Abstract
To understand the role of microorganisms in the functioning of forest ecosystems, the structure of bacterial communities and the enzymatic activity were determined in forest soils representing the following soil subtypes: Eutric/Dystric Brunic Arenosols (A), Eutric/Endocalcaric Cambisols (C), and Haplic/Albic Luvisols (L). Their microbiological and biochemical properties were compared based on bacterial counts and diversity, and activities of seven soil enzymes: dehydrogenases, catalase, urease, acid phosphatase, alkaline phosphatase, arylsulfatase, and β-glucosidase. Organotrophic bacteria and actinobacteria were the most abundant and featured the highest values of the EP (ecophysiological diversity index) in the Haplic/Albic Luvisol soil. In turn, the CD (colony development index) values of these bacterial groups were the highest in the Eutric/Endocalcaric Cambisols. The OTU number of bacteria allowed concluding that, at the class level, the Eutric/Dystric Brunic Arenosols and Haplic/Albic Luvisols were predominated by Alphaproteobacteria belonging to Proteobacteria, whereas the Eutric/Endocalcaric Cambisols by Actinobacteria. At the family rank, the Eutric/Dystric Brunic Arenosols were colonized in the highest numbers by Mycobacteriaceae, Rhodospirillaceae, Koribacteriaceae, and Acidobacteriaceae; the Eutric/Endocalcaric Cambisols by Nocardiaceae, Bradyrhizobiaceae, and Mycobacteriaceae, whereas Haplic/Albic Luvisols by Sinobacteriaceae and Rhodospirillaceae. Four bacterial genera, i.e., Rhodoplanes, Burkholderia belonging to Proteobacteria, Mycobacterium belonging to Actinobacteria, and Candidatus Solibacter belonging to Acidobacteria, were identified in all soils tested. The genetic diversity of bacteria was proved the highest in Eutric/Endocalcaric Cambisols. In turn, the highest enzymatic activity was found for Haplic/Albic Luvisols, while the lowest one for Eutric/Endocalcaric Cambisols. The present study results point out to significant differences between the soil types analyzed in terms of the diversity and structure of their bacterial communities and their enzymatic properties.
Similar content being viewed by others
1 Introduction
Forests are one of the most valuable and most important ecosystems on the globe. They offer the habitat to many living organisms, diverse in terms of species. The processes taking place in these ecosystems are of global importance, making it necessary to understand the composition and functions of their microbiome and the processes occurring therein (Lladó et al. 2017). Each soil is unique due to the presence of parent material and the activities of fauna and flora. It is a natural habitat for various living organisms such as bacteria, archaea, fungi, annelids, insects, small invertebrates, and plants. However, the most significant role in the functioning of soil ecosystems is ascribed to microorganisms (Baćmaga et al. 2020). The enormous wealth and diversity of soil microorganisms are the core of the trophic chain. Microorganisms are responsible for the functioning of terrestrial ecosystems, e.g., they provide nutrients to other organisms inhabiting the soil, transform sparingly available compounds into easily digestible ones, or enter into ecological interactions with organisms and biogeochemical processes (Baldrian 2017). Bacteria represent an integral part of the forest soil microbial community. They contain genes encoding for enzymes capable of plant cell wall degradation and enhance dead organic matter degradation (Berlemont and Martiny 2013). They are also responsible for the nitrogen cycle in forest ecosystems (Lladó et al. 2017) and for the weathering of minerals, leading to the release of inorganic nutrients (Uroz et al. 2011). Forest ecosystems offer suitable habitats to bacteria, including the most frequently colonized soil (Hardoim et al. 2015). According to Lauber et al. (2009), the prevailing bacteria of forest soils are belonging to Proteobacteria, Actinobacteria, Acidobacteria, Firmicutes, and Bacteroidetes. Bacterial populations are significantly affected by soil pH, organic matter content, availability of nutrients, climatic conditions, and interactions with other organisms. The variety of these parameters determines the number, structure, diversity, and activity of these microorganisms (Urbanová et al. 2015). The microbiological and biochemical properties of forest soils are also significantly influenced by tree species. Forest soils are characterized by a high plant material content, which makes them rich in organic carbon compounds and microbial biomass. The primary sources of carbon in forest soils include forest litter and plant roots (Hajnal-Jafari et al. 2016). Root secretions and plant debris offer an excellent source of carbon and nutrients to groups of microorganisms inhabiting specific ecological niches (Klimek et al. 2016). The structure of the communities is a key factor influencing the functioning of ecosystems and the sustainability of soil resources. In addition, due to the great responsiveness of microorganisms to changes in the soil microenvironment, they can serve as indicators of soil condition. The functions of most microorganisms inhabiting the soils of forest ecosystems, including bacteria, are still not fully explored (Liu et al. 2019; Wang et al. 2017). Determination of the population numbers and structure of soil microbial communities and the biochemical processes taking place will allow understanding the influence these organisms have on the functioning of forest ecosystems in varying environmental conditions (Lladó et al. 2017).
The ecological functions of soils are largely determined by the microorganisms colonizing them, which have a positive effect not only on the course of soil processes, but also on plant communities. Soil microorganisms and enzymes respond very quickly and differently to changes in the ecological processes in soil and plant cover, which makes them reliable indicators reflecting the quality of forest soils (Pająk et al. 2016).
Considering the above, a study was undertaken to determine the population numbers and diversity of bacteria as well as the activity of enzymes in Eutric/Dystric Brunic Arenosols, Eutric/Endocalcaric Cambisols, and Haplic/Albic Luvisol soils, and to identify correlations between the microbiological, enzymatic, and physicochemical properties of these soils.
