Abstract
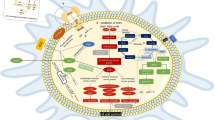
Tuberculosis (TB), caused by Mycobacterium tuberculosis (Mtb), remains a serious global health problem that kills over 1 million people annually. Understanding the pathogenesis of TB and the emergence of drug-resistant strains of Mtb is a priority for the development of strategies against TB. Its DNA damage and repair systems are essential for maintaining genome stability in Mtb. Their aberrant work leads to hypermutability and is often associated with the emergence of resistant bacteria. On the other hand, Mtb infection also induces genome instability of host cells, which are involved in the pathogenesis of TB, including the formation of giant cells. Collectively, this review is an attempt to summarize our current understanding of the role of genome instability in the pathogenesis of TB and to shed light on the development of new strategies for TB treatment.

Similar content being viewed by others
References
Ackermann, M. (2015). A functional perspective on phenotypic heterogeneity in microorganisms. Nature Reviews Microbiology, 13(8), 497–508. https://doi.org/10.1038/nrmicro3491
Adams, L. B., Dinauer, M. C., Morgenstern, D. E., & Krahenbuhl, J. L. (1997). Comparison of the roles of reactive oxygen and nitrogen intermediates in the host response to Mycobacterium tuberculosis using transgenic mice. Tubercle and Lung Disease, 78(5–6), 237–246. https://doi.org/10.1016/s0962-8479(97)90004-6
Amundsen, S. K., Spicer, T., Karabulut, A. C., Londono, L. M., Eberhart, C., Fernandez Vega, V., & Smith, G. R. (2012). Small-molecule inhibitors of bacterial AddAB and RecBCD helicase-nuclease DNA repair enzymes. ACS Chemical Biology, 7(5), 879–891. https://doi.org/10.1021/cb300018x
Baharoglu, Z., & Mazel, D. (2014). SOS, the formidable strategy of bacteria against aggressions. FEMS Microbiology Reviews, 38(6), 1126–1145. https://doi.org/10.1111/1574-6976.12077
Balaban, N. Q., Helaine, S., Lewis, K., Ackermann, M., Aldridge, B., Andersson, D. I., & Zinkernagel, A. (2019). Publisher correction: Definitions and guidelines for research on antibiotic persistence. Nature Reviews Microbiology, 17(7), 460. https://doi.org/10.1038/s41579-019-0207-4
Balganesh, M., Dinesh, N., Sharma, S., Kuruppath, S., Nair, A. V., & Sharma, U. (2012). Efflux pumps of Mycobacterium tuberculosis play a significant role in antituberculosis activity of potential drug candidates. Antimicrobial Agents and Chemotherapy, 56(5), 2643–2651. https://doi.org/10.1128/AAC.06003-11
Belenky, P., Ye, J. D., Porter, C. B., Cohen, N. R., Lobritz, M. A., Ferrante, T., & Collins, J. J. (2015). Bactericidal antibiotics induce toxic metabolic perturbations that lead to cellular damage. Cell Reports, 13(5), 968–980. https://doi.org/10.1016/j.celrep.2015.09.059
Ben-David, U. (2015). Genomic instability, driver genes and cell selection: Projections from cancer to stem cells. Biochimica Et Biophysica Acta, 1849(4), 427–435. https://doi.org/10.1016/j.bbagrm.2014.08.005
Booth, J. A., Spirek, M., Lobie, T. A., Skarstad, K., Krejci, L., & Bjoras, M. (2020). Antibiotic-induced DNA damage results in a controlled loss of pH homeostasis and genome instability. Science and Reports, 10(1), 19422. https://doi.org/10.1038/s41598-020-76426-2
