Abstract
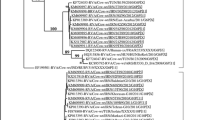
Stool samples were collected from calves from nine family-based small dairy farms in the state of Rio de Janeiro, for detection and characterization of rotavirus (RV) species A, B, and C (RVA, RVB, and RVC, respectively) by reverse transcription polymerase chain reaction. Twenty-six samples (27.7%) were positive for at least one of the species: 22 (23.4%) samples were positive only for RVA, 3 (3.2%) were positive for RVC, and one sample (1.1%) had co-infection of RVA and RVC. RVB was not detected. Seven (21.9%; n = 32) animals with diarrhea and 19 (30.1% n = 62) asymptomatic animals were positive, with no significant difference in positivity (p = 0.3677). RV was detected in all properties studied, at rates between 14.3 and 80%, demonstrating the widespread circulation of RV in four of the seven geographic regions of the state of Rio de Janeiro. Infection was more prevalent among animals ≤ 6 months of age. Sequence analysis of a portion of the RVA VP6-encoding gene identified the I2 genotype. RVC was also detected; to our knowledge, this is the first description of this agent in cattle in Brazil. The data presented here should add knowledge regarding the importance and prevalence of RV in our national territory, and may facilitate the planning and implementation of control and prevention measures for bovine rotavirus infections in Brazil.

Similar content being viewed by others
References
Geletu US, Usmael MA, Bari FD (2021) Rotavirus in calves and its zoonotic importance. Vet Med Int 2021:6639701. https://doi.org/10.1155/2021/6639701
Gomez DE, Weese JS (2017) Viral enteritis in calves. Can Vet J 58:1267–1274. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5680732/
Dhama K, Chauhan RS, Mahendran M, Malik SVS (2009) Rotavirus diarrhea in bovines and other domestic animals. Vet Res Commun 33:1–23. https://doi.org/10.1007/s11259-008-9070-x
Papp H, László B, Jakab F, Ganesh B, De Grazia S, Matthijnssens J, Ciarlet M, Martella V, Bányai K (2013) Review of group A rotavirus strains reported in swine and cattle. Vet Microbiol 165:190–199. https://doi.org/10.1016/j.vetmic.2013.03.020
ABIEC (2021). Associação Brasileira das Indústrias Exportadoras de Carnes. Beef Report. Perfil da Pecuária no Brasil. http://abiec.com.br/publicacoes/beef-report-2021/. Accessed 16 September 2021.
Sadiq A, Bostan N, Yinda KC, Naseem S, Sattar S (2018) Rotavirus: genetics, pathogenesis and vaccine advances. Rev Med Virol 28:e2003. https://doi.org/10.1002/rmv.2003
MAPA (2013). Ministério da Agricultura, Pecuária e Abastecimento. Instrução normativa nº 50, de 24 de Setembro de 2013. https://www.gov.br/agricultura/pt-br/assuntos/sanidade-animal-e-vegetal/saude-animal/arquivos-sisa/Listadedoencasanimaisdenotificaoobrigatoria.pdf/view. Accessed 25 September 2021.
EMBRAPA (2015). Empresa Brasileira de Pesquisa Agropecuária. Circular Técnica. Manual de boas práticas de vacinação e imunização de bovinos. http://ainfo.cnptia.embrapa.br/digital/bitstream/item/128128/1/CiT-47-15-online.pdf. Accessed 25 September 2021.
