9 participates in K+ and Cs+ transport and distribution in plants in cooperation with other transport systems such as HAK5.
Material and methods
Plant material and growth
Sterilized seeds were vernalized for 48 h at 4 °C in 1.5-mL centrifuge tubes. Plants were grown on agar medium containing 2 mM or 100 μM K+ at 22 °C under 16 h light/8 h dark conditions for 2 weeks. Light intensity was adjusted to around 100 μmolm− 2 s− 1. The composition of hydroponic medium was 1.25 mM NH4H2PO4, 3 mM Ca(NO3)2, 1.5 mM MgSO4, 100 μM Fe3-EDTA, 70 μM H3BO3, 20 μM MnCl2・4H2O, 0.3 μM CuSO4・5H2O, 1 μM ZnSO4・7H2O, 0.2 μM K2MoO4, 0.1 μM CoCl2・6H2O, 0.1% (v/v), Gamborg’s Vitamin Solution (SIGMA-ALDRICH, USA). The pH of the medium was adjusted to pH 5.8 with KOH and then adjusted to 2 mM or 100 μM K+ with added KCl. The synthetic agar medium contained the same components as the hydroponic medium plus 0.8% agar and 0.1% sucrose. To create K-limited conditions during reproductive growth, plants were first grown on 1/2 MS agar medium for 25 d and then transferred to liquid medium (2 mM MgSO4・7H2O, 2 mM NaH2PO4, 2 mM NH4NO3, 2 mM Ca(NO3)2, 0.05 mM Fe(III)EDTA and 2 μM KI) containing 2 mM KCl for 8 d to acclimate to hydroponic culture. Then, plants were transferred to liquid medium containing 0.1, 0.2 or 0.3 mM KCl and grown for 30 d at 23 °C under 8 h light/16 h dark conditions. When bolting occurred, the stems were removed to encourage the growth of leaves. The Arabidopsis Columbia ecotype (Col-0) and T-DNA insertion lines including kup1 (SALK_051343), kup2 (SALK_097636), kup3 (SALK_002622), kup4 (SALK_043791), kup5 (SALK_072850), kup7 (SALK_206158), kup8 (SALK_041357), kup9 (SALK_108080), kup10 (SALK_072956), kup12 (SALK_083613) and hak5 (SALK_130604) were obtained from the Arabidopsis Biological Resource Center (ABRC).
qPCR analysis
Total RNA was extracted from the younger four leaves (including meristematic tissue), the rest of older the leaves, and roots of 2-week-old plants using TRI REAGENT (Molecular Research Center, lnc). ReverTra Ace® qPCR RT Master Mix with gDNA Remover (TOYOBO CO., LTD) was used for genomic DNA removal and cDNA synthesis. The reverse transcription reaction protocol consisted of 37 °C for 15 min, 50 °C for 5 min, and 98 °C for 5 min. qPCR was performed using a StepOnePlus Real-Time PCR System (Applied Biosystems, USA). Expression level is shown as values relative to the expression level of the reference gene of Ubiquitin 10 (UBQ10) (Tong et al. 2009). For each gene, the relative expression under various conditions (2 mM K+ and 100 μM K+) and in different tissues (the four youngest leaves, the remaining older leaves and roots) was then calculated in relation to its expression level in younger leaves with 2 mM K+.
Chlorophyll content
Shoot tissue was sampled in 1.5 ml tubes and fresh weight was determined. 80% (v/v) of cold acetone solution was added and left overnight at 4 °C in dark. Samples were then centrifuged at 4 °C, 15,000 rpm for 15 minutes, and absorbance of the supernatant solution was measured at 646 nm, 663 nm, and 750 nm with a double-beam spectrophotometer (HITACHI UH5300). The amount of chlorophyll in the aboveground tissue was determined from the concentration of chlorophyll in the solution, which was calculated from the following formula using the measured values.