Brauner, A., Fridman, O., Gefen, O., & Balaban, N. Q. (2016). Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nature Reviews Microbiology, 14(5), 320–330. https://doi.org/10.1038/nrmicro.2016.34
Brissett, N. C., Pitcher, R. S., Juarez, R., Picher, A. J., Green, A. J., Dafforn, T. R., & Doherty, A. J. (2007). Structure of a NHEJ polymerase-mediated DNA synaptic complex. Science, 318(5849), 456–459. https://doi.org/10.1126/science.1145112
Cadet, J., Douki, T., & Ravanat, J. L. (2010). Oxidatively generated base damage to cellular DNA. Free Radical Biology and Medicine, 49(1), 9–21. https://doi.org/10.1016/j.freeradbiomed.2010.03.025
Cadet, J., & Wagner, J. R. (2014). Oxidatively generated base damage to cellular DNA by hydroxyl radical and one-electron oxidants: Similarities and differences. Archives of Biochemistry and Biophysics, 557, 47–54. https://doi.org/10.1016/j.abb.2014.05.001
Castaneda-Garcia, A., Prieto, A. I., Rodriguez-Beltran, J., Alonso, N., Cantillon, D., Costas, C., & Blazquez, J. (2017). A non-canonical mismatch repair pathway in prokaryotes. Nature Communications, 8, 14246. https://doi.org/10.1038/ncomms14246
Cebrian-Sastre, E., Martin-Blecua, I., Gullon, S., Blazquez, J., & Castaneda-Garcia, A. (2021). Control of genome stability by EndoMS/NucS-mediated non-canonical mismatch repair. Cells. https://doi.org/10.3390/cells10061314
Chakaya, J., Khan, M., Ntoumi, F., Aklillu, E., Fatima, R., Mwaba, P., & Zumla, A. (2021). Global tuberculosis report 2020—reflections on the global TB burden, treatment and prevention efforts. International Journal of Infectious Diseases. https://doi.org/10.1016/j.ijid.2021.02.107
Chen, Y., Jiang, Q., Zou, J., Yang, T., Liu, Q., Luo, G., & Gao, Q. (2021). Deep whole-genome sequencing reveals no evidence for heteroresistance influencing treatment outcomes among drug-susceptible tuberculosis patients. Tuberculosis (edinburgh, Scotland), 130, 102120. https://doi.org/10.1016/j.tube.2021.102120
Choudhary, E., Sharma, R., Kumar, Y., & Agarwal, N. (2019). Conditional Silencing by CRISPRi reveals the role of DNA gyrase in formation of drug-tolerant persister population in Mycobacterium tuberculosis. Frontiers in Cellular and Infection Microbiology, 9, 70. https://doi.org/10.3389/fcimb.2019.00070
Cole, S. T., Brosch, R., Parkhill, J., Garnier, T., Churcher, C., Harris, D., & Barrell, B. G. (1998). Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature, 393(6685), 537–544. https://doi.org/10.1038/31159
da Silva, A. L., Bresciani, M. J., Karnopp, T. E., Weber, A. F., Ellwanger, J. H., Henriques, J. A., & Possuelo, L. G. (2015). DNA damage and cellular abnormalities in tuberculosis, lung cancer and chronic obstructive pulmonary disease. Multidisciplinary Respiratory Medicine, 10, 38. https://doi.org/10.1186/s40248-015-0034-z
Darwin, K. H., Ehrt, S., Gutierrez-Ramos, J. C., Weich, N., & Nathan, C. F. (2003). The proteasome of Mycobacterium tuberculosis is required for resistance to nitric oxide. Science, 302(5652), 1963–1966. https://doi.org/10.1126/science.1091176
