Abstract
Zika virus (ZIKV) is an arbovirus maintained in nature in two distinct cycles of transmission: urban and sylvatic. Each cycle includes specific vertebrate and invertebrate hosts, and through alternate infections, a conserved consensus sequence is maintained that might vary depending on the cycle. The current study aimed to investigate the ability of ZIKVAF and ZIKVBR to maintain an infectious cycle by alternating passages in cells mimicking the urban (UC) and semi-sylvatic (SC) cycles. The complete genome of the original inoculum and the last passages for each cycle were sequenced by Sanger. Ten passages were performed, as planned, for ZIKVBR UC, ZIKVAF SC, and ZIKVBR SC. ZIKVBR SC showed significant variation in viral titers along the passages, suggesting that the virus is not well adapted to the non-human primate host. ZIKVAF passage in UC was abrogated in the third passage, showing the inability of the African lineage to sustain cycles in human cells, suggesting a low capacity to establish an urban cycle. Several mutations were found in both strains along the passages, but not occurring at equivalent positions. Further studies are needed to elucidate whether any of these specific mutations affect viral fitness.
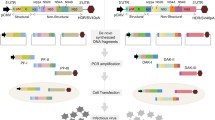
Graphical Abstract
ZIKV strains behave differently in artificial transmission cycles in vitro: Brazilian ZIKV was able to establish urban and semi-sylvatic cycles in vitro. African ZIKV proved unable to cycle among human and mosquito cells and is compatible only with the semi-sylvatic cycle. The main mutations arose in the NS2A region after artificial transmission cycles for both ZIKV strains but not at equivalent positions.







Similar content being viewed by others
Data availability
The data that support the findings of this study are openly available in GenBank, reference numbers OK569755, OK569756, OK569757, OK569758, OK569759, and OK569760, and in the supplementary material of this article.
References
Dick GWA (1953) Epidemiological notes on some viruses isolated in Uganda (Yellow fever, Rift Valley fever, Bwamba fever, West Nile, Mengo, Semliki forest, Bunyamwera, Ntaya, Uganda S and Zika viruses). Trans R Soc Trop Med Hyg 47(1):13–48. https://doi.org/10.1016/0035-9203(53)90021-2
Dick GWA, Kitchen SF, Haddow AJ (1952) Zika virus (I). Isolations and serological specificity. Trans Royal Soc Trop Med Hyg 46(5):509–520. https://doi.org/10.1016/0035-9203(52)90042-4
França GVAd et al (2018) Síndrome congênita associada à infecção pelo vírus Zika em nascidos vivos no Brasil: descrição da distribuição dos casos notificados e confirmados em 2015–2016. Epidemiologia e Serviços de Saúde 27(2):e2017473. https://doi.org/10.5123/S1679-49742018000200014
Cauchemez S et al (2016) Association between Zika virus and microcephaly in French Polynesia, 2013–15: a retrospective study. Lancet 387(10033):2125–2132. https://doi.org/10.1016/S0140-6736(16)00651-6
Magalhães-Barbosa MC et al (2016) Trends of the microcephaly and Zika virus outbreak in Brazil, January-July 2016. Travel Med Infect Dis 14(5):458–463. https://doi.org/10.1016/J.TMAID.2016.09.006
de Oliveira WK et al (2017) Infection-related microcephaly after the 2015 and 2016 Zika virus outbreaks in Brazil: a surveillance-based analysis. Lancet 390(10097):861–870. https://doi.org/10.1016/J.TMAID.2016.09.006
Calvet G et al (2016) Detection and sequencing of Zika virus from amniotic fluid of fetuses with microcephaly in Brazil: a case study. Lancet Infect Dis 16(6):653–660. https://doi.org/10.1016/S1473-3099(16)00095-5