2 Material and Methods
2.1 Characteristics of Study Location
The research was conducted in the Stare Jabłonki Forest District area located in the central-western part of the Warmian-Masurian Voivodeship, north-eastern part of Poland. The Stare Jabłonki Forest District is located between 53° 37′ 28″ and 53° 48′ 20″ of the northern latitude and between 20° 01′ 03″ and 20° 12′ 39″ of the eastern longitude, and its total area is 9948.96 ha. It consists of one forest area, which is divided into 8 forest units: Śmieszny Kąt, Perkunicha, Laski, Draby, Barduń, Gąsiory, Białe Błota, and Ostrowin. The Forest District lands are located in Eastern Europe, the sub-area of the East European Lowlands, the Eastern Baltic-Belarusian Lowlands province, the Eastern Baltic Lake District sub-province, the Masurian Lake District macroregion, and the Olsztyn Lakeland mesoregion. The District is located at the collision site of the Atlantic climate and the continental climate influences. According to the data of the Meteorological Station in Olsztyn, in the years 1993–2016, the average annual air temperature in the study area was + 7.8 °C, the temperature of the growing season was + 14.7 °C, and the average annual precipitation was 636 mm. The highest amount of precipitation occurs from May to July (248 mm), while in the growing season spanning from April to September, it reaches 430 mm. The length of the growing season is approx. 200 days; snow cover maintains for approx. 80 days on average (on average from December 17 to March 7). The sunniest days are in June and July, while the least sunny ones are recorded from November to January. According to the data of the Institute of Meteorology and Water Management, in 2018, the average annual air temperature in this area was + 9 °C, the annual precipitation was 550 mm, and the average air humidity was 81%. The growing season lasted about 206 days, while the snow cover maintained for 70 days. The forest cover in the territorial range of the Stare Jabłonki Forest District is 70.4%, whereas in the Warmian-Masurian Voivodeship, it accounts for 31.2%, and in Poland for 30.5%. The area of the Stare Jabłonki Forest District is characterized by a varied topography, which is a consequence of the Vistula glaciation. Forms of glacial and hydro-glacial origin dominate in its geomorphological structure. The majority of this area is outwash plain made of sand and sand with gravel.
The study was conducted at three Forest Units stands located in the forest complex of the Stare Jabłonki Forest District, i.e., Perkunicha, Laski, and Draby. The Units were characterized by varied soil conditions and proximity to each other. The Perkunicha Forest Unit (53° 74′ 68″ N, 20° 04′ 14″ E) was predominated by soils belonging to the Brunic Arenosols type (subtype Eutric/Dystric Brunic Arenosols), where the dominant tree stands were oak (Quercus L.) and hornbeam (Carpinus betulus L.). The soils of the Laski Forst Unit (53° 71′ 58″ N, 20° 10′ 93″ E) belonged to the Cambisols type (subtype Eutric/Endocalcaric Cambisols) and were covered with stands predominated by Norway spruce (Picea abies L.), small-leaved lime (Tilia cordata L.), and warty birch (Betula pendula L.). In turn, the soils found at the Draby Forest Unit (53° 69′ 256″ N, 20° 09′ 59″ E) were classified to the Luvisol type (Haplic/Albic Luvisol subtype) and were covered in majority by Scots pine (Pinus sylvestris L.), Norway spruce (Picea abies L.), hornbeam (Carpinus betulus L.), and oak (Quercus L.).
2.2 Sampling Procedure
In November 2018, samples were collected from soils classified according to the World Reference Base for Soil Resources (2014) to the subtypes Eutric/Dystric Brunic Arenosols (A), Eutric/Endocalcaric Cambisols (C), and Haplic/Albic Luvisols (L). The selected properties of these soils are provided in Table 1. The soil material was sampled from the areas belonging to the Stare Jabłonki Forest District, Poland. The soil classified to the subtype Eutric/Dystric Brunic Arenosols (A) was sampled from the area of the Perkunicha Forest Unit that classified to Eutric/Endocalcaric Cambisols (C)—from the Laki Forest Unit, and that classified as Haplic/Albic Luvisols (L)—from the Draby Forest Unit. Three plots were established on the areas of all three Forest Units examined. After removing ca. 2 cm of the upper layer of plant litter, 3 samples of soil were randomly collected from a depth of 20 cm from each plot. The samples were collected with using Egner-Riehm’s rod. Immediately after collection, the soil samples were divided into two equal portions. One portion (in the moist state) was intended for microbiological and biochemical analyses, whereas the second one for the physicochemical analyses of soil. The fresh soil material to be used for microbiological and biochemical assays was sieved through a screen with a mesh diameter of 2 mm and stored at 4 °C until analyzed, whereas the soil samples intended for physicochemical analyses were dried and sieved through the same screen.
2.3 Physicochemical Analyses of Soil
The granulometric soil composition was determined using a Mastersizer 2000 laser diffraction particle size analyzer (Malvern, Worcestershire, UK). The selected physicochemical properties of soil were established acc. to the methods described by Harris (2006). Soil pH was measured potentiometrically in 1 mol dm−3 KCl (1:2.5). Hydrolytic activity (HAC) and the sum of exchangeable base cations (EBC) were analyzed with the Kappen method, whereas organic carbon content with the Tiurin method. The total nitrogen content of soil was determined with an analyzer VarioMaxCube CN Elementar. The determined HAC and EBC values allowed computing the capacity of exchangeable cations (CEC) and soil saturation with base cations (BS).
2.4 Population Numbers, Colony Development Index, and Ecophysiological Diversity Index
Population numbers of organotrophic bacteria and actinobacteria were determined with the method of serial dilutions, in 4 replications. Organotrophic bacteria were isolated in the Bunt and Rovira (1955), whereas actinobacteria on the Küster and Williams medium described by Parkinson et al. (1971). The microorganisms were incubated on Petri dishes at a temperature of 28 °C for 10 days, and the grown colonies were counted every day. After 10-day incubation, the colony-forming units (cfu) were determined, which allowed calculating the CD (colony development index) according to the formula provided by Sarathchandra et al. (1997) and the EP (ecophysiological diversity index) according to the formula given by De Leij et al. (1993). The microbiological analyses were carried out as described by Borowik et al. (2020).
2.5 Extraction of Genomic DNA, PCR Amplification, and Illumina MiSeq Sequencing
The genomic DNA of bacteria was extracted from 1 g of soil using a Genomic Mini AX Bacteria+ kit. The resulting bacterial DNA was additionally purified with an Anti-Inhibitor Kit. DNA concentration was determined fluorometrically using a Qubit 4 Fluorometer. The presence of bacterial DNA was confirmed via Real-Time PCR in an Mx3000P thermocycler with an SYBR Green dye (A&A Biotechnology, Gdynia, Poland). The bacterial fragment of the 16S rDNA gene was amplified using universal primers: 1055F (5′-ATGGCTGTCGTCAGCT-3′) and 1392R (5′-ACGGGCGGTGTGTAC-3′). The amplicon of the gene encoding for 16S (SSU rRNA) was analyzed by the next-generation sequencing (NGS) method using a MiSeq Illumina sequencer. The hypervariable region V3–V4 was amplified using 341F and 785R primers. Chimeric and incomplete sequences were filtered off using a QIIME package based on reference databases Greengenes v13_8 (Caporaso et al. 2010). The genomic sequencing and the assembly of the metagenomic data library were made by Genomed S.A. company (Warsaw, Poland).