Darwin, K. H., & Nathan, C. F. (2005). Role for nucleotide excision repair in virulence of Mycobacterium tuberculosis. Infection and Immunity, 73(8), 4581–4587. https://doi.org/10.1128/IAI.73.8.4581-4587.2005
Della, M., Palmbos, P. L., Tseng, H. M., Tonkin, L. M., Daley, J. M., Topper, L. M., & Doherty, A. J. (2004). Mycobacterial Ku and ligase proteins constitute a two-component NHEJ repair machine. Science, 306(5696), 683–685. https://doi.org/10.1126/science.1099824
Dhar, N., McKinney, J., & Manina, G. (2016). Phenotypic heterogeneity in Mycobacterium tuberculosis. Microbiology Spectrum. https://doi.org/10.1128/microbiolspec.TBTB2-0021-2016
Dos Vultos, T., Mestre, O., Rauzier, J., Golec, M., Rastogi, N., Rasolofo, V., & Gicquel, B. (2008). Evolution and diversity of clonal bacteria: The paradigm of Mycobacterium tuberculosis. PLoS ONE, 3(2), e1538. https://doi.org/10.1371/journal.pone.0001538
Dos Vultos, T., Mestre, O., Tonjum, T., & Gicquel, B. (2009). DNA repair in Mycobacterium tuberculosis revisited. FEMS Microbiology Reviews, 33(3), 471–487. https://doi.org/10.1111/j.1574-6976.2009.00170.x
Dutta, N. K., Mehra, S., Didier, P. J., Roy, C. J., Doyle, L. A., Alvarez, X., & Kaushal, D. (2010). Genetic requirements for the survival of tubercle bacilli in primates. Journal of Infectious Diseases, 201(11), 1743–1752. https://doi.org/10.1086/652497
Ehrt, S., & Schnappinger, D. (2009). Mycobacterial survival strategies in the phagosome: Defence against host stresses. Cellular Microbiology, 11(8), 1170–1178. https://doi.org/10.1111/j.1462-5822.2009.01335.x
Fan, X. Y., Tang, B. K., Xu, Y. Y., Han, A. X., Shi, K. X., Wu, Y. K., & Lyu, L. D. (2018). Oxidation of dCTP contributes to antibiotic lethality in stationary-phase mycobacteria. Proceedings of the National Academy of Sciences of the United States of America, 115(9), 2210–2215. https://doi.org/10.1073/pnas.1719627115
Flentie, K., Garner, A. L., & Stallings, C. L. (2016). Mycobacterium tuberculosis transcription machinery: Ready to respond to host attacks. Journal of Bacteriology, 198(9), 1360–1373. https://doi.org/10.1128/JB.00935-15
Foti, J. J., Devadoss, B., Winkler, J. A., Collins, J. J., & Walker, G. C. (2012). Oxidation of the guanine nucleotide pool underlies cell death by bactericidal antibiotics. Science, 336(6079), 315–319. https://doi.org/10.1126/science.1219192
Gorna, A. E., Bowater, R. P., & Dziadek, J. (2010). DNA repair systems and the pathogenesis of Mycobacterium tuberculosis: Varying activities at different stages of infection. Clinical Science (london, England), 119(5), 187–202. https://doi.org/10.1042/CS20100041
Grossman, L., & Thiagalingam, S. (1993). Nucleotide excision repair, a tracking mechanism in search of damage. Journal of Biological Chemistry, 268(23), 16871–16874. Accessed from https://www.ncbi.nlm.nih.gov/pubmed/8349576
Gupta, R., Barkan, D., Redelman-Sidi, G., Shuman, S., & Glickman, M. S. (2011). Mycobacteria exploit three genetically distinct DNA double-strand break repair pathways. Molecular Microbiology, 79(2), 316–330. https://doi.org/10.1111/j.1365-2958.2010.07463.x.
