Abstract
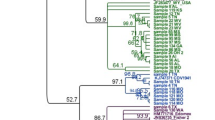
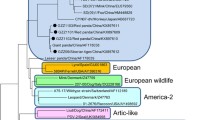
The canine distemper virus (CDV) is responsible for a multisystem infectious disease with high prevalence in dogs and wild carnivores and has vaccination as the main control measure. However, recent studies show an increase in cases including vaccinated dogs in different parts of the world. There are several reasons for vaccine failures, including differences between vaccine strains and wild-type strains. In this study, a phylogenetic analysis of CDV strains from samples of naturally infected, vaccinated, and symptomatic dogs in Goiânia, Goiás, Brazil was performed with partial sequencing of the hemagglutinin (H) gene of CDV. Different sites of amino acid substitutions were found, and one strain had the Y549H mutation, typically present in samples from wild animals. Substitutions in epitopes (residues 367, 376, 379, 381, 386, and 388) that may interfere with the vaccine’s ability to provide adequate protection against infection for CDV were observed. The identified strains were grouped in the South America 1/Europe lineage, with a significant difference from other lineages and vaccine strains. Twelve subgenotypes were characterized, considering a nucleotide identity of at least 98% among the strains. These findings highlight the relevance of canine distemper infection and support the need better monitoring of the circulating strains that contribute to elucidate if there is a need for vaccine update.


Similar content being viewed by others
References
Greene CE (2012) Canine distemper. In: Greene CE (ed) Infectious diseases of the dog and cat, 4th edn. Elsevier Saunders, St. Louis, pp 25–41
Beineke A, Baumgärtner W, Wohlsein P (2015) Cross-species transmission of canine distemper virus-an update. One Health 1:49–59. https://doi.org/10.1016/j.onehlt.2015.09.002
Martinez-Gutierrez M, Ruiz-Saenz J (2016) Diversity of susceptible hosts in canine distemper virus infection: a systematic review and data synthesis. BMC Vet Res 12(1):78. https://doi.org/10.1186/s12917-016-0702-z
von Messling V, Zimmer G, Herrler G, Haas L, Cattaneo R (2001) The hemagglutinin of canine distemper virus determines tropism and cytopathogenicity. J Virol Published online. https://doi.org/10.1128/JVI.75.14.6418-6427.2001
McCarthy AJ, Shaw MA, Goodman SJ (2007) Pathogen evolution and disease emergence in carnivores. Proceedings of the Royal Society B: Biological Sciences 274(1629):3165–3174
Ke GM, Ho CH, Chiang MJ et al (2015) Phylodynamic analysis of the canine distemper virus hemagglutinin gene. BMC Vet Res 11(1):1–15
Nikolin VM, Wibbelt G, Michler FUF, Wolf P, East ML (2012) Susceptibility of carnivore hosts to strains of canine distemper virus from distinct genetic lineages. Vet Microbiol 156(1–2):45–53
Bi Z, Wang Y, Wang X, Xia X (2015) Phylogenetic analysis of canine distemper virus in domestic dogs in Nanjing. China Arch Virol 160(2):523–527
Haas L, Martens W, Greiser-Wilke I et al (1997) Analysis of the haemagglutinin gene of current wild-type canine distemper virus isolates from Germany. Virus Res 48(2):165–171. https://doi.org/10.1016/S0168-1702(97)01449-4
Hashimoto M, Une Y, Mochizuki M (2001) Hemagglutinin genotype profiles of canine distemper virus from domestic dogs in Japan. Arch Virol 146(1):149–155. https://doi.org/10.1007/s007050170198
Fischer CDB, Gräf T, Ikuta N et al (2016) Phylogenetic analysis of canine distemper virus in South America clade 1 reveals unique molecular signatures of the local epidemic. Infect Genet Evol 41:135–141
Duque-Valencia J, Forero-Muñoz NR, Díaz FJ, Martins E, Barato P, Ruiz-Saenz J (2019) Phylogenetic evidence of the intercontinental circulation of a canine distemper virus lineage in the Americas. Sci Rep 9(1):15747. https://doi.org/10.1038/s41598-019-52345-9
