Abstract
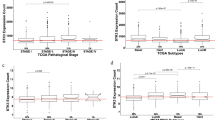
The inhibition of KIF18A selectively reduces the viability of chromosomally unstable cancers due to increased mitotic vulnerability. KIF18A expression was also reported to be upregulated and associated with tumor aggressiveness in certain cancer types including breast cancer. Here, I first showed that KIF18A mRNA expression is higher in triple-negative breast cancer (TNBC) than in non-TNBC. I also found that ER (estrogen receptor)-negative and PR (progesterone receptor)-negative breast cancer cells have higher KIF18A mRNA expression compared to ER-positive and PR-positive breast cancer cells, respectively. In contrast, HER2-positive breast tumors have higher KIF18A expression compared to HER2-negative breast tumors. In terms of PAM50 breast cancer subtypes, KIF18A transcript levels were found to be the highest in basal-like breast cancer, followed by HER2-enriched, luminal B, normal-like and luminal A. Besides, in non-TNBC, cells with high AR (androgen receptor) mRNA expression have higher KIF18A mRNA expression than cells with low AR mRNA expression. Both non-TNBC and TNBC cells with high BRCA1 and BRCA2 mRNA expression levels were observed to have higher KIF18A mRNA expression than those with low BRCA1 and BRCA2 mRNA expression levels, respectively. Combined, this study demonstrates that breast tumors with low and high expression of ER, PR, HER2, AR and BRCA1/2 have differential transcript levels of KIF18A, pointing that KIF18A might contribute to the molecular differences between different breast cancer subtypes.





Similar content being viewed by others
Data availability
Data used in the present study is publicly available, for which resources were given in the Methods section.
References
Alfarsi, L. H., Elansari, R., Toss, M. S., Diez-Rodriguez, M., Nolan, C. C., Ellis, I. O., Rakha, E. A., & Green, A. R. (2019). Kinesin family member-18A (KIF18A) is a predictive biomarker of poor benefit from endocrine therapy in early ER+ breast cancer. Breast Cancer Research and Treatment, 173(1), 93–102. https://doi.org/10.1007/s10549-018-4978-5. Epub 2018 Oct 10. PMID: 30306428.
Belete, A. M., Aynalem, Y. A., Gemeda, B. N., Demelew, T. M., & Shiferaw, W. S. (2022). The effect of estrogen receptor status on survival in breast cancer patients in Ethiopia. Retrospective Cohort Study. Breast Cancer (Dove Med Press), 14, 153–161. https://doi.org/10.2147/BCTT.S365295.
Berkel, C. (2023). Estrogen receptor- and progesterone receptor-positive breast tumors have higher mRNA levels of NR3C1 and ZBTB16, with implications in prognosis for luminal A subtype. Human Cell. https://doi.org/10.1007/s13577-023-01014-1. Epub ahead of print. PMID: 37999919.
Berkel, C., & Cacan, E. (2021). Involvement of ATMIN-DYNLL1-MRN axis in the progression and aggressiveness of serous ovarian cancer. Biochemical and Biophysical Research Communications, 17(570), 74–81. https://doi.org/10.1016/j.bbrc.2021.07.004. Epub 2021 Jul 14. PMID: 34273621.
Berkel, C., & Cacan, E. (2023a). Lower expression of NINJ1 (Ninjurin 1), a mediator of plasma membrane rupture, is associated with advanced disease and worse prognosis in serous ovarian cancer. Immunologic Research, 71(1), 15–28. https://doi.org/10.1007/s12026-022-09323-7. Epub 2022 Oct 3. PMID: 36184655.
Berkel, C., & Cacan, E. (2023b). The expression of O-linked glycosyltransferase GALNT7 in breast cancer is dependent on estrogen-, progesterone-, and HER2-receptor status, with prognostic implications. Glycoconjugate Journal. https://doi.org/10.1007/s10719-023-10137-4. Epub ahead of print. PMID: 37947928.
Chang, H. J., Yang, U. C., Lai, M. Y., Chen, C. H., & Fann, Y. C. (2022). High BRCA1 gene expression increases the risk of early distant metastasis in ER+ breast cancers. Science and Reports, 12(1), 77. https://doi.org/10.1038/s41598-021-03471-w. PMID: 34996912; PMCID: PMC8741892.