Mukherjee A, Dutta D, Ghosh S, Bagchi P, Chattopadhyay S, Nagashima S, Kobayashi N, Dutta P, Krishnan T, Naik TN, Chawla-Sarkar M (2009) Full genomic analysis of a human group A rotavirus G9P[6] strain from Eastern India provides evidence for porcine-to-human interspecies transmission. Arch Virol 154:733–746. https://doi.org/10.1007/s00705-009-0363-3
Komoto S, Pongsuwannab Y, Tacharoenmuangb R, Guntapongb R, Idea T, Higo-Moriguchia K, Tsujic T, Yoshikawad T, Taniguchia K (2016) Whole genomic analysis of bovine group A rotavirus strains A5–10 and A5–13 provides evidence for close evolutionary relationship with human rotaviruses. Vet Microbiol 195:37–57. https://doi.org/10.1016/j.vetmic.2016.09.003
Marton S, Dóró R, Fehér E, Forró B, Ihász K, Varga-Kugler R, Farkas SL, Bányai K (2017) Whole genome sequencing of a rare rotavirus from archived stool sample demonstrates independent zoonotic origin of human G8P[14] strains in Hungary. Virus Res 227:96–103. https://doi.org/10.1016/j.virusres.2016.09.012
Castells M, Caffarena RD, Casaux ML, Schild C, Miño S, Castells F, Castells D, Victoria M, Riet-Correa F, Giannitti F, Parreño V, Colina R (2020) Phylogenetic analyses of rotavirus A from cattle in uruguay reveal the circulation of common and uncommon genotypes and suggest interspecies transmission. Pathogens 9:570. https://doi.org/10.3390/pathogens9070570
Cook N, Bridger J, Kendall K, Gomara MI, El-Attar L, Gray J (2004) The zoonotic potential of rotavirus. J Infec 48:289–302. https://doi.org/10.1016/j.jinf.2004.01.018
Martella V, Bányai K, Matthijnssens J, Buonavoglia C, Ciarlet M (2010) Zoonotic aspects of rotaviruses. Vet Microbiol 140:246–255. https://doi.org/10.1016/j.vetmic.2009.08.028
Dóró R, Farkas SL, Martella V, Bányai K (2015) Zoonotic transmission of rotavirus: surveillance and control. Expert Rev Anti Infect Ther 13:1337–1350. https://doi.org/10.1586/14787210.2015.1089171
Fritzen JTT, Oliveira MV, Lorenzetti E, Alfieri AF, Alfieri AA (2020) Genotype constellation of a rotavirus A field strain with an uncommon G8P[11] genotype combination in a rotavirus-vaccinated dairy cattle herd. Arch Virol 165:1855–1861. https://doi.org/10.1007/s00705-020-04675-7
Gouvea V, Allen JR, Glass RI, Fang ZY, Bremont M, Cohen J, McCrae MA, Saif LJ, Sinarachatanant P, Caul EO (1991) Detection of group B and C rotaviruses by polymerase chain reaction. J Clin Microbiol 29:519–523. https://doi.org/10.1128/jcm.29.3.519-523.1991
Flores PS, Costa FB, Amorim AR, Mendes GS, Rojas M, Santos N (2021) Rotavirus A, C, and H in Brazilian pigs: potential for zoonotic transmission of RVA. J Vet Diagn Invest 33:129–135. https://doi.org/10.1177/1040638720967673
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Alfieri AA, Parazzi ME, Takiuchi E, Médici KC, Alfieri AF (2006) Frequency of group A rotavirus in diarrhoeic calves in Brazilian cattle herds, 1998–2002. Trop Anim Health Prod 38:521–526. https://doi.org/10.1007/s11250-006-4349-9
Caruzo TA, Brito WM, Munford V, Rácz ML (2010) Molecular characterization of G and P-types bovine rotavirus strains from Goiás, Brazil: high frequency of mixed P-type infections. Mem Inst Oswaldo Cruz 105:1040–1043. https://doi.org/10.1590/s0074-02762010000800014
Freitas PPS, Uyemura SA, Silva DG, Samara SI, Buzinaro MG (2011) Rotavírus bovino: fatores de risco, prevalência e caracterização antigênica de amostras em rebanhos leiteiros no estado de São Paulo. Arq Bras Med Vet Zootec 63:820–827. https://doi.org/10.1590/S0102-09352011000400005