GUS staining
Arabidopsis kup9 promoter sequence (2864 bp) was amplified by PCR using a pair of primers, 5′-TGATTACGCCAAGCTTCTGCATCATCAAACAGAG-3′ and 5′- AGGGACTGACCACCCGGGTTTTGTAACAAAAGAACT-3′ and cloned into HindIII and Smal-digested pBI101 (provided by Kenzo Nakamura) using the In-Fusion® HD cloning kit (Takara Bio USA, Inc.) (Ohta et al. 1990). The final construct consisting of the β-glucuronidase (GUS) gene under control of the AtKUP9 promoter (ProAtKUP9:GUS) was introduced into Arabidopsis thaliana Col-0 via Agrobacterium-mediated transformation. Seeds of Arabidopsis thaliana expressing GUS gene were grown on solid medium containing 2 mM or 100 μM K+ for two weeks. The ProAtKUP9:GUS plants were analyzed for GUS expression according to standard procedures (Jefferson et al. 1987). For each treatment whole plants were incubated in staining solution containing 0.5 mM X-Gluc (5-bromo-4-chloro-3-indolyl-β-D-glucuronide) at 37 °C for 3–4 h and destained in 70% ethanol. To prepare tissue sections, GUS-stained plants were dehydrated with 80%, 90% and 100% ethanol for one day each, and then fixed in Technovit 7100 (Heraeus Kulzer, Tokyo, Japan). The tissue was sliced into 6 μm sections using a Leica RM2145 microtome (Leica, Wetzlar, Germany). Sections were stained with 0.01% neutral red (FUJIFILM Wako Chemicals, Japan) for 1 min and then rinsed twice with water. After drying the samples, they were covered with Entellan® new (FUJIFILM Wako Chemicals, Japan) and were examined under a microscope.
K+ content determination
Plants grown for 2 weeks with 2 mM or 100 μM K+ were divided into the younger four leaves (including meristematic tissue), the remaining older leaves, the petioles and stems, and roots. Plant materials from four individual plants were placed into 1.5 ml tubes and dried at 60 °C for at least 24 h. Then 400 μl of nitric acid was added to each sample, and were left to digest overnight or longer. The nitric acid extracts were diluted 50-fold with MiliQ water and the amount of K+, Ca2+, and Mg2+ was determined with an inductively coupled plasma optical emission spectrometer; ICP-OES (iCAP 6500; Thermo Fisher Scientific, USA).
Production of 43K+
Potassium-43 (43K+) was produced in the natCa(γ,pxn) reactions using an electron linear accelerator at the Research Center for Electron Photon Science at Tohoku University. The calcium oxide was sealed in quartz tubes and irradiated with bremsstrahlung for several hours under water cooling. Produced 43K+ was separated from the calcium oxide by an oxalate precipitation method and purified by cation exchange chromatography. The 43K+ tracer was dried and dissolved in hydroponic medium containing 2 mM K+. For the 43K+ absorption experiments, plants were grown on 2 mM or 100 μM K+ synthetic agar medium without tracer for 2 weeks. Then plants were incubated in hydroponic medium (10 ml) supplemented with 2 MBq of 43K+ for 1 h and potassium absorption was determined.
Image analysis
After 1 h, the plants were washed twice with hydroponic medium containing 2 mM K+, and dissected with surgical scissors into shoots, hypocotyls and roots. The plant material was put onto imaging plates (Fuji Film, Japan) and exposed for 3–5 h at 4 °C. The imaging plates were scanned in a Typhoon FLA 9500 laser scanner (GE Healthcare Japan K.K., Japan). The plant material was then transferred into individual tubes and the weight was determined.
Quantitative analysis by γ-ray spectrometry
After the image analysis, the plant parts were analyzed for the amount of 43K radioactivity via g-spectrometry with a germanium semiconductor detector of 35% relative efficiency. For measurements, samples were placed at 2 mm from the surface of the detector, which was housed in a lead-shield box. The radioactivity of the 43K was determined from the peak area at 373 keV.