Cohen-Sharir, Y., McFarland, J. M., Abdusamad, M., Marquis, C., Bernhard, S. V., Kazachkova, M., Tang, H., Ippolito, M. R., Laue, K., Zerbib, J., Malaby, H. L. H., Jones, A., Stautmeister, L. M., Bockaj, I., Wardenaar, R., Lyons, N., Nagaraja, A., Bass, A. J., Spierings, D. C. J., … Ben-David, U. (2021). Aneuploidy renders cancer cells vulnerable to mitotic checkpoint inhibition. Nature, 590(7846), 486–491. https://doi.org/10.1038/s41586-020-03114-6. Epub 2021 Jan 27. PMID: 33505028; PMCID: PMC8262644.
Curtis, C., Shah, S. P., Chin, S. F., Turashvili, G., Rueda, O. M., Dunning, M. J., Speed, D., Lynch, A. G., Samarajiwa, S., Yuan, Y., Gräf, S., Ha, G., Haffari, G., Bashashati, A., Russell, R., McKinney, S., METABRIC Group, Langerød, A., Green, A., … Aparicio, S. (2012). The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature, 486(7403), 346–352. https://doi.org/10.1038/nature10983. PMID: 22522925; PMCID: PMC3440846.
Derakhshan, F., & Reis-Filho, J. S. (2022). Pathogenesis of triple-negative breast cancer. Annual Review of Pathology: Mechanisms of Disease, 24(17), 181–204. https://doi.org/10.1146/annurev-pathol-042420-093238. PMID: 35073169; PMCID: PMC9231507.
Fonseca, C. L., Malaby, H. L. H., Sepaniac, L. A., Martin, W., Byers, C., Czechanski, A., Messinger, D., Tang, M., Ohi, R., Reinholdt, L. G., & Stumpff, J. (2019). Mitotic chromosome alignment ensures mitotic fidelity by promoting interchromosomal compaction during anaphase. Journal of Cell Biology, 218(4), 1148–1163. https://doi.org/10.1083/jcb.201807228. Epub 2019 Feb 7. PMID: 30733233; PMCID: PMC6446859.
Goldman, M. J., Craft, B., Hastie, M., Repečka, K., McDade, F., Kamath, A., Banerjee, A., Luo, Y., Rogers, D., Brooks, A. N., Zhu, J., & Haussler, D. (2020). Visualizing and interpreting cancer genomics data via the Xena platform. Nature Biotechnology, 38(6), 675–678. https://doi.org/10.1038/s41587-020-0546-8. PMID: 32444850; PMCID: PMC7386072.
Gorodetska, I., Kozeretska, I., & Dubrovska, A. (2019). BRCA genes: the role in genome stability, cancer stemness and therapy resistance. Journal of Cancer, 10(9), 2109–2127. https://doi.org/10.7150/jca.30410. PMID: 31205572; PMCID: PMC6548160.
Greenup, R., Buchanan, A., Lorizio, W., Rhoads, K., Chan, S., Leedom, T., King, R., McLennan, J., Crawford, B., Kelly Marcom, P., & Shelley, H. E. (2013). Prevalence of BRCA mutations among women with triple-negative breast cancer (TNBC) in a genetic counseling cohort. Annals of Surgical Oncology, 20(10), 3254–3258. https://doi.org/10.1245/s10434-013-3205-1. Epub 2013 Aug 22. PMID: 23975317.
Gudmundsdottir, K., & Ashworth, A. (2006). The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene, 25(43), 5864–5874. https://doi.org/10.1038/sj.onc.1209874. PMID: 16998501.
Győrffy, B. (2021). Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Computational and Structural Biotechnology Journal, 18(19), 4101–4109. https://doi.org/10.1016/j.csbj.2021.07.014. PMID: 34527184; PMCID: PMC8339292.
Hitti, E., Bakheet, T., Al-Souhibani, N., Moghrabi, W., Al-Yahya, S., Al-Ghamdi, M., Al-Saif, M., Shoukri, M. M., Lánczky, A., Grépin, R., Győrffy, B., Pagès, G., & Khabar, K. S. (2016). Systematic analysis of AU-rich element expression in cancer reveals common functional clusters regulated by key RNA-binding proteins. Cancer Research, 76(14), 4068–4080. https://doi.org/10.1158/0008-5472.CAN-15-3110. Epub 2016 May 17. PMID: 27197193.