Gutierrez, A., Laureti, L., Crussard, S., Abida, H., Rodriguez-Rojas, A., Blazquez, J., & Matic, I. (2013). Beta-Lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity. Nature Communications, 4, 1610. https://doi.org/10.1038/ncomms2607
Hans, S., Purkait, D., Nandan, S., Bansal, M., Hameed, S., & Fatima, Z. (2020). Rec A disruption unveils cross talk between DNA repair and membrane damage, efflux pump activity, biofilm formation in Mycobacterium smegmatis. Microbial Pathogenesis, 149, 104262. https://doi.org/10.1016/j.micpath.2020.104262
Heaton, B. E., Barkan, D., Bongiorno, P., Karakousis, P. C., & Glickman, M. S. (2014). Deficiency of double-strand DNA break repair does not impair Mycobacterium tuberculosis virulence in multiple animal models of infection. Infection and Immunity, 82(8), 3177–3185. https://doi.org/10.1128/IAI.01540-14
Herrtwich, L., Nanda, I., Evangelou, K., Nikolova, T., Horn, V., Sagar, A., & Triantafyllopoulou, A. (2016). DNA damage signaling instructs polyploid macrophage fate in granulomas. Cell, 167(5), 1264–1280. https://doi.org/10.1016/j.cell.2016.09.054
Jadaun, A., Sudhakar, D. R., Subbarao, N., & Dixit, A. (2015). In silico screening for novel inhibitors of DNA polymerase III alpha subunit of Mycobacterium tuberculosis (MtbDnaE2, H37Rv). PLoS ONE, 10(3), e0119760. https://doi.org/10.1371/journal.pone.0119760
Jain, R., Kumar, P., & Varshney, U. (2007). A distinct role of formamidopyrimidine DNA glycosylase (MutM) in down-regulation of accumulation of G, C mutations and protection against oxidative stress in mycobacteria. DNA Repair (amst), 6(12), 1774–1785. https://doi.org/10.1016/j.dnarep.2007.06.009
Kha, D. T., Wang, G., Natrajan, N., Harrison, L., & Vasquez, K. M. (2010). Pathways for double-strand break repair in genetically unstable Z-DNA-forming sequences. Journal of Molecular Biology, 398(4), 471–480. https://doi.org/10.1016/j.jmb.2010.03.035
Khanam, T., Afsar, M., Shukla, A., Alam, F., Kumar, S., Soyar, H., & Ramachandran, R. (2020). M. tuberculosis class II apurinic/apyrimidinic-endonuclease/3’-5’ exonuclease (XthA) engages with NAD+-dependent DNA ligase A (LigA) to counter futile cleavage and ligation cycles in base excision repair. Nucleic Acids Research, 48(8), 4325–4343. https://doi.org/10.1093/nar/gkaa188
Kisker, C., Kuper, J., & Van Houten, B. (2013). Prokaryotic nucleotide excision repair. Cold Spring Harbor Perspectives in Biology, 5(3), a012591. https://doi.org/10.1101/cshperspect.a012591
Kling, A., Lukat, P., Almeida, D. V., Bauer, A., Fontaine, E., Sordello, S., & Muller, R. (2015). Antibiotics. Targeting DnaN for tuberculosis therapy using novel griselimycins. Science, 348(6239), 1106–1112. https://doi.org/10.1126/science.aaa4690
Kohanski, M. A., Dwyer, D. J., Hayete, B., Lawrence, C. A., & Collins, J. J. (2007). A common mechanism of cellular death induced by bactericidal antibiotics. Cell, 130(5), 797–810. https://doi.org/10.1016/j.cell.2007.06.049
Krokan, H. E., Drablos, F., & Slupphaug, G. (2002). Uracil in DNA–occurrence, consequences and repair. Oncogene, 21(58), 8935–8948. https://doi.org/10.1038/sj.onc.1205996
Kurahashi, H., Inagaki, H., Ohye, T., Kogo, H., Kato, T., & Emanuel, B. S. (2006). Palindrome-mediated chromosomal translocations in humans. DNA Repair (amst), 5(9–10), 1136–1145. https://doi.org/10.1016/j.dnarep.2006.05.035
Kurthkoti, K., Srinath, T., Kumar, P., Malshetty, V. S., Sang, P. B., Jain, R., & Varshney, U. (2010). A distinct physiological role of MutY in mutation prevention in mycobacteria. Microbiology (reading), 156(Pt 1), 88–93. https://doi.org/10.1099/mic.0.033621-0