Blixenkrone-Möller M, Svansson V, Appel M, Krogsrud J, Have P, Örvell C (1992) Antigenic relationships between field isolates of morbilliviruses from different carnivores. Arch Virol 123(3):279–294. https://doi.org/10.1007/BF01317264
Harder TC, Kenter M, Vos H et al (1996) Canine distemper virus from diseased large felids: biological properties and phylogenetic relationships. J Gen Virol 77(3):397–405
Iwatsuki K, Tokiyoshi S, Hirayama N et al (2000) Antigenic differences in the H proteins of canine distemper viruses. Vet Microbiol 71(3–4):281–286
Zhao JJ, Yan XJ, Chai XL et al (2010) Phylogenetic analysis of the haemagglutinin gene of canine distemper virus strains detected from breeding foxes, raccoon dogs and minks in China. Vet Microbiol 140(1–2):34–42
Panzera Y, Calderon MG, Sarute N et al (2012) Evidence of two co-circulating genetic lineages of canine distemper virus in South America. Virus Res 163(1):401–404. https://doi.org/10.1016/j.virusres.2011.10.008
Radtanakatikanon A, Keawcharoen J, Charoenvisal TN et al (2013) Genotypic lineages and restriction fragment length polymorphism of canine distemper virus isolates in Thailand. Vet Microbiol 166(1–2):76–83. https://doi.org/10.1016/j.vetmic.2013.05.015
Espinal MA, Díaz FJ, Ruiz-Saenz J (2014) Phylogenetic evidence of a new canine distemper virus lineage among domestic dogs in Colombia. South America Vet Microbiol 172(1–2):168–176
Riley MC, Wilkes RP (2015) Sequencing of emerging canine distemper virus strain reveals new distinct genetic lineage in the United States associated with disease in wildlife and domestic canine populations. Virol J 12(1):1–10
Nikolin VM, Olarte-Castillo XA, Osterrieder N et al (2017) Canine distemper virus in the Serengeti ecosystem: molecular adaptation to different carnivore species. Mol Ecol 26(7):2111–2130
Anis E, Holford AL, Galyon GD, Wilkes RP (2018) Antigenic analysis of genetic variants of canine distemper virus. Vet Microbiol 219:154–160
Martella V, Blixenkrone-Møller M, Elia G et al (2011) Lights and shades on an historical vaccine canine distemper virus, the Rockborn strain. Vaccine 29(6):1222–1227. https://doi.org/10.1016/j.vaccine.2010.12.001
Budaszewski R da F, Pinto LD, Weber MN et al (2014) Genotyping of canine distemper virus strains circulating in Brazil from 2008 to 2012. Virus Res 180:76–83. https://doi.org/10.1016/j.virusres.2013.12.024
Martella V, Cirone F, Elia G et al (2006) Heterogeneity within the hemagglutinin genes of canine distemper virus (CDV) strains detected in Italy. Vet Microbiol Published online. https://doi.org/10.1016/j.vetmic.2006.04.019
Demeter Z, Lakatos B, Palade EA, Kozma T, Forgách P, Rusvai M (2007) Genetic diversity of Hungarian canine distemper virus strains. Vet Microbiol 122(3–4):258–269. https://doi.org/10.1016/j.vetmic.2007.02.001
Frisk AL, König M, Moritz A, Baumgärtner W (1999) Detection of canine distemper virus nucleoprotein RNA by reverse transcription-PCR using serum, whole blood, and cerebrospinal fluid from dogs with distemper. Published online, J Clin Microbiol
An DJ, Yoon SH, Park JY, No IS, Park BK (2008) Phylogenetic characterization of canine distemper virus isolates from naturally infected dogs and a marten in Korea. Vet Microbiol Published online. https://doi.org/10.1016/j.vetmic.2008.05.025
Ewing B, Hillier L, Wendl MC, Green P (1998) Base-calling of automated sequencer traces using phred. I Accuracy Assess Genome Res 8(3):175–185. https://doi.org/10.1101/gr.8.3.175
Gordon D, Desmarais C, Green P (2001) Automated finishing with autofinish. Genome Res 11(4):614–625. https://doi.org/10.1101/gr.171401
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Mochizuki M, Hashimoto M, Hagiwara S, Yoshida Y, Ishiguro S (1999) Genotypes of canine distemper virus determined by analysis of the hemagglutinin genes of recent isolates from dogs in Japan. J Clin Microbiol 37(9):2936–2942