Huszno, J., Kołosza, Z., & Grzybowska, E. (2019). BRCA1 mutation in breast cancer patients: Analysis of prognostic factors and survival. Oncology Letters, 17(2), 1986–1995. https://doi.org/10.3892/ol.2018.9770. Epub 2018 Nov 28. PMID: 30675265; PMCID: PMC6341769.
Jin, T. Y., Park, K. S., Nam, S. E., Yoo, Y. B., Park, W. S., & Yun, I. J. (2022). BRCA1/2 serves as a biomarker for poor prognosis in breast carcinoma. International Journal of Molecular Sciences, 23(7), 3754. https://doi.org/10.3390/ijms23073754. PMID: 35409110; PMCID: PMC8998777.
Kasahara, M., Nagahara, M., Nakagawa, T., Ishikawa, T., Sato, T., Uetake, H., & Sugihara, K. (2016). Clinicopathological relevance of kinesin family member 18A expression in invasive breast cancer. Oncology Letters, 12(3), 1909–1914. https://doi.org/10.3892/ol.2016.4823. Epub 2016 Jul 7. PMID: 27588139; PMCID: PMC4998100.
Kassambara, A. (2023). ggpubr: 'ggplot2' based publication ready plots. R package version 0.6.0. https://CRAN.R-project.org/package=ggpubr. Accessed 26 Dec 2023.
Kensler, K. H., Sankar, V. N., Wang, J., Zhang, X., Rubadue, C. A., Baker, G. M., Parker, J. S., Hoadley, K. A., Stancu, A. L., Pyle, M. E., Collins, L. C., Hunter, D. J., Eliassen, A. H., Hankinson, S. E., Tamimi, R. M., & Heng, Y. J. (2019). PAM50 molecular intrinsic subtypes in the nurses’ health study cohorts. Cancer Epidemiology, Biomarkers & Prevention, 28(4), 798–806. https://doi.org/10.1158/1055-9965.EPI-18-0863. Epub 2018 Dec 27. PMID: 30591591; PMCID: PMC6449178.
Lal, A., Ramazzotti, D., Weng, Z., Liu, K., Ford, J. M., & Sidow, A. (2019). Comprehensive genomic characterization of breast tumors with BRCA1 and BRCA2 mutations. BMC Medical Genomics, 12(1), 84. https://doi.org/10.1186/s12920-019-0545-0. PMID: 31182087; PMCID: PMC6558765.
Li, T. F., Zeng, H. J., Shan, Z., Ye, R. Y., Cheang, T. Y., Zhang, Y. J., Lu, S. H., Zhang, Q., Shao, N., & Lin, Y. (2020). Overexpression of kinesin superfamily members as prognostic biomarkers of breast cancer. Cancer Cell International, 15(20), 123. https://doi.org/10.1186/s12935-020-01191-1. PMID: 32322170; PMCID: PMC7161125.
Marquis, C., Fonseca, C. L., Queen, K. A., Wood, L., Vandal, S. E., Malaby, H. L. H., Clayton, J. E., & Stumpff, J. (2021). Chromosomally unstable tumor cells specifically require KIF18A for proliferation. Nature Communications, 12(1), 1213. https://doi.org/10.1038/s41467-021-21447-2. PMID: 33619254; PMCID: PMC7900194.
Mayr, M. I., Hümmer, S., Bormann, J., Grüner, T., Adio, S., Woehlke, G., & Mayer, T. U. (2007). The human kinesin Kif18A is a motile microtubule depolymerase essential for chromosome congression. Current Biology, 17(6), 488–498. https://doi.org/10.1016/j.cub.2007.02.036. Epub 2007 Mar 8. PMID: 17346968.
Morgan, M., Obenchain, V., Hester, J., & Pagès, H. (2022). SummarizedExperiment: SummarizedExperiment container. R package version 1.26.1. https://bioconductor.org/packages/SummarizedExperiment. Accessed 26 Dec 2023.
Morgan, M., Shepherd, L. (2022a). AnnotationHub: Client to access AnnotationHub resources. R package version 3.4.0.