Abbreviations
- HAK/KUP/KT:
-
High-Affinity K+ Transporter/K+ Uptake Permease/K+ Transporter
- CPAs:
-
Cation/Proton Antiporters
- CBL:
-
Calcineurin B-Like
- CIPK:
-
CBL-Interacting Protein Kinase
- CHX:
-
Cation/H+ Exchanger
- CNGC:
-
Cyclic Nucleotide Gated Channel
- IAA:
-
Indole-3-Acetic Acid
- ER:
-
Endoplasmic Reticulum
- GUS:
-
β-Glucuronidase
- UBQ:
-
Ubiquitin
- qPCR:
-
Quantitative PCR
References
Adams E, Abdollahi P, Shin R (2013) Cesium inhibits plant growth through jasmonate signaling in Arabidopsis thaliana. Int J Mol Sci 14:4545–4559. https://doi.org/10.3390/ijms14034545
Adams E, Miyazaki T, Saito S, Uozumi N, Shin R (2019a) Cesium inhibits plant growth primarily through reduction of potassium influx and accumulation in Arabidopsis. Plant Cell Physiol 60:63–76. https://doi.org/10.1093/pcp/pcy188
Adams E, Miyazaki T, Shin R (2019b) Contribution of KUPs to potassium and cesium accumulation appears complementary in Arabidopsis. Plant Signal Behav 14:1–3. https://doi.org/10.1080/15592324.2018.1554468
Ahn SJ, Shin R, Schachtman DP (2004) Expression of KT/KUP genes in Arabidopsis and the role of root hairs in K+ uptake. Plant Physiol 134:1135–1145. https://doi.org/10.1104/pp.103.034660
Alemán F, Caballero F, Ródenas R, Rivero RM, Martínez V, Rubio F (2014) The F130S point mutation in the Arabidopsis high-affinity K+ transporter AtHAK5 increases K+ over Na+ and Cs+ selectivity and confers Na+ and Cs+ tolerance to yeast under heterologous expression. Front Plant Sci 5:1–11. https://doi.org/10.3389/fpls.2014.00430
Al-Younis I, Wong A, Lemtiri-Chlieh F, Schmöckel S, Tester M, Gehring C et al (2018) The Arabidopsis thaliana K+ −uptake permease 5 (AtKUP5) contains a functional cytosolic adenylate cyclase essential for K+ transport. Front Plant Sci 871:1–15. https://doi.org/10.3389/fpls.2018.01645
Dreyer I, Gomez-porras JL, Riedelsberger J (2017) The potassium battery: a mobile energy source for transport processes in plant vascular tissues. New Phytol 216:1049–1053. https://doi.org/10.1111/nph.14667
Fu HH, Luan S (1998) AtKUP1: a dual-affinity K+ transporter from Arabidopsis. Plant Cell 10:63–73. https://doi.org/10.1105/tpc.10.1.63
Gambale F, Uozumi N (2006) Properties of shaker-type potassium channels in higher plants. J Membr Biol 210:1–19. https://doi.org/10.1007/s00232-006-0856-x
Genies L, Orjollet D, Carasco L, Camilleri V, Frelon S, Vavasseur A et al (2017) Uptake and translocation of cesium by Arabidopsis thaliana in hydroponics conditions: links between kinetics and molecular mechanisms. Environ Exp Bot 138:164–172. https://doi.org/10.1016/j.envexpbot.2017.03.013
Genies L, Martin L, Kanno S, Chiarenza S, Carasco L, Camilleri V et al (2021) Disruption of AtHAK/KT/KUP9 enhances plant cesium accumulation under low potassium supply. Physiol Plant 173:1230–1243. https://doi.org/10.1111/ppl.13518
Gierth M, Mäser P (2007) Potassium transporters in plants - involvement in K+ acquisition, redistribution and homeostasis. FEBS Lett 581:2348–2356. https://doi.org/10.1016/j.febslet.2007.03.035
Gierth M, Mäser P, Schroeder JI (2005) The potassium transporter AtHAK5 functions in K+ deprivation-induced high-affinity K+ uptake and AKT1 K+ channel contribution to K+ uptake kinetics in Arabidopsis roots. Plant Physiol 137:1105–1114. https://doi.org/10.1104/pp.104.057216
Han M, Wu W, Wu WH, Wang Y (2016) Potassium transporter KUP7 is involved in K+ acquisition and translocation in arabidopsis rroot under K+-limited conditions. Mol Plant 9:437–446. https://doi.org/10.1016/j.molp.2016.01.012
