Abstract
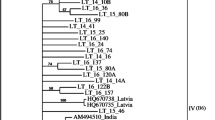
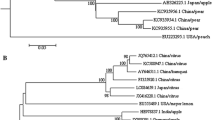
Apple chlorotic leaf spot virus (ACLSV) is one of the most common viruses infecting apple and pear trees, and Iran is among the top ten apple-producing countries in the world. In the present study, incidence and genetic diversity of ACLSV were investigated in the main apple-growing regions of Iran. To achieve this purpose, a total of 1053 leaf samples were collected from orchards located in nine Iranian provinces. ACLSV infection was detected by DAS-ELISA in 48 samples (4.55%) from seven provinces and was confirmed by RT-PCR. Based on the geographical origin, 19 representative isolates were selected for phylogenetic analysis. Nineteen amplicons, with a size of 677 base pairs (bp), containing the 3′ end of the movement protein (MP), the coat protein (CP) gene, and 3′-UTR sequence, were sequenced. Sequence analysis, using data of 45 isolates from 12 different countries, including the 19 Iranian isolates, showed that CP gene among the Iranian isolates were 80–99% identical at both nucleotide and amino acid levels, and these isolates were placed in the B6 (AB326224) and P-205 (D14996) ACLSV groups. This is the first genetic analysis of ACLSV in Iran.


Similar content being viewed by others
References
Adams MJ, Antoniw JF, Bar-Joseph M, Brunt AA, Candresse T, Foster GD, Martelli GP, Milne RG, Zavriev SK, Fauquet CM (2004) The new plant virus family Flexiviridae and assessment of molecular criteria for species demarcation. Arch Virol 149:1045–1060
Al Rwahnih M, Turturo C, Minafra A, Saldarelli P, Myrta A, Pallas V, Savino V (2004) Molecular variability of Apple chlorotic leaf spot virus in different hosts and geographical regions. J Plant Pathol 86:117–122
Bao Y, Chetvernin V, Tatusova T (2012) Pairwise sequence comparison (PASC) and its application in the classification of filoviruses. Viruses 4:1318–1327
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Report 11:113–116
Chen S, Zhou Y, Ye T, Hao L, Guo L, Fan Z, Li S, Zhou T (2014) Genetic variation analysis of Apple chlorotic leaf spot virus coat protein reveals a new phylogenetic type and two recombinants in China. Arch Virol 159:1431–1438
Desvignes J, Boyé R (1989) Different diseases caused by the chlorotic leaf spot virus on the fruit trees. Acta Hortic 235:31–38
Desvignes JC, Boyé R, Cornaggia D, Grasseau N, Hurtt S, Waterworth H (1999) Virus diseases of fruit trees. Centre Technique Interprofessionnel des Fruits et Légumes (CTIFL), Paris, pp 239–252
German S, Candresse T, Lanneau M, Huet J, Pernollet J, Dunez J (1990) Nucleotide sequence and genomic organization of Apple chlorotic leaf spot closterovirus. Virology 179:104–112
German S, Delbos RP, Candresse T, Lannean L, Dunez J (1997) Complete nucleotide sequence of the genome of a severe cherry isolate of Apple chlorotic leaf spot trichovirus (ACLSV). Arch Virol 142:833–841
Guo W, Zheng W, Wang M, Xiaohong L, Ma Y, Dai H (2016) Genome sequences of three Apple chlorotic leaf spot virus isolates from Hawthorns in China. PLoS One 11:e0161099. https://doi.org/10.1371/journal.pone.0161099
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Oxford Journals 41:95–98
Jelkmann W, Kunze L (1995) Plum pseudopox in German plum after infection with an isolate of Apple chlorotic leaf spot virus causing plum line pattern. Acta Hortic 386:122–125
Keshavarz T, Shams-Bakhsh M (2014) Incidence and distribution of Apple chlorotic leaf spot virus in the main fruit growing areas of Iran. Arch Phytopathol Plant Protect 48:306–312
Keshavarz T, Shams-Bakhsh M, Norinejad SH (2009) First report of Apple chlorotic leaf spot virus infection of apple trees in Iran. J Plant Pathol 91:233
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Liu P, Li Z, Song S, Wu Y (2014) Molecular variability of Apple chlorotic leaf spot virus in Shaanxi, China. Phytoparasitica 42:445–454
Mahfoudhi N, Elair M, Moujahed R, Salleh W, Djelouah K (2013) Occurrence and distribution of pome fruit viruses in Tunisia. Phytopathol Mediterr 52:136–140
Marini D, Gibson P, Scott S (2008) The complete nucleotide sequence of an isolate of Apple chlorotic leaf spot virus from peach (Prunus persica (L.) Batch). Arch Virol 153:1003–1005
Mathioudakis MM, Maliogka VI, Katsiani AT, Katis NI (2010) Incidence and molecular variability of apple stem pitting and apple chlorotic leaf spot viruses in apple and pear orchards in Greece. J Plant Pathol 92:139–147
Menzel W, Jelkmann W, Maiss E (2002) Detection of four apple viruses by multiplex RT-PCR assays with co-amplification of plant mRNA as internal control. J Virol Methods 99:81–92
Németh M (1986) Virus, mycoplasma and rickettsia diseases of fruit trees. Akademiai Kiado, Budapest
Niu F, Pan S, Wu Z, Jiang D, Li S (2012) Complete nucleotide sequences of the genomes of two isolates of Apple chlorotic leaf spot virus from peach (Prunus persica) in China. Arch Virol 157:783–786
Paduch-Cichal E, Szyndel MS, Tomala K (2005) Preliminary results of the study on viruses occurring in ‘Mutsu’ apple cultivar trees. Phytopathol Pol 37:87–90
Rana T, Chandel V, Kumar Y, Ram R, Hallan V, Zaidi AA (2010) Molecular variability analyses of Apple chlorotic leaf spot virus capsid protein. J Biosci 35:605–615
Salem N, Mansour A, Al-Musa A (2005) Viruses of pome fruit trees in Jordan. J Plant Pathol 87:123–126
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Laboratory Press, New York
Sato K, Yoshikawa N, Takahashi T (1993) Complete nucleotide sequence of the genome of an apple isolate of Apple chlorotic leaf spot virus. J Gen Virol 74:1927–1931
Song YS, Hong N, Wang LP, Xu WX, Hu HJ, Tian R, Wang GP (2011) Molecular and serological diversity in Apple chlorotic leaf spot virus from sand pear (Pyrus pyrilofia) in China. Eur J Plant Pathol 130:183–196
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Yaegashi H, Isogai M, Tajima H, Sano T, Yoshikawa N (2007) Combinations of two amino acids (Ala40 and Phe75 or Ser40 and Tyr75) in the coat protein of Apple chlorotic leaf spot virus are crucial for infectivity. J Gen Virol 88:2611–2618
Acknowledgements
We would like to thank the Iranian Ministry of Science, Research and Technology, for supporting F. Abtahi’s internship in Italy.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 38 kb)
Rights and permissions
About this article
Cite this article
Abtahi, F., Shams-Bakhsh, M., Safaie, N. et al. Incidence and genetic diversity of apple chlorotic leaf spot virus in Iran. J Plant Pathol 101, 513–519 (2019). https://doi.org/10.1007/s42161-018-00224-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-018-00224-z