2.6 Activity of Soil Enzymes
Activities of 5 soil enzymes representing the class of hydrolases involved in the conversion of phosphorus compounds (acid phosphatase, alkaline phosphatase), nitrogen (urease), sulfur (arylsulfatase), and carbon (β-glucosidase, and activities of 2 enzymes belonging to the class of oxidoreductases (dehydrogenases, catalase) responsible for the course of redox reactions were determined in the study. The activity of hydrolases was determined following the procedure provided by Alef and Nannipieri (1998), that of dehydrogenases acc. to the Öhlinger method (1996), whereas that of catalase acc. to Johnson and Temple (1964). The classification of soil enzymes tested, their acronyms, units used to present analytical data, as well as substrates and products used during the above assays are presented in Table 2. The activities of enzymes were determined using a Perkin-Elmer Lambda 25 spectrophotometer, at the following wavelengths: λ = 485 nm for dehydrogenases, λ = 410 nm for alkaline phosphatase and acid phosphatase, λ = 420 nm for arylsulfatase, and λ = 400 nm for β-glucosidase. Catalase activity was analyzed employing the titration method with potassium permanganate.
2.7 Statistical Analyses
The statistical analyses were performed in Statistica 13.1 analytical package (Dell Inc. 2016) employing a one-way analysis of variance (ANOVA) at a significance level of p ≤ 0.05. Homogenous groups were calculated with the honestly significant difference (HSD) Tukey test. The counts of organotrophic bacteria and actinobacteria were presented as a dendrogram using cluster analysis (CA) and multi-dimensional exploratory techniques. Also, Pearson correlation coefficients were computed for the analyzed parameters. The relative abundance of the bacteria was computed by means of STAMP 2.1.3. software based on a two-way test of statistical hypotheses: G-test (w/Yates’) + Fisher’s (Parks et al. 2014). The metagenomic analysis data were presented in a circular graph prepared using Circos 0.68 package. The diagrams illustrate sequence similarities, proportional to each band used for grouping. Grouped data are arranged radially into segments. The outer ring represents the total percentage of the sequence while the inner ring represents the operational taxonomic unit (OTU) values. In turn, the heat map depicting bacterial families in the soils was prepared with RStudio v1.2.5033 software (RStudio Team 2019), gplots library (Warnes et al. 2020), and v3.6.2 system (R Core Team. R 2019). Data from the metagenomic analysis of bacteria were presented for the classification range of ≥ 1%. Finally, the determined OTU values of bacteria allowed computing their Shannon-Wiener (H′) and Simpson (D) indices.
3 Results
3.1 Diversity and Structure of Bacterial Communities
The Haplic/Albic Luvisols offered the most favorable conditions for bacteria proliferation. The count of organotrophic bacteria in this soil subtype was at 3.63 109 cfu kg−1 DM of soil and that of actinobacteria at 2.02 109 cfu kg−1 DM of soil. The counts of organotrophic bacteria and actinobacteria in Haplic/Albic Luvisols were higher than in Eutric/Endocalcaric Cambisols (2.9-fold and 3-fold, respectively) and in Eutric/Dystric Brunic Arenosols (1.7-fold and 1.1-fold, respectively) (Table 3).
The effect of soil subtype on differences in the counts of organotrophic bacteria and actinobacteria is depicted in a dendrogram presented in Fig. 1. Three clusters of organotrophic bacteria and actinobacteria were distinguished in three soil subtypes. The first cluster was formed by organotrophic bacteria and actinobacteria colonizing Dystric Brunic Arenosols and actinobacteria colonizing Haplic/Albic Luvisols. The second cluster included organotrophic bacteria and actinobacteria from Eutric/Endocalcaric Cambisols, whereas the last one, only organotrophs colonizing Haplic/Albic Luvisols.
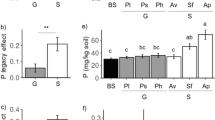
The condition of the soil environment is reliably indicated by values of the colony development index (CD) and the ecophysiological diversity index (EP). The CD values determined for the organotrophic bacteria (Fig. 2) showed their highest growth rate in Eutric/Endocalcaric Cambisols, then in Eutric/Dystric Brunic Arenosols and Haplic/Albic Luvisols (CD = 45.85, CD = 43.37, and CD = 34.78, respectively). In the case of actinobacteria, the highest CD value was recorded in Eutric/Endocalcaric Cambisols (CD = 26.95), followed by Haplic/Albic Luvisols (CD = 21.95), and Eutric/Dystric Brunic Arenosols (CD = 19.44). Opposite dependencies were noted regarding the ecophysiological diversity index (Fig. 2). The highest EP value was determined in Haplic/Albic Luvisols and reached EP = 0.74 for organotrophic bacteria and EP = 0.77 for actinobacteria. In contrast, the lowest EP values were determined for organotrophic bacteria and actinobacteria from Eutric/Dystric Brunic Arenosols (EP = 0.45 and EP = 0.61, respectively).
Values of the colony development index and ecophysiological diversity index of organotrophic bacteria and actinobacteria in the soils tested. Org, organotrophic bacteria; Act, actinobacteria; CD, colony development index; EP, ecophysiological diversity index; A, Eutric/Dystric Brunic Arenosols; C, Eutric/Endocalcaric Cambisols; L, Haplic/Albic Luvisols
The predominating bacteria in Eutric/Dystric Brunic Arenosols and Eutric/Endocalcaric Cambisols were Proteobacteria, and their OTU numbers reached 33.24% and 28.90%, respectively. In turn, the Haplic/Albic Luvisol soil subtype was most abundantly colonized by Actinobacteria, which accounted for 45.31% of the total microbiome. The analyzed soils were also colonized by Acidobacteria, which accounted for 18.84% in Eutric/Dystric Brunic Arenosols, for 21.97% in Eutric/Endocalcaric Cambisols, and for 6.11% in Haplic/Albic Luvisols (Fig. 3).