Hasanuzzaman M, Bhuyan MHMB, Nahar K, Hossain MS, Mahmud JA, Hossen MS et al (2018) Potassium: a vital regulator of plant responses and tolerance to abiotic stresses. Agronomy 8:31. https://doi.org/10.3390/agronomy8030031
Held K, Pascaud F, Eckert C, Gajdanowicz P, Hashimoto K, Corratgé-Faillie C et al (2011) Calcium-dependent modulation and plasma membrane targeting of the AKT2 potassium channel by the CBL4/CIPK6 calcium sensor/protein kinase complex. Cell Res 21:1116–1130. https://doi.org/10.1038/cr.2011.50
Hosy E, Vavasseur A, Mouline K, Dreyer I, Gaymard F, Porée F et al (2003) The Arabidopsis outward K+ channel GORK is involved in regulation of stomatal movements and plant transpiration. Proc Natl Acad Sci U S A 100:5549–5554. https://doi.org/10.1073/pnas.0733970100
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907. https://doi.org/10.1002/j.1460-2075.1987.tb02730.x
Kanter U, Hauser A, Michalke B, Dräxl S, Schäffner AR (2010) Caesium and strontium accumulation in shoots of Arabidopsis thaliana: genetic and physiological aspects. J Exp Bot 61:3995–4009. https://doi.org/10.1093/jxb/erq213
Kim EJ, Kwak JM, Uozumi N, Schroeder JI (1998) AtKUP1: an Arabidopsis gene encoding high-affinity potassium transport activity. Plant Cell 10:51–62. https://doi.org/10.1105/tpc.10.1.51
Kobayashi D, Uozumi N, Hisamatsu S, Yamagami M (2010) AtKUP/HAK/KT9, a K+ transporter from Arabidopsis thaliana, mediates Cs+ uptake in Escherichia coli. Biosci Biotechnol Biochem 74:203–205. https://doi.org/10.1271/bbb.90638
Lee SC, Lan WZ, Kim BG, Li L, Cheong YH, Pandey GK et al (2007) A protein phosphorylation/dephosphorylation network regulates a plant potassium channel. Proc Natl Acad Sci U S A 104:15959–15964. https://doi.org/10.1073/pnas.0707912104
María AB, Ronald DK, Susan JAB, Alonso RN (1995) A potassium transporter of the yeast Schwanniomyces occidentalis homologous to the Kup system of Escherichia coli has a high concentrative capacity. EMBO J 14:3021–3027. https://doi.org/10.1002/j.1460-2075.1995.tb07304.x
Mouline K, Véry AA, Gaymard F, Boucherez J, Pilot G, Devic M et al (2002) Pollen tube development and competitive ability are impaired by disruption of a shaker K+ channel in Arabidopsis. Genes Dev 16:339–350. https://doi.org/10.1101/gad.213902
Nieves-Cordones M, Alemán F, Martínez V, Rubio F (2010) The Arabidopsis thaliana HAK5 K+ transporter is required for plant growth and K+ acquisition from low K+ solutions under saline conditions. Mol Plant 3:326–333. https://doi.org/10.1093/mp/ssp102
Nieves-Cordones M, Mohamed S, Tanoi K, Kobayashi NI, Takagi K, Vernet A et al (2017) Production of low-Cs+ rice plants by inactivation of the K+ transporter OsHAK1 with the CRISPR-Cas system. Plant J 92:43–56. https://doi.org/10.1111/tpj.13632
Nieves-Cordones M, Lara A, Ródenas R, Amo J, Rivero RM, Martínez V et al (2019) Modulation of K+ translocation by AKT1 and AtHAK5 in Arabidopsis plants. Plant Cell Environ 42:2357–2371. https://doi.org/10.1111/pce.13573
Ohta S, Mita S, Hattori T, Nakamura K (1990) Construction and expression in tobacco of a β-glucuronidase (GUS) reporter gene containing an intron within the coding sequence. Plant Cell Physiol 31:805–813. https://doi.org/10.1093/oxfordjournals.pcp.a077982
Osakabe Y, Arinaga N, Umezawa T, Katsura S, Nagamachi K, Tanaka H et al (2013) Osmotic stress responses and plant growth controlled by potassium transporters in Arabidopsis. Plant Cell 25:609–624. https://doi.org/10.1105/tpc.112.105700
Osmolovskaya N, Shumilina J, Bureiko K, Chantseva V, Bilova T, Kuchaeva L et al (2020) Ion homeostasis response to nutrient-deficiency stress in plants. cell growth 61. https://doi.org/10.5772/intechopen.89398. https://www.intechopen.com/chapters/69690