Kurthkoti, K., & Varshney, U. (2010). Detrimental effects of hypoxia-specific expression of uracil DNA glycosylase (Ung) in Mycobacterium smegmatis. Journal of Bacteriology, 192(24), 6439–6446. https://doi.org/10.1128/JB.00679-10
Kurthkoti, K., & Varshney, U. (2011). Base excision and nucleotide excision repair pathways in mycobacteria. Tuberculosis (edinburgh, Scotland), 91(6), 533–543. https://doi.org/10.1016/j.tube.2011.06.005
Lahiri, S., Rizzi, M., Rossi, F., & Miggiano, R. (2018). Mycobacterium tuberculosis UvrB forms dimers in solution and interacts with UvrA in the absence of ligands. Proteins, 86(1), 98–109. https://doi.org/10.1002/prot.25412
Li, Z., Pearlman, A. H., & Hsieh, P. (2016). DNA mismatch repair and the DNA damage response. DNA Repair (amst), 38, 94–101. https://doi.org/10.1016/j.dnarep.2015.11.019
Liu, Q., Liu, H., Shi, L., Gan, M., Zhao, X., Lyu, L. D., & Gao, Q. (2021). Local adaptation of Mycobacterium tuberculosis on the Tibetan Plateau. Proceedings of the National Academy of Sciences of the United States of America. https://doi.org/10.1073/pnas.2017831118
Liu, Q., Wei, J., Li, Y., Wang, M., Su, J., Lu, Y., & Gao, Q. (2020). Mycobacterium tuberculosis clinical isolates carry mutational signatures of host immune environments. Science Advances, 6(22), eaba4901. https://doi.org/10.1126/sciadv.aba4901
Lochab, S., Singh, Y., Sengupta, S., & Nandicoori, V. K. (2020). Mycobacterium tuberculosis exploits host ATM kinase for survival advantage through SecA2 secretome. eLife. https://doi.org/10.7554/eLife.51466
Lyu, L. D., Tang, B. K., Fan, X. Y., Ma, H., & Zhao, G. P. (2013). Mycobacterial MazG safeguards genetic stability via housecleaning of 5-OH-dCTP. PLoS Pathogens, 9(12), e1003814. https://doi.org/10.1371/journal.ppat.1003814
MacMicking, J. D., North, R. J., LaCourse, R., Mudgett, J. S., Shah, S. K., & Nathan, C. F. (1997). Identification of nitric oxide synthase as a protective locus against tuberculosis. Proceedings of the National Academy of Sciences of the United States of America, 94(10), 5243–5248. https://doi.org/10.1073/pnas.94.10.5243
Malshetty, V. S., Jain, R., Srinath, T., Kurthkoti, K., & Varshney, U. (2010). Synergistic effects of UdgB and Ung in mutation prevention and protection against commonly encountered DNA damaging agents in Mycobacterium smegmatis. Microbiology (reading), 156(Pt 3), 940–949. https://doi.org/10.1099/mic.0.034363-0
Manina, G., Griego, A., Singh, L. K., McKinney, J. D., & Dhar, N. (2019). Preexisting variation in DNA damage response predicts the fate of single mycobacteria under stress. EMBO Journal, 38(22), e101876. https://doi.org/10.15252/embj.2019101876
Mazloum, N., Stegman, M. A., Croteau, D. L., Van Houten, B., Kwon, N. S., Ling, Y., & Nathan, C. (2011). Identification of a chemical that inhibits the mycobacterial UvrABC complex in nucleotide excision repair. Biochemistry, 50(8), 1329–1335. https://doi.org/10.1021/bi101674c
McGrath, M., Gey van Pittius, N. C., van Helden, P. D., Warren, R. M., & Warner, D. F. (2014). Mutation rate and the emergence of drug resistance in Mycobacterium tuberculosis. Journal of Antimicrobial Chemotherapy, 69(2), 292–302. https://doi.org/10.1093/jac/dkt364
Mittal, P., Sinha, R., Kumar, A., Singh, P., Ngasainao, M. R., Singh, A., & Singh, I. K. (2020). Focusing on DNA repair and damage tolerance mechanisms in Mycobacterium tuberculosis: an emerging therapeutic theme. Current Topics in Medicinal Chemistry, 20(5), 390–408. https://doi.org/10.2174/1568026620666200110114322