Considering the OTU number, Alphaproteobacteria belonging to Proteobacteria were the most representative bacteria at the phylum rank in Eutric/Dystric Brunic Arenosols and Haplic/Albic Luvisols. Their OTUs reached 22,945 and 18,811, respectively. In both these soil subtypes, high OTU numbers were also determined for Actinobacteria and Termoleophilia belonging to Actinobacteria, and for Acidobacteria representing the Acidobacteria phylum. In the case of Haplic/Albic Luvisols, high counts were also recorded for Gammaproteobacteria representing Proteobacteria and for DAO52 belonging to Acidobacteria. In turn, in the Eutric/Endocalcaric Cambisols, the predominating bacteria belong to Actinobacteria, and their OTU number reached 20,039. This soil subtype was also characterized by high OTUs of Termoleophilia and Alphaproteobacteria. The prevailing orders found in the class Alphaproteobacteria, belonging to Proteobacteria, included Rhizobiales (from 4.98 to 14.91%) and Rhodospirillales (from 5.78 to 9.61%). In turn, the Actinomycetales order prevailed in the class Actinobacteria with its OTUs ranging from 9.81% (Eutric/Endocalcaric Cambisols) to 23.58% (Haplic/Albic Luvisols). The predominating bacteria among the Termoleophilia were these representing the order Solirubrobacterales, which constituted from 4.60% in Eutric/Endocalcaric Cambisols to 9.29% in Haplic/Albic Luvisols (Fig. 4).
In Eutric/Dystric Brunic Arenosols, the lower taxon was predominated by Mycobacteriaceae belonging to Actinobacteria, Rhodospirillaceae belonging to Proteobacteria, as well as Koribacteriaceae and Acidobacteriaceae belonging to Acidobacteria. The most abundant bacteria found in Eutric/Endocalcaric Cambisols were from the Nocardiaceae and Mycobacteriaceae families representing the Actinobacteria phylum and from Bradyrhizobiaceae belonging to Proteobacteria. In turn, Haplic/Albic Luvisols were mostly colonized by Sinobacteriaceae and Rhodospirillaceae belonging to Proteobacteria (Fig. 5).
The Venne analysis (Fig. 6) allowed distinguishing bacterial genera unique for the three subtypes of forest soils. The highest number of unique bacterial genera was identified in Eutric/Endocalcaric Cambisols (Aeromicrobium, Pilimelia, Methylibium, Pseudonocardia, Kribbella, Cellulomonas, Streptomyces, Nocardioides, DA101), then in Haplic/Albic Luvisols (Paenibacillus, Bdellovibrio, Bacillus, FFCH10602, Sporosarcina, Clostridium, Solibacillus), whereas the lowest number in Eutric/Dystric Brunic Arenosols (Pedosphaera, Chthoniobacter). Four bacterial genera were identified as common for all three soil subtypes, i.e., Rhodoplanes and Burkholderia belonging to Proteobacteria, Mycobacterium to Actinobacteria, and Candidatus Solibacter to Acidobacteria.
The calculated values of the Shannon-Wiener index (H′) and the Simpson (D) index showed that the most diversified in terms of bacterial structure turned out to be Eutric/Endocalcaric Cambisols, followed by Haplic/Albic Luvisols, and Eutric/Dystric Brunic Arenosols (Table 4). The highest value of the Shannon-Wiener index was noted in Eutric/Endocalcaric Cambisols at the family rank (H′ = 3.20), whereas the highest value of the Simpson index in Haplic/Albic Luvisols at the order level (D = 0.94).
3.2 Activity of Soil Enzymes
The activities of all soil enzymes, except for β-glucosidases, were the highest in Haplic/Albic Luvisols, whereas β-glucosidase activity in Eutric/Dystric Brunic Arenosols. The largest differences were noted among the soil subtypes regarding urease activity, which in Eutric/Dystric Brunic Arenosols and Eutric/Endocalcaric Cambisols was almost 4-fold lower than in Haplic/Albic Luvisols. In turn, the greatest differences in the activities of dehydrogenases, catalase, alkaline phosphatase, and arylsulfatase were noticeable between Haplic/Albic Luvisols and Eutric/Endocalcaric Cambisols. Their activities determined in Eutric/Endocalcaric Cambisols were 2.7-fold, 1.8-fold, 2.1-fold, and 3-fold lower, respectively, than these assayed in Haplic/Albic Luvisols. In turn, acid phosphatase activity was found the lowest in Eutric/Dystric Brunic Arenosols (Table 5).
3.3 Correlations between Bacterial Diversity, the Structure of Bacterial Communities, Enzymatic Activity, and the Physiochemical Properties of Soil
Table 6 presents the coefficients of a simple Pearson correlation between the microbiological and enzymatic properties of soil and its physicochemical parameters. The microbiological and enzymatic properties of soil were significantly influenced by its pH, which was significantly positively correlated with the count of organotrophic bacteria, EP of organotrophic bacteria and actinobacteria, Shannon-Wiener index at the taxonomic levels from phylum to order, Simpson index at the class level, and activities of dehydrogenases, catalase, alkaline phosphatase, and acid phosphatase. In turn, negative correlations were observed between soil pH and CD of organotrophs, Shannon-Wiener index at the species level, Simpson index at the family level, and β-glucosidase activity. The microbiological and enzymatic properties of soil were also significantly affected by organic carbon content, which was significantly positively correlated with β-glucosidase activity and Shannon-Wiener index at the species level, and negatively correlated with the CD value of actinobacteria, the EP values of organotrophic bacteria and actinobacteria, the Simpson index at the species level, and acid phosphatase activity. The significant negative correlations were observed between a total nitrogen content and the count of organotrophic bacteria and actinobacteria; activities of dehydrogenases, catalase, alkaline phosphatase, arylsulfatase, and urease; Shannon-Wiener index; and Simpson index at the phylum, class, and order level. There was a significant positive correlation between the total nitrogen content and CD value of organotrophic bacteria and actinobacteria, Shannon-Wiener index at the family and species level, and Simpson index at the family, genus, and species level.