Fleming AM et al (2016) Zika virus genomic RNA possesses conserved G-quadruplexes characteristic of the Flaviviridae family. ACS Infect Dis 2(10):674–681. https://doi.org/10.1021/ACSINFECDIS.6B00109/SUPPL_FILE/ID6B00109_SI_001.PDF
Weaver SC et al (2016) Zika virus: History, emergence, biology, and prospects for control. Antiviral Res 130:69–80. https://doi.org/10.1016/J.ANTIVIRAL.2016.03.010
Kuno G, Chang GJ (2007) Full-length sequencing and genomic characterization of Bagaza, Kedougou, and Zika viruses. Arch Virol 152(4):687–696. https://doi.org/10.1007/S00705-006-0903-Z
Wang A et al (2017) Zika virus genome biology and molecular pathogenesis. Emerg Microbes Infect 6(3):e13. https://doi.org/10.1038/EMI.2016.141
Liu ZY, Shi WF, Qin CF (2019) The evolution of Zika virus from Asia to the Americas. Nat Rev Microbiol 17(3):131–139. https://doi.org/10.1038/S41579-018-0134-9
Pettersson JH-O et al (2016) How did Zika virus emerge in the Pacific Islands and Latin America? mBio 7(5):e01239-16. https://doi.org/10.1128/MBIO.01239-16
Wang L et al (2016) From mosquitos to humans: genetic evolution of Zika virus. Cell Host Microbe 19(5):561–565. https://doi.org/10.1016/J.CHOM.2016.04.006
Metsky HC et al (2017) Zika virus evolution and spread in the Americas. Nature 546(7658):411–415. https://doi.org/10.1038/NATURE22402
Ye Q et al (2016) Genomic characterization and phylogenetic analysis of Zika virus circulating in the Americas. Infect Genet Evol 43:43–49. https://doi.org/10.1016/J.MEEGID.2016.05.004
Althouse BM et al (2016) Potential for Zika virus to establish a sylvatic transmission cycle in the Americas. Plos Negl Trop Dis 10(12):e0005055–e0005055. https://doi.org/10.1371/JOURNAL.PNTD.0005055
Vasilakis N, Weaver SC (2017) Flavivirus transmission focusing on Zika. Curr Opin Virol 22:30–35. https://doi.org/10.1016/J.COVIRO.2016.11.007
Jenkins GM et al (2002) Rates of molecular evolution in RNA viruses: a quantitative phylogenetic analysis. J Mol Evol 54(2):156–165. https://doi.org/10.1007/S00239-001-0064-3
Ciota AT, Kramer LD (2010) Insights into arbovirus evolution and adaptation from experimental studies. Viruses 2(12):2594–2617. https://doi.org/10.3390/V2122594
Grubaugh ND, Ebel GD (2016) Dynamics of West Nile virus evolution in mosquito vectors. Curr Opin Virol 21:132–138. https://doi.org/10.1016/J.COVIRO.2016.09.007
Moser LA et al (2018) Growth and adaptation of Zika virus in mammalian and mosquito cells. Plos Negl Trop Dis 12(11):e0006880. https://doi.org/10.1371/JOURNAL.PNTD.0006880
Deardorff ER et al (2011) West Nile virus experimental evolution in vivo and the trade-off hypothesis. Plos Pathog 7(11):e1002335. https://doi.org/10.1371/JOURNAL.PPAT.1002335
van Boheemen S et al (2017) Quasispecies composition and evolution of a typical Zika virus clinical isolate from Suriname. Sci Rep 7(1):2368. https://doi.org/10.1038/S41598-017-02652-W
Musso D, Gubler DJ (2016) Zika virus. Clin Microbiol Rev 29(3):487–524. https://doi.org/10.1128/CMR.00072-15
Aliota MT et al (2017) Zika in the Americas, year 2: what have we learned? What gaps remain? A report from the Global Virus Network. Antiviral Res 144:223–246. https://doi.org/10.1016/J.ANTIVIRAL.2017.06.001
Faye O et al (2014) Molecular evolution of Zika virus during its emergence in the 20(th) century. Plos Negl Trop Dis 8(1):e2636. https://doi.org/10.1371/JOURNAL.PNTD.0002636
Faria NR et al (2016) Zika virus in the Americas: early epidemiological and genetic findings. Science (New York, N.Y.) 352(6283):345–349. https://doi.org/10.1371/JOURNAL.PNTD.0002636