Rocha TG, Silva FD, Gregori F, Alfieri AA, Buzinaro MD, Fagliari JJ (2017) Longitudinal study of bovine rotavirus group A in newborn calves from vaccinated and unvaccinated dairy herds. Trop Anim Health Prod 49:783–790. https://doi.org/10.1007/s11250-017-1263-2
Fritzen JTT, Oliveira MV, Lorenzetti E, Miyabe FM, Viziack MP, Rodrigues CA, Ayres H, Alfieri AF, Alfieri AA (2019) Longitudinal surveillance of rotavirus A genotypes circulating in a high milk yield dairy cattle herd after the introduction of a rotavirus vaccine. Vet Microbiol 230:260–264. https://doi.org/10.1016/j.vetmic.2019.02.022
Fritzen JTT, Lorenzetti E, Oliveira MV, Bon VR, Ayres H, Alfieri AF, Alfieri AA (2019) Cross-sectional study of the G and P genotypes of rotavirus A field strains circulating in regularly vaccinated dairy cattle herds. Trop Anim Health Prod 51:887–892. https://doi.org/10.1007/s11250-018-1769-2
Barros BCV, Chagas EN, Bezerra LW, Ribeiro LG, Duarte Júnior JWB, Pereira D, da Penha Junior ET, Silva JR, Bezerra DAM, Bandeira RS, Pinheiro HHC, Guerra SFDS, Guimarães RJPSE, Mascarenhas JDP (2018) Rotavirus A in wild and domestic animals from areas with environmental degradation in the Brazilian Amazon. PLoS ONE 13:e0209005. https://doi.org/10.1371/journal.pone.0209005
Cruvinel LB, Ayres H, Zapa D, Nicaretta JE, Couto L, Heller LM, Bastos T, Cruz BC, Soares VE, Teixeira WF, de Oliveira JS, Fritzen JT, Alfieri AA, Freire RL, Lopes W (2020) Prevalence and risk factors for agents causing diarrhea (Coronavirus, Rotavirus, Cryptosporidium spp., Eimeria spp., and nematodes helminthes) according to age in dairy calves from Brazil. Trop Anim Health Prod 52:777–791. https://doi.org/10.1007/s11250-019-02069-9
Abe M, Ito N, Morikawa S, Takasu M, Murase T, Kawashima T, Kawai JK, Sugiyama M (2009) Molecular epidemiology of rotaviruses among healthy calves in Japan: Isolation of a novel bovine rotavirus bearing new P and G genotypes. Virus Res 144:250–257. https://doi.org/10.1016/j.virusres.2009.05.005
Steyer A, Poljšak-Prijatelj M, Barlič-Maganja D, Marin J (2008) Human, porcine and bovine rotaviruses in Slovenia: evidence of interspecies transmission and genome reassortment. J Gen Virology 89:1690–1698. https://doi.org/10.1099/vir.0.2008/001206-0
Tsunemitsu H, Jiang B, Saif LJ (1996) Sequence comparison of the VP7 gene encoding the outer capsid glycoprotein among animal and human group C rotaviruses. Arch Virol 141:705–713. https://doi.org/10.1007/BF01718328
Chang KO, Nielsen PR, Ward LA, Saif LJ (1999) Dual infection of gnotobiotic calves with bovine strains of group A and porcine-like group C rotaviruses influences pathogenesis of the group C rotavirus. J Virol 73:9284–9293. https://doi.org/10.1128/JVI.73.11.9284-9293.1999
Soma J, Tsunemitsu H, Miyamoto T, Suzuki G, Sasaki T, Suzuki T (2013) Whole-genome analysis of two bovine rotavirus C strains: Shintoku and Toyama. J Gen Virol 94:128–135. https://doi.org/10.1099/vir.0.046763-0
Mawatari T, Hirano K, Tsunemitsu H, Suzuki T (2014) Whole-genome analysis of bovine rotavirus species C isolates obtained in Yamagata, Japan, 2003–2010. J Gen Virol 95:1117–1125. https://doi.org/10.1099/vir.0.062166-0
Mawatari T, Taneichi A, Kawagoe T, Hosokawa M, Togashi K, Tsunemitsu H (2004) Detection of a bovine group C rotavirus from adult cows with diarrhea and reduced milk production. J Vet Med Sci 66:887–890. https://doi.org/10.1292/jvms.66.887
Chang KO, Parwani AV, Smith D, Saif LJ (1997) Detection of group B rotaviruses in fecal samples from diarrheic calves and adult cows and characterization of their VP7 genes. J Clin Microbiol 35:2107–2110. https://doi.org/10.1128/jcm.35.8.2107-2110.1997