Mizrahi, V., & Andersen, S. J. (1998). DNA repair in Mycobacterium tuberculosis. What have we learnt from the genome sequence? Molecular Microbiology, 29(6), 1331–1339. https://doi.org/10.1046/j.1365-2958.1998.01038.x
Mohanty, S., Dal Molin, M., Ganguli, G., Padhi, A., Jena, P., Selchow, P., & Sonawane, A. (2016). Mycobacterium tuberculosis EsxO (Rv2346c) promotes bacillary survival by inducing oxidative stress mediated genomic instability in macrophages. Tuberculosis (edinburgh, Scotland), 96, 44–57. https://doi.org/10.1016/j.tube.2015.11.006
Nandakumar, M., Nathan, C., & Rhee, K. Y. (2014). Isocitrate lyase mediates broad antibiotic tolerance in Mycobacterium tuberculosis. Nature Communications, 5, 4306. https://doi.org/10.1038/ncomms5306
Naz, S., Dabral, S., Nagarajan, S. N., Arora, D., Singh, L. V., Kumar, P., & Nandicoori, V. K. (2021). Compromised base excision repair pathway in Mycobacterium tuberculosis imparts superior adaptability in the host. PLoS Pathogens, 17(3), e1009452. https://doi.org/10.1371/journal.ppat.1009452
Pitcher, R. S., Green, A. J., Brzostek, A., Korycka-Machala, M., Dziadek, J., & Doherty, A. J. (2007). NHEJ protects mycobacteria in stationary phase against the harmful effects of desiccation. DNA Repair (amst), 6(9), 1271–1276. https://doi.org/10.1016/j.dnarep.2007.02.009
Poetsch, A. R. (2020). The genomics of oxidative DNA damage, repair, and resulting mutagenesis. Computational and Structural Biotechnology Journal, 18, 207–219. https://doi.org/10.1016/j.csbj.2019.12.013
Prammananan, T., Phunpruch, S., Jaitrong, S., & Palittapongarnpim, P. (2012). Mycobacterium tuberculosis uvrC essentiality in response to UV-induced cell damage. The Southeast Asian Journal of Tropical Medicine and Public Health, 43(2), 370–375. Accessed on https://www.ncbi.nlm.nih.gov/pubmed/23082589
Rao, V. V., Gupta, E. V., & Thomas, I. M. (1990). Chromosome damage in untreated tuberculosis patients. Tubercle, 71(3), 169–172. https://doi.org/10.1016/0041-3879(90)90070-o
Rengarajan, J., Bloom, B. R., & Rubin, E. J. (2005). Genome-wide requirements for Mycobacterium tuberculosis adaptation and survival in macrophages. Proceedings of the National Academy of Sciences of the United States of America, 102(23), 8327–8332. https://doi.org/10.1073/pnas.0503272102
Rossi, F., Khanduja, J. S., Bortoluzzi, A., Houghton, J., Sander, P., Guthlein, C., & Rizzi, M. (2011). The biological and structural characterization of Mycobacterium tuberculosis UvrA provides novel insights into its mechanism of action. Nucleic Acids Research, 39(16), 7316–7328. https://doi.org/10.1093/nar/gkr271
Sassetti, C. M., & Rubin, E. J. (2003). Genetic requirements for mycobacterial survival during infection. Proceedings of the National Academy of Sciences of the United States of America, 100(22), 12989–12994. https://doi.org/10.1073/pnas.2134250100
Selek, S., Aslan, M., Horoz, M., Celik, H., Cosar, N., Gunak, F., & Kocyigit, A. (2012). Peripheral DNA damage in active pulmonary tuberculosis. Environmental Toxicology, 27(6), 380–384. https://doi.org/10.1002/tox.20674
Singh, A. (2017). Guardians of the mycobacterial genome: A review on DNA repair systems in Mycobacterium tuberculosis. Microbiology (reading), 163(12), 1740–1758. https://doi.org/10.1099/mic.0.000578
