Abstract
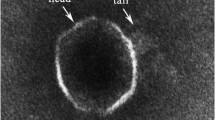
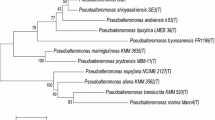
Seamounts are hotspots for marine life, but to date, no bacteriophages have been reported. Here, a novel Bacillus podophage (named as Bacillus phage Gxv1) was isolated from deep-sea seamount sediments of the western Pacific Ocean (~ 5790 m). Phage Gxv1 has a hexameric head ~ 42–53 nm in diameter and a short tail of ~ 30 nm long, which is a typical feature of the Podoviridae family. One-step curve analysis showed that Gxv1 is a lytic phage that can initiate host lysis within 3.5 h post-infection, and has a relatively large burst size. The 21,781-bp genome contains 34 predicted genes, and the G + C content of phage Gxv1 is 39.69%. Whole-genome comparison of phage Gxv1 with known bacteriophages, using BlastN analysis against the IMG/VR database, revealed that phage Gxv1 is closely related to Bacillus phage phi29 that infects Bacillus subtilis, and their genome-wide similarity is 93.62%. Phylogenetic analysis based on DNA polymerase showed that phage Gxv1 belongs to the Salasvirus genus. Multiple genome alignment showed that phage Gxv1 shares a high level of sequence similarity and common gene order with Bacillus phage phi29. However, some sequences are unique to phage Gxv1, and this region contains genes encoding DNA packing protein, DNA replication protein, and unknown protein. These sequences exhibit low sequence similarity to known bacteriophages, highlighting an unknown origin of these sequences. This study will help improve our understanding of the Salasvirus genus and phage diversity in deep-sea seamounts.





Similar content being viewed by others
Data availability
The complete genome sequence for Bacillus phage Gxv1 was deposited in GenBank database under the accession number MT459794.
References
Ackermann HW (2001) Frequency of morphological phage descriptions in the year 2000. Arch Virol 146:843–857
Alicia DP, Laurentino V, Miguel DV, Margarita S (2012) Involvement of residues of the 29 terminal protein intermediate and priming domains in the formation of a stable and functional heterodimer with the replicative DNA polymerase. Nucleic Acids Res 40:3886–3897
Ana MA, Javier A, Carlos VJ, Fernanda MM, Alicia O, Minerva E, Ana M, Stuart H (2012) Is there a seamount effect on microbial community structure and biomass? The case study of Seine and Sedlo seamounts (northeast Atlantic). PLoS ONE 7:e29526
Anderson DL, Hickman DD, Reilly BE (1966) Structure of Bacillus subtilis bacteriophage phi 29 and the length of phi 29 deoxyribonucleic acid. J Bacteriol 91:2081–2089
Blanco L, Bernad A, Lázaro JM, Martí G, Garmendia C, Salas M (1989) Highly efficient DNA synthesis by the phage phi 29 DNA polymerase. Symmetrical mode of DNA replication. J Biol Chem 264:8935–8940
Chen A, Gui GF, Zhuo Y, Chai YQ, Xiang Y, Yuan R (2015) Signal-off electrochemiluminescence biosensor based on phi29 DNA polymerase mediated strand displacement amplification for microRNA detection. Anal Chem 87:6328–6334
Chiang OE, Quiñones RA (2007) Relationship between viral and prokaryotic abundance on the Bajo O’Higgins Seamount (Humboldt Current System off Chile). Sci Mar 71:37–46
Colombet J, Robin A, Lavie L, Bettarel Y, Cauchie HM, Sime-Ngando T (2008) Virioplankton ‘pegylation’: use of PEG (polyethylene glycol) to concentrate and purify viruses in pelagic ecosystems. J Microbiol Methods 71:212–219
Forges BRD, Koslow JA, Poore GCB (2000) Diversity and endemism of the benthic seamount fauna in the southwest Pacific. Nature 405:944–947
Gascón I, Lázaro JM, Salas M (2000) Differential functional behavior of viral phi29, Nf and GA-1 SSB proteins. Nucleic Acids Res 28:2034–2042
Gollner S, Kaiser S, Menzel L, Jones DOB, Brown A, Mestre NC, van Oevelen D, Menot L, Colaço A, Canals M, Cuvelier D, Durden JM, Gebruk A, Egho GA, Haeckel M, Marcon Y, Mevenkamp L, Morato T, Pham CK, Purser A et al (2017) Resilience of benthic deep-sea fauna to mining activities. Mar Environ Res 129:76–101
Hamidi SV, Ghourchian H, Tavoosidana G (2015) Real-time detection of H5N1 influenza virus through hyperbranched rolling circle amplification. Analyst 140:1502–1509
Hillier JK, Watts AB (2007) Global distribution of seamounts from ship-track bathymetry data. Geophys Res Lett 34:173–180