Tsunemitsu H, Morita D, Takaku H, Nishimori T, Imai K, Saif LJ (1999) First detection of bovine group B rotavirus in Japan and sequence of its VP7 gene. Arch Virol 144:805–815. https://doi.org/10.1007/s007050050546
Matthijnssens J, Ciarlet M, Heiman E, Arijs I, Delbeke T, McDonald SM, Palombo EA, Iturriza-Gómara M, Maes P, Patton JT, Rahman M, Van Ranst M (2008) Full genome-based classification of rotaviruses reveals a common origin between human Wa-Like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J Virol 82:3204–3219. https://doi.org/10.1128/JVI.02257-07
Medeiros TNS, Lorenzetti E, Alfieri AF, Alfieri AA (2015) Phylogenetic analysis of a G6P[5] bovine rotavirus strain isolated in a neonatal diarrhea outbreak in a beef cattle herd vaccinated with G6P[1] and G10P[11] genotypes. Arch Virol 160:447–451. https://doi.org/10.1007/s00705-014-2271-4
Ghosh S, Varghese V, Sinha M, Kobayashi N, Naik TN (2007) Evidence for interstate transmission and increase in prevalence of bovine group B rotavirus strains with a novel VP7 genotype among diarrhoeic calves in Eastern and Northern states of India. Epidemiol Infect 135:1324–1330. https://doi.org/10.1017/S0950268806007813
Abe M, Ito N, Masatani T, Nakagawa K, Yamaoka S, Kanamaru Y, Suzuki H, Shibano K, Arashi Y, Sugiyama M (2011) Whole genome characterization of new bovine rotavirus G21P[29] and G24P[33] strains provides evidence for interspecies transmission. J Gen Virol 92:952–960. https://doi.org/10.1099/vir.0.028175-0
Sawant PM, Digraskar SS, Gopalkrishna V (2020) Molecular characterization of unusual G10P[33], G6P[14] genomic constellations of group A rotavirus and evidence of zooanthroponosis in bovines. Infect Genet Evol 84:104385. https://doi.org/10.1016/j.meegid.2020.104385
Malik YS, Kumar N, Sharma K, Saurabh S, Dhama K, Prasad M, Ghosh S, Bányai K, Kobayashi N, Singh RK (2016) Multispecies reassortant bovine rotavirus strain carries a novel simian G3-like VP7 genotype. Infect Genet Evol 41:63–72. https://doi.org/10.1016/j.meegid.2016.03.023
Barreiros MA, Alfieri AF, Médici KC, Leite JP, Alfieri AA (2004) G and P genotypes of group A rotavirus from diarrhoeic calves born to cows vaccinated against the NCDV (P[1], G6) rotavirus strain. J Vet Med B Infect Dis Vet Public Health 51:104–109. https://doi.org/10.1111/j.1439-0450.2004.00737.x
Matthijnssens J, Desselberger U (2012) Genome diversity and evolution of rotaviruses. In: Hacker J, Dobrindt U, Kurth R (eds) Genome Plasticity and Infectious Diseases. ASM Press, Washington, DC, pp 219–241
Matthijnssens J, Potgieter CA, Ciarlet M, Parreño V, Martella V, Bányai K, Garaicoechea L, Palombo EA, Novo L, Zeller M, Arista S, Gerna G, Rahman M, Ranst M (2009) Are human P[14] rotavirus strains the result of interspecies transmissions from sheep or other ungulates that belong to the mammalian order Artiodactyla? J Virol 83:2917–2929. https://doi.org/10.1128/JVI.02246-08
Jeong S, Than VT, Lim I, Kim W (2014) Whole-genome analysis of a rare human Korean G3P rotavirus strain suggests a complex evolutionary origin potentially involving reassortment events between feline and bovine rotaviruses. PLoS ONE 9:e97127. https://doi.org/10.1371/journal.pone.0097127
Tacharoenmuang R, Komoto S, Guntapong RIT, Sinchai P, Upachai S, Yoshikawa T, Tharmaphornpilas P, Sangkitporn S, Taniguchi K (2016) Full genome characterization of novel DS-1-Like G8P[8] rotavirus strains that have emerged in Thailand: reassortment of bovine and human rotavirus gene segments in emerging DS-1-like intergenogroup reassortant strains. PLoS ONE 11:e0165826. https://doi.org/10.1371/journal.pone.0165826