Panzera Y, Sarute N, Iraola G, Hernandez M, Perez R (2015) Molecular phylogeography of canine distemper virus: geographic origin and global spreading. Mol Phylogenet Evol 92:147–154. https://doi.org/10.1016/j.ympev.2015.06.015
Cortez A, Heinemann MB, Junior AAF, da Costa LF, de Souza VAF, Megid J (2017) Genetic characterization of the haemagglutinin gene in canine distemper virus strains from naturally infected dogs in Brazil Caracterização genética do gene da hemaglutinina em vírus da cinomose caninca de cães naturalmente infectados no Brasil. Braz J Vet Res Anim Sci 54(4):445–449
Day MJ, Horzinek MC, Schultz RD, Squires RA (2016) WSAVA guidelines for the vaccination of dogs and cats. J Small Anim Pract 57(1):E1
Bae CW, Lee JB, Park SY et al (2013) Deduced sequences of the membrane fusion and attachment proteins of canine distemper viruses isolated from dogs and wild animals in Korea. Virus Genes 47(1):56–65
Benetka V, Leschnik M, Affenzeller N, Möstl K (2011) Phylogenetic analysis of Austrian canine distemper virus strains from clinical samples from dogs and wild carnivores. Veterinary Record 168(14):377
Nava AFD, Cullen L Jr, Sana DA et al (2008) First evidence of canine distemper in Brazilian free-ranging felids. EcoHealth 5(4):513–518
Megid J, de Souza VAF, Teixeira CR et al (2009) Canine distemper virus in a crab-eating fox (Cerdocyon thous) in Brazil: case report and phylogenetic analyses. J Wildl Dis 45(2):527–530
de Hübner OS, Pappen FG, Ruas JL, Vargas GD, Fischer G, Vidor T (2010) Exposure of pampas fox (Pseudalopex gymnocercus) and crab-eating fox (Cerdocyon thous) from the Southern region of Brazil to canine distemper virus (CDV), canine parvovirus (CPV) and canine coronavirus (CCoV). Brazilian Arch Biol Technol 53(3):593–597
Furtado MM, Hayashi EMK, Allendorf SD et al (2016) Exposure of free-ranging wild carnivores and domestic dogs to canine distemper virus and parvovirus in the Cerrado of Central Brazil. EcoHealth 13(3):549–557. https://doi.org/10.1007/s10393-016-1146-4
Sugiyama M, Ito N, Minamoto N, Tanaka S (2002) Identification of immunodominant neutralizing epitopes on the hemagglutinin protein of rinderpest virus. J Virol 76(4):1691–1696
von Messling V, Oezguen N, Zheng Q, Vongpunsawad S, Braun W, Cattaneo R (2005) Nearby clusters of hemagglutinin residues sustain SLAM-dependent canine distemper virus entry in peripheral blood mononuclear cells. J Virol 79(9):5857–5862
Liebert UG, Flanagan SG, Löffler S, Baczko K, ter Meulen V, Rima BK (1994) Antigenic determinants of measles virus hemagglutinin associated with neurovirulence. J Virol 68(3):1486–1493
Anis E, Newell TK, Dyer N, Wilkes RP (2018) Phylogenetic analysis of the wild-type strains of canine distemper virus circulating in the United States. Virol J 15(1):1–10
Pardo IDR, Johnson GC, Kleiboeker SB (2005) Phylogenetic characterization of canine distemper viruses detected in naturally infected dogs in North America. J Clin Microbiol 43(10):5009–5017
Gámiz C, Martella V, Ulloa R, Fajardo R, Quijano-Hernandéz I, Martínez S (2011) Identification of a new genotype of canine distemper virus circulating in America. Vet Res Commun 35(6):381–390. https://doi.org/10.1007/s11259-011-9486-6
Acknowledgements
The authors thank the Graduate Support Program (Programa de Apoio à Pós-Graduação, Brazil; PROAP) and the Coordination Office for Improvement of Higher-Education Personnel (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, CAPES; Brazil) for their assistance in acquiring materials to carry out the research and for granting the researcher a scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Fernando R. Spilki
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Menezes, K.M.F., Dábilla, N., Souza, M. et al. Identification of a new polymorphism on the wild-type canine distemper virus genome: could this contribute to vaccine failures?. Braz J Microbiol 54, 665–678 (2023). https://doi.org/10.1007/s42770-023-00971-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-00971-x