4 Discussion
4.1 Diversity and Structure of Bacterial Communities
Forest soils are considered an environment whose functioning is determined by multiple factors like, e.g., soil fraction size, organic matter content, pH value, and nutrient content. All these factors can radically modify the composition of soil microbiota (Baldrian 2017). In the present study, the counts of organotrophic bacteria and actinobacteria were the highest in Haplic/Albic Luvisols characterized by the highest pH value and, thus, by the highest sum of exchangeable base cations and soil saturation with base cations, which could consequently promote their development. Wasak et al. (2020) have confirmed that the activity of microorganisms is significantly affected by the physicochemical properties of soil, which may change under the influence of natural or anthropogenic factors. Among all microorganisms, bacteria are the major constituents of the live biomass of organisms colonizing the soil environment. They are the most abundant in the soil environment and the most diverse in terms of their metabolism (Liu et al. 2019). They play a meaningful role in ecological systems as they degrade organic matter (e.g., plant litter or dead animals), provide nutrients to other organisms, promote plant growth and development, and are involved in biochemical cycles (Baldrin 2017). The activity of bacteria in forest soils is largely determined by the forest stand, which affects both the quantity and quality of organic compounds provided in the form of leaves and root litter (Žifčáková et al. 2016). The forest litter can offer an adequate medium for the growth of bacteria and a reservoir of nutrients indispensable for their development. Plant root secretions can also positively affect their proliferation and survivability, suggesting correlations between the microorganisms and the forest flora (Wu et al. 2011). Diversified species composition of trees in forest ecosystems can influence the quantity and quality of organic matter in soil and the microclimate, which may—in turn—contribute to changes in the structure of bacterial communities. Coniferous trees are more resistant to degradation than deciduous trees due to their waxy surface layer and a high concentration of sparingly degradable phenolics (Burton et al. 2010). Hackl et al. (2004) have demonstrated that the structure of bacterial communities in Australian forests was strongly associated with the species composition of the forest stand, i.e., Alphaproteobacteria prevailed in pine forests, Holophagae and Acidobacteria in oak forests, while Verrucomicrobia and Alphaproteobacteria in spruce-fir-beech forests. In turn, forest soils analyzed by Tripathi et al. (2012) and Miyashita et al. (2013) were mainly colonized by Proteobacteria, including copiotrophic bacteria featuring various carbon metabolism, that can proliferate under various environmental conditions (Bastida et al. 2015). In the present study, Proteobacteria were found to predominate in Eutric/Dystric Brunic Arenosols and Eutric/Endocalcaric Cambisols, whereas Actinobacteria in Haplic/Albic Luvisols. He et al. (2006) have also demonstrated the predominance of Proteobacteria, Actinobacteria, and Acidobacteria in forest soils. Actinobacteria were also detected in forest soils by Žifčáková et al. (2016), Liu et al. (2019), and Zhang et al. (2019), while Acidobacteria by Deng et al. (2019a) and Li et al. (2019). These differences in the structure of bacterial communities in particular soil types can be associated with their physicochemical properties, particularly with their pH value being the major influencing factor in this respect (Preem et al. 2012). Lazzaro et al. (2006) have noticed considerable changes in the structure of bacterial communities in the soil having pH 5.8, while no changes in the soil with pH 6.7. The acidic soils are heavily colonized by bacteria belonging to Acidobacteria and Alphaproteobacteria, whereas soils with increased pH are predominated by Actinobacteria. Still, the prevalence of Acidobacteria in acidic soils depends on their organic carbon content. Bacteria belonging to Betaproteobacteria, including, i.e., the genus Burkholderia and Collimonas that are involved in the weathering of minerals, are commonly found in forest soils. The Burkholderia genus bacteria were detected in all soil subtypes analyzed in the present study. As reported by Žifčáková et al. (2018), forest soils were predominated by Pseudomonas, Beijerinckia, and Acidiphilia bacteria. Furthermore, they were strongly colonized by Candidatus Koribacter and Rhodoplanes. The forest soils are characterized by vertical stratification due to organic litter degradation and mineral horizon weathering. A reduction in the organic matter content of soil can contribute to a smaller biomass of microorganism, poorer soil respiration, and lower activity of extracellular enzymes (Lladó et al. 2017).
Comparing the diversity of forest soil bacteria in terms of their physicochemical properties has allowed identifying some unique bacterial communities (Carnovale et al. 2019). In the present study, the highest number of unique bacterial genera was found in Eutric/Endocalcaric Cambisols, whereas the lowest one in Haplic/Albic Luvisols. As demonstrated by Zhang et al. (2019), the OTU number of unique bacteria was higher in the forest than in the arable soils. They showed more significant differences in the structure of bacterial communities in the soils having a diversified vegetation cover than in the soils covered with one plant species.
The bacterial diversity was determined based on the values of the Shannon-Wiener and Simpson indices, which are reliable indicators of the diversity of soil systems and thereby enable comparing the structure of bacterial communities and their sensitivity to various environmental factors. The higher the values of these indices, the greater is the diversity of microorganism communities (Zhao et al. 2020). In the present study, the greatest diversity of bacteria was observed in Eutric/Endocalcaric Cambisols and the lowest one in Eutric/Dystric Brunic Arenosols. As reported by Lauber et al. (2009), the diversity of forest soil bacteria can change upon the influence of environmental factors and species composition of the forest tree stand. Presumably, the structure of bacterial communities and their diversity can be determined by the interactions of physical and chemical factors in the soil environment (Wei et al. 2018; Deng et al. 2019b). What is more, Zhang et al. (2019) claimed that the bacterial diversity of forest soils could be directly associated with the plant cover and soil type.