de Melo Freire CC, et al (2015) Spread of the pandemic Zika virus lineage is associated with NS1 codon usage adaptation in humans [Preprint]. BioRxiv, p 032839. https://doi.org/10.1101/032839
Guth S et al (2020) Ecological processes underlying the emergence of novel enzootic cycles: arboviruses in the neotropics as a case study. Plos Negl Trop Dis 14(8):e0008338. https://doi.org/10.1371/JOURNAL.PNTD.0008338
Terzian ACB et al (2018) Evidence of natural Zika virus infection in neotropical non-human primates in Brazil. Sci Rep 8(1):16034. https://doi.org/10.1038/S41598-018-34423-6
Favoretto SR et al (2019) Zika virus in peridomestic neotropical primates. Northeast Brazil EcoHealth 16(1):61–69. https://doi.org/10.1007/s10393-019-01394-7
Favoretto S et al (2016) First detection of Zika virus in neotropical primates in Brazil: a possible new reservoir [Preprint]. BioRxiv, p 049395. https://doi.org/10.1101/049395
Vanchiere JA et al (2018) Experimental Zika virus infection of neotropical primates. Am J Trop Med Hyg 98(1):173–177. https://doi.org/10.4269/AJTMH.17-0322
Ng WC et al (2017) The 5’ and 3’ untranslated regions of the flaviviral genome. Viruses 9(6):137. https://doi.org/10.3390/V9060137
Queiroz, S.R.d.A., (2011) Estratégias moleculares para utilização do vírus da febre amarela como vetor vacinal. Dissertation, Fundação Oswaldo Cruz - Instituto Aggeu Magalhães. Recife, PE, Brasil.
Faye M et al (2020) Biological characteristics and patterns of codon usage evolution for the African genotype Zika virus. Viruses 12(11):1306. https://doi.org/10.3390/V12111306
Rosenstierne MW et al (2018) Zika virus IgG in infants with microcephaly, Guinea-Bissau, 2016. Emerg Infect Dis 24(5):948–950. https://doi.org/10.3201/EID2405.180153
Bowen JR et al (2017) Zika virus antagonizes type I interferon responses during infection of human dendritic cells. Plos Pathog 13(2):e1006164–e1006164. https://doi.org/10.1371/JOURNAL.PPAT.1006164
Weger-Lucarelli J et al (2016) Vector competence of American mosquitoes for three strains of Zika virus. Plos Negl Trop Dis 10(10):e0005101. https://doi.org/10.1371/JOURNAL.PNTD.0005101
Lazear HM et al (2016) A mouse model of Zika virus pathogenesis. Cell Host Microbe 19(5):720–730. https://doi.org/10.1016/J.CHOM.2016.03.010
Azar SR et al (2017) Differential vector competency of Aedes albopictus populations from the Americas for Zika virus. Am J Trop Med Hyg 97(2):330–339. https://doi.org/10.4269/AJTMH.16-0969
Roundy CM et al (2017) Variation in Aedes aegypti mosquito competence for Zika virus transmission. Emerg Infect Dis 23(4):625–632. https://doi.org/10.3201/EID2304.161484
Simonin Y et al (2017) Differential virulence between Asian and African lineages of Zika virus. Plos Negl Trop Dis 11(9):e0005821–e0005821. https://doi.org/10.1371/JOURNAL.PNTD.0005821
Haddow AD et al (2012) Genetic characterization of Zika virus strains: geographic expansion of the Asian lineage. Plos Negl Trop Dis 6(2):e1477. https://doi.org/10.1371/JOURNAL.PNTD.0001477
Laiton-Donato K et al (2019) Complete genome sequence of a Colombian Zika virus strain obtained from BALB/c mouse brain after intraperitoneal inoculation. Microbiol Resour Announc 8(46):e01719-e1818. https://doi.org/10.1128/MRA.01719-18
Shrivastava S et al (2018) Whole genome sequencing, variant analysis, phylogenetics, and deep sequencing of Zika virus strains. Sci Rep 8(1):15843. https://doi.org/10.1038/S41598-018-34147-7