Hwang CY, Cho BC (2002) Virus-infected bacteria in oligotrophic open waters of the East Sea, Korea. Aquat Microb Ecol 30:1–9
Ito J, Meinke W, Hathaway G, Spizizen J (1973) Studies on Bacillus subtilis bacteriophage φ15. Virology 56:110–122
Jin M, Ye T, Zhang X (2013) Roles of bacteriophage GVE2 endolysin in host lysis at high temperatures. Microbiol SGM 159:1597–1605
Jin M, Guo X, Zhang R, Qu W, Gao B, Zeng R (2019) Diversities and potential biogeochemical impacts of mangrove soil viruses. Microbiome 7:58
Koslow JA, Gowlettholmes K, Lowry JK (2001) Seamount benthic macrofauna off southern Tasmania: community structure and impacts of trawling. Mar Ecol Prog 4:111–125
Marc BB, Massimo V, Eduardo R (2007) Causes for the intriguing presence of tRNAs in phages. Genome Res 17:1486–1495
Meijer WJ, Horcajadas JA, Salas M (2001) φ29 family of phages. Microbiol Mol Biol Rev 65:261–287
Morato T, Hoyle SD, Allain V, Nicol SJ (2010) Seamounts are hotspots of pelagic biodiversity in the open ocean. Proc Nat Acad Sci USA 107:9707–9711
Paez-Espino D, Chen IMA, Palaniappan K, Ratner A, Chu K, Szeto E, Pillay M, Huang J, Markowitz VM, Nielsen T (2016) IMG/VR: a database of cultured and uncultured DNA viruses and retroviruses. Nucleic Acids Res 45:457–465
Parada V, Herndl GJ, Weinbauer MG (2006) Viral burst size of heterotrophic prokaryotes in aquatic systems. J Mar Biol Assoc UK 86:613–621
Pečenková T, Pačes V (1999) Molecular phylogeny of φ29-like phages and their evolutionary relatedness to other protein-primed replicating phages and other phages hosted by gram-positive bacteria. J Mol Evol 48:197–208
Pedulla ML, Ford ME, Houtz JM, Karthikeyan T, Wadsworth C, Lewis JA, Jacobs-Sera D, Falbo J, Gross J, Pannunzio NR (2003) Origins of highly mosaic mycobacteriophage genomes. Cell 113:171–182
Rogers AD (2019) Threats to seamount ecosystems and their management. In: Sheppard C (ed) World seas: an environmental evaluation, 2nd edn. Academic Press, New York, pp 427–451
Rohwer F, Thurber RV (2009) Viruses manipulate the marine environment. Nature 459:207–212
Rowden AA, Dower JF, Schlacher TA, Consalvey M, Clark MR (2010) Paradigms in seamount ecology: fact, fiction and future. Mar Ecol 31:226–241
Salas M (1983) A new mechanism for the initiation of replication of Φ 29 and adenovirus DNA: priming by the terminal protein. In: Doerfler W (ed) The molecular biology of adenoviruses 1. Springer, Berlin, pp 89–106
Salas M (2007) 40 years with bacteriophage ø29. Annu Rev Microbiol 61:1–22
Schilling T, Hoppert M, Daniel R, Hertel R (2018) Complete genome sequence of vB_BveP-Goe6, a virus infecting Bacillus velezensis FZB42. Genome Announc 6:e00008–00018
Suttle CA (2005) Viruses in the sea. Nature 437:356–361
Taehyun P, Struck DK, Deaton JF, Ry Y (2007) Topological dynamics of holins in programmed bacterial lysis. Proc Nat Acad Sci USA 103:19713–19718
Wessel P, Sandwell D, Kim SS (2010) The global seamount census. Oceanography 23:24–33
Ye X, Rossmann MG (2011) Structure of bacteriophage φ29 head fibers has a supercoiled triple repeating helix-turn-helix motif. Proc Nat Acad Sci USA 108:4806–4810
Zhao Y, Zhao Y, Zheng S, Zhao L, Li X, Zhang W, Grégori G, Xiao T (2020) Virioplankton distribution in the tropical western Pacific Ocean in the vicinity of a seamount. MicrobiologyOpen 9:e1031
Acknowledgements
This work was financially supported by China Ocean Mineral Resources R&D Association (No. DY135-B-04), the National Natural Science Foundation of China (No. 41976084), and the Scientific Research Foundation of Third Institute of Oceanography, MNR (No. 2019013).
Author information
Authors and Affiliations
Contributions
MJ and RZ designed the experiments. XG, TZ, and MJ performed the experiments, analyzed the data, and wrote the manuscript. All authors edited and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All the authors declare that they have no competing interests.
Animal and human rights statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Jiamei Li.
Rights and permissions
About this article
Cite this article
Guo, X., Zhang, T., Jin, M. et al. Characterization of Bacillus phage Gxv1, a novel lytic Salasvirus phage isolated from deep-sea seamount sediments. Mar Life Sci Technol 3, 13–19 (2021). https://doi.org/10.1007/s42995-020-00074-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42995-020-00074-8