Bwogi J, Jere KC, Karamagi C, Byarugaba DK, Namuwulya P, Baliraine FN, Desselberger U, Iturriza-Gomara M (2017) Whole genome analysis of selected human and animal rotaviruses identified in Uganda from 2012 to 2014 reveals complex genome reassortment events between human, bovine, caprine and porcine strains. PLoS ONE 12:e0178855. https://doi.org/10.1371/journal.pone.0178855
Moutelíková R, Sauer P, Dvořáková Heroldová M, Holá V, Prodělalová J (2019) Emergence of Rare bovine-human reassortant DS-1-like rotavirus A strains with G8P[8] genotype in human patients in the Czech Republic. Viruses 11:1015. https://doi.org/10.3390/v11111015
Mishra N, Reslan L, El-Husseini M, Raoof H, Finianos M, Guo C, Thakkar R, Inati A, Dbaibo G, Lipkin WI, Zaraket H (2020) Full genome characterization of human G3P[6] and G3P[9] rotavirus strains in Lebanon. Infect Genet Evol 78:104133. https://doi.org/10.1016/j.meegid.2019.104133
Santos N, Lima RC, Pereira CF, Gouvea V (1998) Detection of rotavirus types G8 and G10 among Brazilian children with diarrhea. J Clin Microbiol 36:2727–2729. https://doi.org/10.1128/JCM.36.9.2727-2729.1998
Volotão EM, Soares CC, Maranhão AG, Rocha LN, Hoshino Y, Santos N (2006) Rotavirus surveillance in the city of Rio de Janeiro-Brazil during 2000–2004: detection of unusual strains with G8P[4] or G10P[9] specificities. J Med Virol 78:263–272. https://doi.org/10.1002/jmv.20535
Luchs A, Timenetsky MC (2014) Unexpected detection of bovine G10 rotavirus in a Brazilian child with diarrhea. J Clin Virol 59:74–76. https://doi.org/10.1016/j.jcv.2013.11.001
Luchs A, Timenetsky MDCST (2014) G8P[6] rotaviruses isolated from Amerindian children in Mato Grosso do Sul, Brazil, during 2009: close relationship of the G and P genes with those of bovine and bat strains. J Gen Virol 95:627–641. https://doi.org/10.1099/vir.0.058099-0
Acknowledgements
The authors are grateful for the assistance of Luz Alba M. G. Fornells Arentz and Camila Correia Banks da Rocha with bovine sample collection.
Funding
This study was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq grants 404984/2018–5 and 301469/2018–0), and the Fundação Carlos Chagas de Amparo à Pesquisa do Estado do Rio de Janeiro, Brazil (FAPERJ, grant E-26/202.909/2017).
Author information
Authors and Affiliations
Contributions
A. R. M. M. was responsible for sample analysis and data interpretation, and drafted the manuscript. G. S. M. contributed to methodology, sample analysis, and the reviewing and editing of the manuscript. N. S. was responsible for the conception and design of the study, contributed substantially to data analysis and interpretation, and critically revised the manuscript for important intellectual content. The final manuscript has been reviewed and approved by all the authors.
Corresponding author
Ethics declarations
Ethical approval.
The study was approved by the ethics committee on animal research of the Empresa de Pesquisa Agropecuária do Estado do Rio de Janeiro (PESAGRO, Rio de Janeiro, RJ, Brazil), process number 1/2014.
Conflict of interest
The authors declare no conflict of interest.
Disclaimer.
The funders were not involved in the study design, data collection, data interpretation, or the decision to submit the work for publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Miranda, A.R.M., da Silva Mendes, G. & Santos, N. Rotaviruses A and C in dairy cattle in the state of Rio de Janeiro, Brazil. Braz J Microbiol 53, 1657–1663 (2022). https://doi.org/10.1007/s42770-022-00764-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-022-00764-8