Morgan, M., Shepherd, L. (2022b). ExperimentHub: Client to access ExperimentHub resources. R package version 2.4.0.
Ooms, J. (2023). magick: Advanced graphics and image-processing in R. R package version 2.7.4. https://CRAN.R-project.org/package=magick. Accessed 26 Dec 2023.
Pareja, F., Geyer, F. C., Marchiò, C., Burke, K. A., Weigelt, B., & Reis-Filho, J. S. (2016). Triple-negative breast cancer: The importance of molecular and histologic subtyping, and recognition of low-grade variants. NPJ Breast Cancer, 16(2), 16036. https://doi.org/10.1038/npjbcancer.2016.36. PMID: 28721389; PMCID: PMC5515338.
Patel, A., Unni, N., & Peng, Y. (2020). The Changing paradigm for the treatment of HER2-positive breast cancer. Cancers (basel), 12(8), 2081. https://doi.org/10.3390/cancers12082081. PMID: 32731409; PMCID: PMC7464074.
Payton, M., Belmontes, B., Hanestad, K., Moriguchi, J., Chen, K., McCarter, J. D., Chung, G., Ninniri, M. S., Sun, J., Manoukian, R., Chambers, S., Ho, S. M., Kurzeja, R. J. M., Edson, K. Z., Dahal, U. P., Wu, T., Wannberg, S., Beltran, P. J., Canon, J., … Hughes, P. E. (2023). Small-molecule inhibition of kinesin KIF18A reveals a mitotic vulnerability enriched in chromosomally unstable cancers. Nature Cancer. https://doi.org/10.1038/s43018-023-00699-5. Epub ahead of print. PMID: 38151625.
Quinton, R. J., DiDomizio, A., Vittoria, M. A., Kotýnková, K., Ticas, C. J., Patel, S., Koga, Y., Vakhshoorzadeh, J., Hermance, N., Kuroda, T. S., Parulekar, N., Taylor, A. M., Manning, A. L., Campbell, J. D., & Ganem, N. J. (2021). Whole-genome doubling confers unique genetic vulnerabilities on tumour cells. Nature, 590(7846), 492–497. https://doi.org/10.1038/s41586-020-03133-3. Epub 2021 Jan 27. Erratum in: Nature. 2021 May;593(7860):E15. PMID: 33505027; PMCID: PMC7889737.
R Core Team. (2022). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
Rath, O., & Kozielski, F. (2012). Kinesins and cancer. Nature Reviews Cancer, 12(8), 527–539. https://doi.org/10.1038/nrc3310. PMID: 22825217.
Ravaioli, S., Maltoni, R., Pasculli, B., Parrella, P., Giudetti, A. M., Vergara, D., Tumedei, M. M., Pirini, F., & Bravaccini, S. (2022). Androgen receptor in breast cancer: the “5W” questions. Frontiers in Endocrinology (lausanne)., 30(13), 977331. https://doi.org/10.3389/fendo.2022.977331. PMID: 36111296; PMCID: PMC9468319.
Royston, P. (1995). Remark AS R94: a remark on algorithm AS 181: The WW test for normality. Journal of Applied Statistics, 44, 547–551. https://doi.org/10.2307/2986146
Savci-Heijink, C. D., Halfwerk, H., Koster, J., Horlings, H. M., & van de Vijver, M. J. (2019). A specific gene expression signature for visceral organ metastasis in breast cancer. BMC Cancer, 19(1), 333. https://doi.org/10.1186/s12885-019-5554-z. PMID: 30961553; PMCID: PMC6454625.
Stumpff, J., von Dassow, G., Wagenbach, M., Asbury, C., & Wordeman, L. (2008). The kinesin-8 motor Kif18A suppresses kinetochore movements to control mitotic chromosome alignment. Developmental Cell, 14(2), 252–262. https://doi.org/10.1016/j.devcel.2007.11.014. PMID: 18267093; PMCID: PMC2267861.
Stumpff, J., Wagenbach, M., Franck, A., Asbury, C. L., & Wordeman, L. (2012). Kif18A and chromokinesins confine centromere movements via microtubule growth suppression and spatial control of kinetochore tension. Developmental Cell, 22(5), 1017–1029. https://doi.org/10.1016/j.devcel.2012.02.013. PMID: 22595673; PMCID: PMC3356572.