Pyo YJ, Gierth M, Schroeder JI, Cho MH (2010) High-affinity K+ transport in Arabidopsis: AtHAK5 and AKT1 are vital for seedling establishment and post germination growth under low-potassium conditions. Plant Physiol 153:863–875 https://doi.org/pp.110.154369
Qi Z, Hampton CR, Shin R, Barkla BJ, White PJ, Schachtman DP (2008) The high affinity K+ transporter AtHAK5 plays a physiological role in planta at very low K+ concentrations and provides a caesium uptake pathway in Arabidopsis. J Exp Bot 59:595–607. https://doi.org/10.1093/jxb/erm330
Quintero FJ, Blatt MR (1997) A new family of K+ transporters from Arabidopsis that are conserved across phyla. FEBS Lett 415:206–211. https://doi.org/10.1016/S0014-5793(97)01125-3
Ragel P, Ródenas R, García-Martín E, Andrés Z, Villalta I, Nieves-Cordones M et al (2015) CIPK23 regulates HAK5-mediated high-affinity K+ uptake in Arabidopsis roots. Plant Physiol 169:01401. https://doi.org/10.1104/pp.15.01401
Rai H, Yokoyama S, Satoh-Nagasawa N, Furukawa J, Nomi T, Ito Y et al (2017) Cesium uptake by rice roots largely depends upon a single gene, HAK1, which encodes a potassium transporter. Plant Cell Physiol 58:1486–1493. https://doi.org/10.1093/pcp/pcx094
Rajappa S, Krishnamurthy P, Kumar PP (2020) Regulation of AtKUP2 expression by bHLH and WRKY transcription factors helps to confer increased salt tolerance to Arabidopsis thaliana plants. Front Plant Sci 11:1–16. https://doi.org/10.3389/fpls.2020.01311
Rigas S, Ditengou FA, Ljung K, Daras G, Tietz O, Palme K et al (2013) Root gravitropism and root hair development constitute coupled developmental responses regulated by auxin homeostasis in the Arabidopsis root apex. New Phytol 197:1130–1141. https://doi.org/10.1111/nph.12092
Rubio F, Nieves-Cordones M, Alemán F, Martínez V (2008) Relative contribution of AtHAK5 and AtAKT1 to K+ uptake in the high-affinity range of concentrations. Physiol Plant 134:598–608. https://doi.org/10.1111/j.1399-3054.2008.01168.x
Rubio F, Alemán F, Nieves-Cordones M, Martínez V (2010) Studies on Arabidopsis athak5, atakt1 double mutants disclose the range of concentrations at which AtHAK5, AtAKT1 and unknown systems mediate K+ uptake. Physiol Plant 139:220–228. https://doi.org/10.1111/j.1399-3054.2010.01354.x
Saito S, Hamamoto S, Moriya K, Matsuura A, Sato Y, Muto J et al (2018) N-myristoylation and S-acylation are common modifications of Ca2+-regulated Arabidopsis kinases and are required for activation of the SLAC1 anion channel. New Phytol 218:1504–1521. https://doi.org/10.1111/nph.15053
Santa-maría GE, Rubio F, Dubcovsky J, Rodríguez-navarroa A (1997) The HAK1 gene of barley is a member of a large gene family and encodes a high-affinity potassium transporter. Plant Cell 9:2281–2289. https://doi.org/10.1105/tpc.9.12.2281
Santa-María GE, Oliferuk S, Moriconi JI (2018) KT-HAK-KUP transporters in major terrestrial photosynthetic organisms: a twenty years tale. J Plant Physiol 226:77–90. https://doi.org/10.1016/j.jplph.2018.04.008
Sanz-Fernández M, Rodríguez-González A, Sandalio LM, Romero-Puertas MC (2021) Role of potassium transporter KUP8 in plant responses to heavy metals. Physiol Plant 173:180–190. https://doi.org/10.1111/ppl.13345
Sato Y, Nanatani K, Hamamoto S, Shimizu M, Takahashi M, Tabuchi-Kobayashi M et al (2014) Defining membrane spanning domains and crucial membrane-localized acidic amino acid residues for K+ transport of a Kup/HAK/KT-type Escherichia coli potassium transporter. J Biochem 155:315–323. https://doi.org/10.1093/jb/mvu007
Schleyer M, Bakker EP (1993) Nucleotide sequence and 3′-end deletion studies indicate that the K+- uptake protein kup from Escherichia coli is composed of a hydrophobic core linked to a large and partially essential hydrophilic C terminus. J Bacteriol 175:6925–6931. https://doi.org/10.1128/jb.175.21.6925-6931.1993