Singh, A., Bhagavat, R., Vijayan, M., & Chandra, N. (2016). A comparative analysis of the DNA recombination repair pathway in mycobacterial genomes. Tuberculosis (edinburgh, Scotland), 99, 109–119. https://doi.org/10.1016/j.tube.2016.04.011
Sinha, K. M., Stephanou, N. C., Gao, F., Glickman, M. S., & Shuman, S. (2007). Mycobacterial UvrD1 is a Ku-dependent DNA helicase that plays a role in multiple DNA repair events, including double-strand break repair. Journal of Biological Chemistry, 282(20), 15114–15125. https://doi.org/10.1074/jbc.M701167200
Smith, I. (2003). Mycobacterium tuberculosis pathogenesis and molecular determinants of virulence. Clinical Microbiology Reviews, 16(3), 463–496. https://doi.org/10.1128/CMR.16.3.463-496.2003
Springall, L., Hughes, C. D., Simons, M., Azinas, S., Van Houten, B., & Kad, N. M. (2018). Recruitment of UvrBC complexes to UV-induced damage in the absence of UvrA increases cell survival. Nucleic Acids Research, 46(3), 1256–1265. https://doi.org/10.1093/nar/gkx1244
Thakur, M., Badugu, S., & Muniyappa, K. (2020). UvrA and UvrC subunits of the Mycobacterium tuberculosis UvrABC excinuclease interact independently of UvrB and DNA. FEBS Letters, 594(5), 851–863. https://doi.org/10.1002/1873-3468.13671
Thakur, M., Kumar, M. B., & Muniyappa, K. (2016). Mycobacterium tuberculosis UvrB Is a robust DNA-stimulated ATPase that also possesses structure-specific ATP-dependent DNA helicase activity. Biochemistry, 55(41), 5865–5883. https://doi.org/10.1021/acs.biochem.6b00558
van der Veen, S., & Tang, C. M. (2015). The BER necessities: The repair of DNA damage in human-adapted bacterial pathogens. Nature Reviews Microbiology, 13(2), 83–94. https://doi.org/10.1038/nrmicro3391
Veening, J. W., Smits, W. K., & Kuipers, O. P. (2008). Bistability, epigenetics, and bet-hedging in bacteria. Annual Review of Microbiology, 62, 193–210. https://doi.org/10.1146/annurev.micro.62.081307.163002
Warner, D. F., Ndwandwe, D. E., Abrahams, G. L., Kana, B. D., Machowski, E. E., Venclovas, C., & Mizrahi, V. (2010). Essential roles for imuA’- and imuB-encoded accessory factors in DnaE2-dependent mutagenesis in Mycobacterium tuberculosis. Proceedings of the National Academy of Sciences of the United States of America, 107(29), 13093–13098. https://doi.org/10.1073/pnas.1002614107
Williams, A., Guthlein, C., Beresford, N., Bottger, E. C., Springer, B., & Davis, E. O. (2011). UvrD2 is essential in Mycobacterium tuberculosis, but its helicase activity is not required. Journal of Bacteriology, 193(17), 4487–4494. https://doi.org/10.1128/JB.00302-11
Wipperman, M. F., Heaton, B. E., Nautiyal, A., Adefisayo, O., Evans, H., Gupta, R., & Glickman, M. S. (2018). Mycobacterial mutagenesis and drug resistance are controlled by phosphorylation- and cardiolipin-mediated inhibition of the RecA coprotease. Molecular Cell, 72(1), 152–161. https://doi.org/10.1016/j.molcel.2018.07.037
Acknowledgements
This work was supported by the Natural Science Foundation of China (grant nos. 82072252 and 91942315) and Guangdong Provincial Key Laboratory of Regional Immunity and Diseases (grant no. 2019B030301009). We thank Dr. Jessica Tamanini at Shenzhen University Health Science Center for editing the manuscript prior to submission.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, K., Ning, Y., Kong, F. et al. Genome instability in pathogenesis of tuberculosis. GENOME INSTAB. DIS. 2, 331–338 (2021). https://doi.org/10.1007/s42764-021-00057-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42764-021-00057-8