4.2 Enzymatic Activity
The enzymatic activity plays a key role in the course of biochemical processes in soil. Therefore, it can serve as a reliable quality indicator of soil ecosystems (Baćmaga et al. 2019; Notaro et al. 2018). The soil enzymes are natural mediators and catalysts of, e.g., transformation of organic matter, production and degradation of humus, conversion of organic compounds into mineral compounds available to plants, or participation in the organic matter cycle and energy flow (Błońska et al. 2017). Determining the enzymatic activity in the soils of the forest ecosystems is necessary to assess soil fertility and quality, soil metabolic potential, and correlations between its biochemical, microbiological, and physicochemical properties. Januszek et al. (2015) have claimed that the activity of soil enzymes, dehydrogenases in particular, depends on the soil granulometric composition, organic matter content, and the physicochemical properties of soil, including its pH, hydrolytic activity, sum of exchangeable base cations, sorptive capacity, and saturation with base cations. In the present study, the highest activities of soil enzymes, except for β-glucosidase, were determined in Haplic/Albic Luvisol soil subtype, which was characterized by the highest pH value and the lowest value of hydrolytic activity. The statistical analyses demonstrated positive correlations between soil pH and activities of dehydrogenases, catalase, alkaline phosphatase, and acid phosphatase, and also a negative correlation between soil pH and β-glucosidase. The above results enable concluding that soil pH has the greatest impact on the activities of soil enzymes. This factor could significantly affect the counts and diversity of organotrophic bacteria and actinobacteria, thereby contributing to the increased enzyme secretion by these microorganisms (Błońska et al. 2017). According to Rous et al. (2010), strongly acidified soils are characterized by a low activity of soil enzymes, including dehydrogenase, probably due to the predominance of fungi over bacteria. Also, Januszek et al. (2015) have reported a low dehydrogenase activity in poor and strongly acidified forest soils. In the present study, the highest activity of dehydrogenases was determined in Haplic/Albic Luvisols, whose pH was the highest among all soils analyzed. Bueis et al. (2018) have determined higher activities of dehydrogenases, catalase, and urease in lime than acid soils, whereas Turner (2010) have claimed that soil pH can significantly affect enzyme content in the soil environment through the modification of the ionic active form of enzymes, three-dimensional structure of enzymes, and substrate affinity to the enzyme. In the present study, the highest activity of β-glucosidase was demonstrated in Eutric/Dystric Brunic Arenosols, probably due to the abundance of organic matter, which turned out to be a rich source of nutrients to the microorganisms capable of producing enzymes involved in carbon compounds degradation, like, e.g., β-glucosidase (Błońska et al. 2017). In turn, Madejón et al. (2012) have reported organic matter degradation to be the main process in biogeochemical cycles. Poor degradation of organic matter, resulting from climate conditions or soil acidification, can limit the availability of organic matter to the microorganisms colonizing soil, leading to changes in its enzymatic activity. Veres et al. (2015) indicate that the forest soils have the highest contribution of the light fraction composed in part of degraded plant and animal debris and a high number of cells of microorganisms, which may contribute to their higher enzymatic activity. In addition, changes in the enzymatic activity of these soils can be determined by the species composition of tree stands. For instance, Błońska et al. (2017) have reported a higher β-glucosidase activity in the soils of fir stands than in hornbeam or maple stands. They have demonstrated that firs stimulate β-glucosidase activity by root secretions of nutrients available to microorganisms, thus increasing their number and activity. According to these authors, β-glucosidase activity could be rapidly decreased by the small mass of plant roots and depleting forest litter layer (Veres et al. 2015), whereas activities of dehydrogenases, urease, and protease did not differ significantly among the soils of fir, hornbeam, and maple stands. In the present study, the activity of β-glucosidase was the highest in Eutric/Dystric Brunic Arenosols overgrown with a tree stand predominated by hornbeam and oak. This tree stand composition could contribute to greater accumulation of organic matter, as evidenced by the highest organic carbon content in this soil type. In turn, the activities of the other studied enzymes were the highest in Haplic/Albic Luvisols overgrown with Scots pine, hornbeam, Norway spruce, and oak. Błońska et al. (2017) have determined the highest activities of dehydrogenases and urease in soils covered with deciduous trees. Finally, Wang et al. (2017) have reported that tree species determine soil properties due to the varying content of organic matter pervading to the soil environment.
5 Conclusions
The study results allow concluding that the microbiological and enzymatic properties of forest soils were strongly correlated with their physicochemical properties. The most beneficial conditions for bacteria proliferation occurred in Haplic/Albic Luvisols, as evidenced by the highest counts of organotrophic bacteria and actinobacteria and by the highest value of the ecophysiological diversity index (EP) determined for this soil subtype. In turn, the highest value of the colony development index (CD) of organotrophic bacteria and actinobacteria was determined for Eutric/Endocalcaric Cambisols. The Eutric/Dystric Brunic Arenosols and Eutric/Endocalcaric Cambisols soils were predominated by Proteobacteria, whereas Haplic/Albic Luvisols by Actinobacteria. The highest genetic diversity of bacteria was found in the Eutric/Endocalcaric Cambisol soil subtype. These differences can be due to the analytical methods used. The Haplic/Albic Luvisols showed the highest activities of dehydrogenases, catalase, acid phosphatase, alkaline phosphatase, arylsulfatase, and urease, whereas Eutric/Dystric Brunic Arenosols, β-glucosidase. The evaluation of soils of forest ecosystems based on microbiological and biochemical analyses allows for a better understanding of their functioning and helps planning further activities associated with the monitoring and improvement of this environment quality.