Strottmann DM et al (2019) Genetic and biological characterisation of Zika virus isolates from different Brazilian regions. Mem Inst Oswaldo Cruz 114:e190150–e190150. https://doi.org/10.1590/0074-02760190150
Alpuche-Lazcano SP et al (2018) Higher cytopathic effects of a Zika virus Brazilian isolate from Bahia compared to a Canadian-imported Thai strain. Viruses 10(2):53. https://doi.org/10.3390/V10020053
Ávila-Pérez G et al (2019) A natural polymorphism in Zika virus NS2A protein responsible of virulence in mice. Sci Rep 9(1):19968. https://doi.org/10.1038/s41598-019-56291-4
Collins ND et al (2019) Inter- and intra-lineage genetic diversity of wild-type Zika viruses reveals both common and distinctive nucleotide variants and clusters of genomic diversity. Emerg Microbes Infect 8(1):1126–1138. https://doi.org/10.1080/22221751.2019.1645572
Wang B et al (2018) Structure and function of Zika virus NS5 protein: perspectives for drug design. Cell Mol Life Sci 75(10):1723–1736. https://doi.org/10.1007/s00018-018-2751-x
Upadhyay AK et al (2017) Crystal structure of full-length Zika virus NS5 protein reveals a conformation similar to Japanese encephalitis virus NS5. Acta Crystallogr Section F, Struct Biol Commun 73(Pt 3):116–122. https://doi.org/10.1107/S2053230X17001601
Zhao B et al (2017) Structure and function of the Zika virus full-length NS5 protein. Nat Commun 8(1):14762. https://doi.org/10.1038/ncomms14762
Longdon B et al (2014) The evolution and genetics of virus host shifts. Plos Pathogens 10(11):e1004395. https://doi.org/10.1371/JOURNAL.PPAT.1004395
Bahir I et al (2009) Viral adaptation to host: a proteome-based analysis of codon usage and amino acid preferences. Mol Syst Biol 5:311. https://doi.org/10.1038/MSB.2009.71
Wang H et al (2016) Analysis of synonymous codon usage bias of Zika virus and its adaption to the hosts. Plos One 11(11):e0166260. https://doi.org/10.1371/JOURNAL.PONE.0166260
Hall TA (1999) BioEdit : a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
TM Calculator. (s.d.). Protein Melting Temperature Calculation. https://tmcalculator.neb.com/#!/main
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A 74(12):5463–5467. https://doi.org/10.1073/PNAS.74.12.5463
Acknowledgements
We thank all the participants who contributed to this study.
Funding
This work is supported by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brazil (CAPES) Finance Code 001.
Author information
Authors and Affiliations
Contributions
M. N. B. conceived the study. All authors designed the study. B. F. M., N. N. M., and C. B. performed the experiments and were involved in the acquisition and analysis of the data; all authors were involved in the interpretation of the data. B. F. M. and N. N. M. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Ethics approval
The study described is exempt from ethics approval.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Flavio Guimaraes Fonseca
Supplementary Information
Table 1 Primers used in the nested-PCR reaction to amplify the complete virus genome. Start and stop sites are based on ZIKV Brazilian strain reference sequence (GenBank NC_035889.1). Primers P4-5F/R and N4-5F/R were only needed for African ZIKV complete genome amplification
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Molina, B.F., Marques, N.N., Bittar, C. et al. African ZIKV lineage fails to sustain infectivity in an in vitro mimetic urban cycle. Braz J Microbiol 54, 1421–1431 (2023). https://doi.org/10.1007/s42770-023-01053-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01053-8