Sharma T, Dreyer I, Riedelsberger J (2013) The role of K(+) channels in uptake and redistribution of potassium in the model plant Arabidopsis thaliana. Front Plant Sci 4:224. https://doi.org/10.3389/fpls.2013.00224
Tanudjaja E, Hoshi N, Su YH, Hamamoto S, Uozumi N (2017) Kup-mediated Cs+ uptake and Kdp-driven K+ uptake coordinate to promote cell growth during excess Cs+ conditions in Escherichia coli. Sci Rep 7:1–9. https://doi.org/10.1038/s41598-017-02164-7
Tascón I, Sousa JS, Corey RA, Mills DJ, Griwatz D, Aumüller N et al (2020) Structural basis of proton-coupled potassium transport in the KUP family. Nat Commun 11:1–10. https://doi.org/10.1038/s41467-020-14441-7
Templalexis D, Tsitsekian D, Liu C, Daras G, Šimura J, Moschou P et al (2021) Potassium transporter TRH1/KUP4 contributes to distinct auxin-mediated root system architecture responses. Plant Physiol 188:1043–1060. https://doi.org/10.1093/plphys/kiab472
Tong Z, Gao Z, Wang F, Zhou J, Zhang Z (2009) Selection of reliable reference genes for gene expression studies in peach using real-time PCR. BMC Mol Biol 10:1–13. https://doi.org/10.1186/1471-2199-10-71
Tsujii M, Tanudjaja E, Uozumi N (2020) Diverse physiological functions of cation proton antiporters across bacteria and plant cells. Int J Mol Sci 21:1–14. https://doi.org/10.3390/ijms21124566
Uozumi N (2001) Escherichia coli as an expression system for K+ transport systems from plants. Am J Physiol Cell Physiol 281:C733–9. https://doi.org/10.1152/ajpcell.2001.281.3.c733
Wang M, Zheng Q, Shen Q, Guo S (2013) The critical role of potassium in plant stress response. Int J Mol Sci 14:7370–7390. https://doi.org/10.3390/ijms14047370
White PJ, Broadley MR (2000) Mechanisms of caesium uptake by plants. New Phytol 147:241–256. https://doi.org/10.1046/j.1469-8137.2000.00704.x
Xu J, Li HD, Chen LQ, Wang Y, Liu LL, He L et al (2006) A protein kinase, interacting with two calcineurin B-like proteins, regulates K+ transporter AKT1 in Arabidopsis. Cell 125:1347–1360. https://doi.org/10.1016/j.cell.2006.06.011
Zhang M, Huang P, Ji Y, Wang S, Wang SS, Li Z et al (2020) KUP9 maintains root meristem activity by regulating K + and auxin homeostasis in response to low K. EMBO Rep 21:1–20. https://doi.org/10.15252/embr.202050164
Acknowledgements
We thank Megumi Kato for plant cultivation, Naomi Hoshi for plasmid construction, and Anke Reinders for critical reading of the manuscript. 43K was supplied through Supply Platform of Short-lived Radioisotopes by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (16H06278).
Availability of data and material
Not applicable.
Funding
This work was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (20KK0127 and 21 K19060 to NU, 21H05266 21KK0268 and 22K19121 to YI and 20 K15447 to SS, respectively), Moonshot R&D (JPMJMS2033) from JST to NU.
Author information
Authors and Affiliations
Contributions
TY, TU., TH and MY performed biological measurements and biochemical experiments, HI and HK produced radioisotope, YI and NU designed the research, and SS and NU wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Not applicable.
Additional information
Handling editor: Ingo Dreyer
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamanashi, T., Uchiyama, T., Saito, S. et al. Potassium transporter KUP9 participates in K+ distribution in roots and leaves under low K+ stress. Stress Biology 2, 52 (2022). https://doi.org/10.1007/s44154-022-00074-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-022-00074-x