References
Alef K, Nannipieri P (1998) Methods in applied soil microbiology and biochemistry. In: Alef K, Nannipieri P (eds), Academic London, UK, pp. 316–365
Baćmaga M, Wyszkowska J, Kucharski W (2019) Effect of the biostimulation of soil exposed to tebuconazole on its biological activity. J Soils Sediment 19:3728–3741. https://doi.org/10.1007/s11368-019-02325-3
Baćmaga M, Wyszkowska J, Kucharski W (2020) Response of soil microorganisms and enzymes to the foliar application of Helicur 250 EW fungicide on Horderum vulgare L. Chemosphere 242:125163. https://doi.org/10.1016/j.chemosphere.2019.125163
Baldrian P (2017) Forest microbiome: diversity, complexity and dynamics. FEMS Microbiol Rev fuw040 41:109–130. https://doi.org/10.1093/femsre/fuw040
Bastida F, Garcia C, von Bergen M, Moreno JL, Richnow HH, Jehmlich N (2015) Deforestation fosters bacterial diversity and the cyanobacterial community responsible for carbon fixation processes under semiarid climate: a metaproteomics study. Appl Soil Ecol 93:65–67. https://doi.org/10.1016/j.apsoil.2015.04.006
Berlemont R, Martiny AC (2013) Phylogenetic distribution of potential cellulases in bacteria. Appl Environ Microbiol 79:1545–1554. https://doi.org/10.1128/AEM.03305-12
Błońska E, Lasota J, Gruba P (2017) Enzymatic activity and stabilization of organic matter in soil with different detritus inputs. Soil Sci Plant Nut 63(3):242–247. https://doi.org/10.1080/00380768.2017.1326281
Borowik A, Wyszkowska J, Kucharski J (2020) Impact of various grass species on soil bacteriobiome. Diversity 12:212. https://doi.org/10.3390/d12060212
Bueis T, Turrión MB, Bravo F, Pando V, Muscolo A (2018) Factors determining enzyme activities in soils under Pinus halepensis and Pinus sylvestris plantations in Spain: a basis for establishing sustainable forest management strategies. Ann For Sci 75:34. https://doi.org/10.1007/s13595-018-0720-z
Bunt JS, Rovira AD (1955) Microbiological studies of some subantarctic soils. J Soil Sci 6:119–128. https://doi.org/10.1111/j.1365-2389.1955.tb00836.x
Burton J, Chen C, Xu Z, Ghadiri H (2010) Soil microbial biomass, activity and community composition in adjacent native and plantation forests of subtropical Australia. J Soils Sediment 10:1267–1277. https://doi.org/10.1007/s11368-010-0238-y
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high through put community sequencing data. Nat Methods 7:335–336
Carnovale D, Bissett A, ThrallPH BG (2019) Plant genus (Acacia and Eucalyptus) alters soil microbial community structure and relative abundance within revegetated shelterbelts. Appl Soil Ecol 133:1–11. https://doi.org/10.1016/j.apsoil.2018.09.001
De Leij FAAM, Whipps JM, Lynch JM (1993) The use of colony development for the characterization of bacterial communities in soil and on roots. Microb Ecol 27:81–97. https://doi.org/10.1007/BF00170116
Dell Inc (2016) Dell statistica (data analysis software system), version 13.1., Dell Inc Tulsa, OK, USA
Deng J, Zhou Y, Bai X, Luo J, Yin Y, Zhu W (2019a) Soil microbial functional diversity responses to different revegetation types in Baishilazi Nature Reserve. Pol J Environ Stud 28(5):3675–3686. https://doi.org/10.15244/pjoes/99100
Deng J, Zhang Y, Yin Y, Zhu X, Zhu WX, Zhou YB (2019b) Comparison of soil bacterial community and functional characteristics following afforestation in the semi-arid areas. PeerJ 7:e7141. https://doi.org/10.7717/peerj.7141
Hackl E, Zechmeister-Boltenstern S, Bodrossy L, Sessitsch A (2004) Comparison of diversities and compositions of bacterial populations inhabiting natural forest soils. Appl Environ Microbiol 70:5057–5065. https://doi.org/10.1128/AEM.70.9.5057-5065.2004
Hajnal-Jafari T, Durić S, Stamenov D, Vasić V, Hackenberger D (2016) Microbial activity in forest soil under beech, spruce, Douglas fir and fir. Contemp Agric 65(1–2):33–38. https://doi.org/10.1515/contagri-2016-0006
Hardoim PR, van Overbeek LS, Berg G, Pirttilä AM, Company S, Campisano A, Döring M, Sessitsch A (2015) The hidden world with in plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79(3):293–320. https://doi.org/10.1128/MMBR.00050-14
Harris DC (2006) Quantitative chemical analysis. Michelson Laboratory Chine Lake USA. WH Freeman and Company 7th edition, 1008
Januszek K, Błońska E, Długa J, Socha J (2015) Dehydrogenase activity of forest soils depends on the assay used. Int Agrophys 29:47–59. https://doi.org/10.1515/intag-2015-0009
Johnson JJ, Temple K (1964) Some variables affecting the measurement of catalase activity in soil. Soil Sci Soc Proc 28:207–209. https://doi.org/10.2136/sssaj1964.03615995002800020024x
Klimek B, Chodak M, Jaźwa M, Solak A, Tarasek A, Niklińska M (2016) The relationship between soil bacteria substrate utilisation patterns and the vegetation structure in temperate forests. Eur J Forest Res 135:179–189. https://doi.org/10.1007/s10342-015-0929-4
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120. https://doi.org/10.1128/AEM.00335-09
Lazzaro A, Hartmann M, Blaser P, Widmer F, Schulin R, Frey B (2006) Bacterial community structure and activity in different Cd-treated forest soils. FEMS Microbiol Ecol 58:278–292. https://doi.org/10.1111/j.1574-6941.2006.00163.x
Li P, Shi RJ, Zhao F, Yu JH, Cui XY, Hu JG, Zhang Y (2019) Soil bacterial community structure and predicted functions in the larch forest during succession at the Greater Khingan Mountains of Northeast China. Chin J Appl Ecol 30:95–107. https://doi.org/10.13287/j.1001-9332.201901.010
Liu Y, Wang S, Wang Z, Zhang Z, Qin H, Wei Z, Feng K, Li S, Wu Y, Yin H, Li H, Deng Y (2019) Soil microbiome mediated nutrients decline during forest degradation process. Soil Ecol Lett 1(1–2):59–71. https://doi.org/10.1007/s42832-019-0009-7
Lladó S, López-Mondéjar R, Baldrian P (2017) Forest soil bacteria: diversity, involvement in ecosystem processes, and response to global change. Microbiol Mol Biol Rev 81:e00063–16. https://doi.org/10.1128/MMBR.00063-16
Madejón P, Soler-Rovira P, Ciadamidaro L, Cabrera F, Madejón E (2012) Trace element-rich litter in soils: influence on biochemical properties related to the carbon cycle. J Soils Sediment 12:663–673. https://doi.org/10.1007/s11368-012-0493-1
Miyashita NT, Iwanaga H, Charles S, Diway B, Sabang J, Chong L (2013) Soil bacterial community structure in five tropical forests in Malaysia and one temperate forest in Japan revealed by pyrosequencing analyses of 16S rRNA gene sequence variation. Genes Genet Syst 88:93–103. https://doi.org/10.1266/ggs.88.93
