Abstract
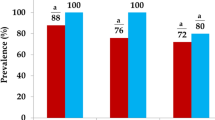
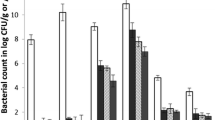
In this study, the occurrence of tetracycline resistance genes among bacteria present on the raw egg surface was analysed. Metagenomic method was used in this study for the rapid detection and better surveillance on antibiotic resistance transfer. From the study, meta DNA extracted from the raw egg surface wash was found to be positive for tetB gene. Whereas the meta DNA from the poultry slaughter house cutting surface, was found to have both tetA and tetB genes. The antibiotic resistance pattern of bacteria isolated from the raw egg samples further showed 76% of them to be resistant to more than 3 different categories of antibiotics. As the antibiotics used in the poultry feed are known to contribute to the fast emergence of resistance in bacteria associated with poultry derived food, it is important to rigorously monitor the feed in poultry farms in order to minimise the rapid spread of antibiotic resistance.


Similar content being viewed by others
Data availability
The sequences of the bacterial 16S rDNA are available in NCBI under the accession numbers OQ195798- OQ195814.
References
Abd El-Ghany, W.A.: Pseudomonas aeruginosa infection of avian origin: zoonosis and one health implications. Veterinary World. 14(8), 2155 (2021)
Burgos, M.J., Márquez, M.L., Pulido, R.P., et al.: Virulence factors and antimicrobial resistance in Escherichia coli strains isolated from hen egg shells. Int. J. Food Microbiol. 5(238), 89–95 (2016)
Chousalkar, K.K., Flynn, P., Sutherland, M., et al.: Recovery of salmonella and Escherichia coli from commercial egg shells and effect of translucency on bacterial penetration in eggs. Int. J. Food Microbiol. 142(1–2), 207 (2010)
Dutta, T.K., Yadav, S.K., Chatterjee, A.: Antibiotics as feed additives for livestock: human health concerns. Indian J. Anim. Health. 58(2), 121–136 (2019)
Jochum, J.M., Redweik, G.A., Ott, L.C., et al.: Bacteria broadly-resistant to last resort antibiotics detected in commercial chicken farms. Microorganisms. 9(1), 141 (2021)
Kowalczyk, J., Czokajło, I., Gańko, M., et al.: Identification and antimicrobial resistance in Klebsiella spp. Isolates from Turkeys in Poland between 2019 and 2022. Animals. 12(22), 3157 (2022)
Krumperman, P.H.: Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl. Environ. Microbiol. 46(1), 165–170 (1983)
Musgrove, M.T., Northcutt, J.K., Jones, D.R., et al.: Enterobacteriaceae and related organisms isolated from shell eggs collected during commercial processing. Poult. Sci. 87(6), 1211–1218 (2008)
Nordstrom, L., Liu, C.M., Price, L.B.: Foodborne urinary tract infections: a new paradigm for antimicrobial-resistant foodborne illness. Front. Microbiol. 6(4), 29 (2013)
Shah, D.H., Board, M.M., Crespo, R., et al.: The occurrence of Salmonella, extended-spectrum β-lactamase producing Escherichia coli and carbapenem resistant non-fermenting Gram-negative bacteria in a backyard poultry flock environment. Zoonoses Public Health 67(6), 742–753 (2020)
Shah, D.H., Board, M.M., Crespo, R., Guard, J., Paul, N.C., Faux, C.: The occurrence of Salmonella, extended-spectrum β-lactamase producing Escherichia coli and carbapenem resistant non-fermenting Gram-negative bacteria in a backyard poultry flock environment. Zoonoses Public Health 67(6), 742–753 (2020)
Sivakumar, S., Girija, A.S., Priyadharsini, J.V.: Evaluation of the inhibitory effect of caffeic acid and gallic acid on tetR and tetM efflux pumps mediating tetracycline resistance in Streptococcus sp., using computational approach. J King Saud Univ Sci. 32(1), 904–9 (2020)
Sreejith, S., Shajahan, S., Prathiush, P.R., et al.: Healthy broilers disseminate antibiotic resistance in response to tetracycline input in feed concentrates. Microb. Pathog. 1(149), 104562 (2020)
Sreejith, S., Shajahan, S., Prathiush, P.R., et al.: Rapid detection of mobile resistance genes tetA and tetB from metaplasmid isolated from healthy broiler feces. Microb. Pathog. 1(166), 105504 (2022)
Wang, Y., Hu, Y., Cao, J., et al.: Antibiotic resistance gene reservoir in live poultry markets. J. Infect. 78(6), 445–453 (2019)
Zhou, S., Zhu, Y., Yan, Y., et al.: Deciphering extracellular antibiotic resistance genes (eARGs) in activated sludge by metagenome. Water Res. 15(161), 610–620 (2019)
Acknowledgement
The authors are grateful to Kairali Gaveshana Puraskaram 2021 by KSCSTE for providing facilities to conduct this work.
Funding
The research work is not funded by any agencies.
Author information
Authors and Affiliations
Contributions
Maya Mathew: conceptualization, writing—original draft, methodology, investigation, formal analysis. Swetha Haridas: methodology, investigation. Sreejith S. : methodology, resources. Sandhya C.: supervision. Jyothis Mathew: supervision. Radhakrishnan E.K.: writing—review & editing, resources, visualization, project administration, conceptualization.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mathew, M., Haridas, S., Sreejith, S. et al. Antibiotic resistance genes on raw egg surface with potential to transmit through supply chain. Proc.Indian Natl. Sci. Acad. (2024). https://doi.org/10.1007/s43538-024-00303-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s43538-024-00303-z