TečićVuger, A., Šeparović, R., Vazdar, L., Pavlović, M., Lepetić, P., Šitić, S., Bajić, Ž, Šarčević, B., & Vrbanec, D. (2020). Characteristics and prognosis of triple-negative breast cancer patients: a croatian single institution retrospective cohort study. Acta Clinica Croatica, 59(1), 97–108. https://doi.org/10.20471/acc.2020.59.01.12. PMID: 32724280; PMCID: PMC7382886.
Turner, N. C., & Reis-Filho, J. S. (2013). Tackling the diversity of triple-negative breast cancer. Clinical Cancer Research, 19(23), 6380–6388. https://doi.org/10.1158/1078-0432. CCR-13-0915. PMID: 24298068.
Wang, Z., Zhang, J., Zhang, Y., Deng, Q., & Liang, H. (2018). Expression and mutations of BRCA in breast cancer and ovarian cancer: Evidence from bioinformatics analyses. International Journal of Molecular Medicine, 42(6), 3542–3550. https://doi.org/10.3892/ijmm.2018.3870. Epub 2018 Sep 11. PMID: 30221688.
Wickham, H., Averick, M., Bryan, J., Chang, W., McGowan, L. D., François, R., Grolemund, G., Hayes, A., Henry, L., Hester, J., Kuhn, M., Pedersen, T. L., Miller, E., Bache, S. M., Müller, K., Ooms, J., Robinson, D., Seidel, D. P., Spinu, V., … Yutani, H. (2019). Welcome to the tidyverse. Journal of Open Source Software, 4(43), 1686. https://doi.org/10.21105/joss.01686
You, C. P., Tsoi, H., Man, E. P. S., Leung, M. H., & Khoo, U. S. (2022). Modulating the activity of androgen receptor for treating breast cancer. International Journal of Molecular Sciences, 23(23), 15342. https://doi.org/10.3390/ijms232315342. PMID: 36499670; PMCID: PMC9739178.
Zhang, C., Zhu, C., Chen, H., Li, L., Guo, L., Jiang, W., & Lu, S. H. (2010). Kif18A is involved in human breast carcinogenesis. Carcinogenesis, 31(9), 1676–1684. https://doi.org/10.1093/carcin/bgq134. Epub 2010 Jul 1. PMID: 20595236.
Zou, J. X., Duan, Z., Wang, J., Sokolov, A., Xu, J., Chen, C. Z., Li, J. J., & Chen, H. W. (2014). Kinesin family deregulation coordinated by bromodomain protein ANCCA and histone methyltransferase MLL for breast cancer cell growth, survival, and tamoxifen resistance. Molecular Cancer Research, 12(4), 539–549. https://doi.org/10.1158/1541-7786.MCR-13-0459. Epub 2014 Jan 3. PMID: 24391143; PMCID: PMC4139106.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
42764_2024_126_MOESM1_ESM.pdf
Supplementary Figure 1. Kaplan-Meier curves showing overall survival (OS) for breast cancer patients with low and high expression of KIF18A. Left: all subtypes combined. Right: HER2-negative. HR: Hazard ratio. Data from [Győrffy, 2021] (PDF 971 KB)
42764_2024_126_MOESM2_ESM.pdf
Supplementary Figure 2 KIF18A mRNA expression levels based on ER-, PR- and HER2 status analysed using data from another dataset to validate findings from TCGA dataset. Data from METABRIC (Molecular Taxonomy of Breast Cancer International Consortium), accessed via https://www.mercuriolab.umassmed.edu/metabric [Curtis et al., 2012] (PDF 3930 KB)
42764_2024_126_MOESM3_ESM.pdf
Supplementary Figure 3 KIF18A mRNA and protein levels from cell lines based on breast cancer subtypes and BRCA1/2 mutation status. Data analysed using Dependency Map (DepMap) portal (https://depmap.org/portal/interactive/) (PDF 219 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Berkel, C. KIF18A as a potential biomarker to distinguish different breast cancer subtypes based on receptor status. GENOME INSTAB. DIS. 5, 89–96 (2024). https://doi.org/10.1007/s42764-024-00126-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42764-024-00126-8