Notaro KA, de Medeiros EV, Duda GP, Moreira KA, de Barros JA, dos Santos UJ, Lima JRS, Moraes WS (2018) Enzymatic activity, microbial biomass, and organic carbon of Entisols from Brazilian tropical dry forest and annual and perennial crops. Chil J Agric Res 78(1):68–77. https://doi.org/10.4067/S0718-58392018000100068
Öhlinger R (1996) Dehydrogenase activity with the substrate TTC. In: Schinner F, Ohlinger R, Kandler E, Margesin R (eds) Methods in Soil Biology. Springer, Berlin, Germany, pp 241–243
Pająk M, Błońska E, Frąc M, Oszust K (2016) Functional diversity and microbial activity of forest soils that are heavily contaminated by lead and zinc. Water Air Soil Pollut 227:348. https://doi.org/10.1007/s11270-016-3051-4
Parkinson D, Gray FRG, Williams ST (1971) Methods for studying the ecology of soil micro-organism. Blackwell Scientific Publication, Oxford, IBP Handbook 19
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3312. https://doi.org/10.1093/bioinformatics/btu494
Preem JK, Truu J, Truu M, Mander Ü, Oopkaup K, Lõhmus,K Lõhmus K, Helmisaari HS, Uri V, Zobel M (2012) Bacterial community structure and its relationship to soil physico-chemical characteristics in alder stands with different management histories. Ecol Eng 49:10–17. https://doi.org/10.1016/j.ecoleng.2012.08.034
R Core Team R: A language and environment for statistical computing; R foundation for statistical computing: Vienna, Austria (2019) Available online: https://www.R-project.org/ (accessed on 23 February 2020)
Rous J, Brookes PC, Bååth E (2010) Investigating the mechanisms for the opposing pH relationships of fungal and bacterial growth in soil. Soil Biol Biochem 42:926–934. https://doi.org/10.1016/j.soilbio.2010.02.009
RStudio Team. RStudio: Integrated Development for R, RStudio, Inc.: Boston. MA, USA (2019) Available online: http://www.rstudio.com/ (accessed on 23 February 2020)
Sarathchandra SU, Burch G, Cox NR (1997) Growth patterns of bacterial communities in the rhizoplane and rhizosphere of with clover (Trifolium repens L) and perennial ryegrass (Lolium perenne L) in long-term pasture. Appl Soil Ecol 6:293–299. https://doi.org/10.1016/S0929-1393(97)00015-2
Tripathi BM, Kim M, Singh D, Lee-Cruz L, Lai-Hoe A, Ainuddin AN, Go R, Rahim RA, Husni MHA, Chun J, Adams JM (2012) Tropical soil bacterial communities in Malaysia: pH dominates in the equatorial tropics too. Microb Ecol 64:474–484. https://doi.org/10.1007/s00248-012-0028-8
Turner BL (2010) Variation in pH optima of hydrolytic enzyme activities in tropical rain forest soils. Appl Environ Microb 76:6485–6493. https://doi.org/10.1128/AEM.00560-10
Uroz S, Oger P, Lepleux C, Collignon C, Frey-Klett P, Turpault MP (2011) Bacterial weathering and its contribution to nutrient cycling in temperate forest ecosystems. Res Microbiol 162:820–831. https://doi.org/10.1016/j.resmic.2011.01.013
Veres Z, Kotroczó Z, Fekete I, Tóth JA, Lajtha K, Townsend K, Tóthmérész B (2015) Soil extracellular enzyme activities are sensitive indicators of detrital inputs and carbon availability. Appl Soil Ecol 92:18–23. https://doi.org/10.1016/j.apsoil.2015.03.006
Wang J, Ji Y, Yuan Z, Guan J, Liu C, Lv L (2017) Analysis of bacterial community structure and diversity in different restoration methods in Qixing River wetland. Adv J Toxicol Curr Res 1(2):049–055
Warnes GR, Bolker B, Bonebakker L, Gentleman R, Huber W, Liaw A, Lumley T, Maechler M, Magnusson M, Moeller S, Schwartz M, Venables B, Galili T (2020) Gplots: various R programming tools for plotting data. R Package Version 2.17.0. Available online: https://CRAN.R-project.org/package=gplots (accessed on 23 February 2020)
Wasak K, Klimek B, Drewnik M (2020) Rapid effects of windfall on soil microbial activity and substrate utilization patterns in the forest belt in the Tatra Mountains. J Soils Sediment 20:801–815. https://doi.org/10.1007/s11368-019-02439-8
Wei H, Peng C, Yang B, Song H, Li Q, Jiang L, Wei G, Wang K, Wang H, Liu S, Liu X, Chen D, Li Y, Wang M (2018) Contrasting soil bacterial community, diversity, and function in two forests in China. Front Microbiol 9:1693. https://doi.org/10.3389/fmicb.2018.01693
World Reference Base for Soil Resources (2014) International soil classification system for naming soils and creating legends for soil maps. World Soil Resources Reports, 106, FAO: Rome, Italy
Wu L, Feinstein LM, Valverde-Barrantes O, Kershner MW, Leff LG, Blackwood CB (2011) Placing the effects of leaf litter diversity on saprotrophic microorganisms in the context of leaf type and habitat. Microb Ecol 61:399–409. https://doi.org/10.1007/s00248-010-9760-0
Zhang J, Xin Y, Zhao Y (2019) Diversity and functional potential of soil bacterial communities in different types of farmland shelterbelts in mid-western Heilongjiang, China. Forests 10:1115. https://doi.org/10.3390/f10121115
Zhao Q, Liu W, Li Y, Ke M, Qu Q, Yuan W, Pan X, Qian H (2020) Enantioselective effects of imazethapyr residues on Arabidopsis thaliana metabolic profile and phyllosphere microbial communities. J Environ Sci 93:57–65. https://doi.org/10.1016/j.jes.2020.03.009
Žifčáková L, Větrovský T, Howe A, Baldrian P (2016) Microbial activity in forest soil reflects the changes in ecosystem properties between summer and winter. Environ Microbiol 18(1):288–301. https://doi.org/10.1111/1462-2920.13026
Urbanová M, Snajdr J, Baldrian P (2015) Composition of fungal and bacterial communities in forest litter and soil is largely determined by dominant trees. Soil Biol Biochem 84:53–64. https://doi.org/10.1016/j.soilbio.2015.02.011
Funding
The results presented in this paper were obtained as part of a comprehensive study financed by the University of Warmia and Mazury in Olsztyn, Faculty of Agriculture and Forestry, Department of Soil Science and Microbiology (grant No. 30.610.006-110). Project financially supported by Minister of Science and Higher Education in the range of the program entitled “Regional Initiative of Excellence” for the years 2019–2022, Project No. 010/RID/2018/19, amount of funding 12.000.000 PLN.
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Human and Animal Rights
This research did not involve human participants and animals.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baćmaga, M., Wyszkowska, J., Borowik, A. et al. Microbiological and Biochemical Properties in Eutric/Dystric Brunic Arenosols, Eutric/Endocalcaric Cambisols, and Haplic/Albic Luvisols Soils. J Soil Sci Plant Nutr 21, 1277–1292 (2021). https://doi.org/10.1007/s42729-021-00439-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-021-00439-7