Abstract
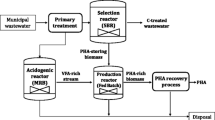
The objective of this study was to assess the possibility of retrofitting an existing full-scale wastewater treatment plant (WWTP) based on a sequencing batch reactor (SBR) technology with the enhanced biological phosphorus removal (EBPR) process. Wastewater characterisation showed highly variable influent composition that fluctuated throughout the year with a rather low and unstable SBOD/TP ratio (SBOD—soluble biological oxygen demand; TP—total phosphorus), which is considered unfavourable for EBPR. Characterisation of the sludge showed that the non-EBPR SBR sludge from the WWTP Koprivnica contained no detectable phosphorus accumulating organisms (PAO), but could be transformed in a laboratory into EBPR performing sludge in less than 45 days under favourable conditions for PAOs. The microbial community composition was assessed using an FISH (fluorescence in situ hybridization) analysis, which confirmed that the original sludge from the WWTP, which did not have detectable PAOs, was transformed into the sludge enriched by PAOs belonging to the genus ‘Candidatus Accumulibacter phosphatis’ after 43 days of cultivation. A plant retrofit, based on the results of laboratory experiments, was proposed with the enrichment of the wastewater with volatile fatty acids via primary anaerobic fermentation and step feeding. Results of mathematical modelling (BioWin) showed that such strategy could lead to sufficient P removal through EBPR in this WWTP.
Article Highlights
-
Wastewater composition was unfavourable for sufficient P removal by the EBPR process.
-
The current sludge had no detectable PAO.
-
Cultivation of the sludge in laboratory SBR enriched current sludge with PAOs to perform substantial EBPR.
-
Simulations showed that fermentation of the influent in full-scale WWTP could be used to feed the PAO to achieve sufficient bio-P removal.






Similar content being viewed by others
References
Amann R, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analysing mixed microbial populations. Appl Environ Microbiol 56:1919–1925
APHA, AWWA, WEF (1998) Standard methods for the examination of water and wastewater, 22nd edn. American Water Works Association, Denver
Barajas MG, Escalas A, Mujeriego R (2002) Fermentation of a low VFA wastewater in an activated primary tank. Water SA 28(1):89–98. https://doi.org/10.4314/wsa.v28i1.4872
Cao G, Wang S, Peng Y, Miao Z (2013) Biological nutrient removal by applying modified four step-feed technology to treat weak wastewater. Bioresour Technol 128:604–611. https://doi.org/10.1016/j.biortech.2012.09.078
Coats ER, Brinkman CK, Lee S (2017) Characterizing and contrasting the microbial ecology of laboratory and full-scale EBPR systems cultured on synthetic and real wastewaters. Water Res 108:124–136. https://doi.org/10.1016/j.watres.2016.10.069
Crocetti GR, Hugenholtz P, Bond PL, Schuler A, Keller J, Jenkins D, Blackall LL (2000) Identification of polyphosphate- accumulating organisms and design of 16S rRNA directed probes for their detection and quantitation. Appl Environ Microbiol 66:1175–1182. https://doi.org/10.1128/aem.66.3.1175-1182.2000
Crocetti GR, Banfield JF, Keller J, Bond PL, Blackall LL (2002) Glycogen-accumulating organisms in laboratory-scale and full-scale wastewater treatment processes. Microbiology 148:3353–3364. https://doi.org/10.1099/00221287-148-11-3353
Daims H, Bruhl A, Amann R, Schleifer KH, Wagner M (1999) The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22:434–444. https://doi.org/10.1016/S0723-2020(99)80053-8
Fang W, Zhang X, Zhang P, Wan J, Guo H, Ghasimi DSM, Morera XC, Zhang T (2020) Overview of key operation factors and strategies for improving fermentative volatile fatty acid production and product regulation from sewage sludge. J Environ Sci 87:94–110. https://doi.org/10.1016/j.jes.2019.05.027
Gu AZ, Saunders A, Neethling JB, Stensel HD, Blackall LL (2008) Functionally relevant microorganisms to enhanced biological phosphorus removal performance at full-scale wastewater treatment plants in the United States. Water Environ Res 80:688–698. https://doi.org/10.2175/106143008x276741
Henze M, van Loosdrecht MCM, Ekama GA, Brdjanovic D (2008) Biological wastewater treatment: principles, modelling and design. IWA Publishing, London
Liu Y, Shi H, Li W, Hou Y, He M (2011) Inhibition of chemical dose in biological phosphorus and nitrogen removal in simultaneous chemical precipitation for phosphorus removal. Bioresour Technol 102:4008–4012. https://doi.org/10.1016/j.biortech.2010.11.107
Lopez-Vazquez CM, Brdjanovic D, Hooijmans CM, Gijzen HJ, van Loosdrecht MCM (2008) Factors affecting the occurrence of phosphorus accumulating organisms (PAO) and glycogen accumulating organisms (GAO) at full-scale enhanced biological phosphorus removal (EBPR) wastewater treatment plants. Water Res 42:2349–2360. https://doi.org/10.1016/j.watres.2008.01.001
Meijer SCF, Brdjanovic D (2012) A practical guide to activated sludge modeling, UNESCO-IHE lecture notes, published by UNESCO-IHE. Institute for Water Education, ISBN:9789073445260
Metcalf and Eddy (2003) Wastewater engineering: treatment and reuse, 4th edn. McGraw-Hill Higher Education, New York
Nielsen PH, McIlroy SJ, Albertsen M, Nierychlo M (2019) Re-evaluating the microbiology of the enhanced biological phosphorus removal process. Curr Opin Biotechnol 57:111–118. https://doi.org/10.1016/j.copbio.2019.03.008
Qiu G, Zuniga-Montanez R, Law Y, Thi SS, Nguyen TQN, Eganathan K, Liu X, Nielsen PH, Williams RBH, Wuertz S (2019) Polyphosphate-accumulating organisms in full-scale tropical wastewater treatment plants use diverse carbon sources. Water Res 149:496–510. https://doi.org/10.1016/j.watres.2018.11.011
Randall CW, Barnard JL, Stensel HD (1992) Design and retrofit of wastewater treatment plants for biological nutrient removal. In: Eckenfelder WW, Malina JF, Patterson JW (eds) Water quality management library, vol 5. Technomic Publishing Company Co. Inc., Lancaster
Shen N, Zhou Y (2016) Enhanced biological phosphorus removal with different carbon sources. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-016-7518-4
Sikic T, Curko J, Crnek V, Horvat S, Matosic M (2016) Feasibility of wastewater treatment plant upgrade by enhanced biological phosphorus removal: case study Koprivnica, Croatia. Fresen Environ Bull 25:1811–1819
Smolders GJF, van Der Meij J, van Loosdrecht MCM, van Heijnen JJ (1994) Model of the anaerobic metabolism of the biological phosphorus removal process: stoichiometry and pH influence. Biotechnol Bioeng 43:461–470. https://doi.org/10.1002/bit.260430605
Tang J, Wang XC, Hu Y, Pu Y, Huang J, Hao Ngo H, Zeng Y, Li Y (2018) Nitrogen removal enhancement using lactic acid fermentation products from food waste as external carbon sources: performance and microbial communities. Bioresour Technol. https://doi.org/10.1016/j.biortech.2018.02.033
Tong JA, Chen YG (2007) Enhanced biological phosphorus removal driven by short-chain fatty acids produced from waste activated sludge alkaline fermentation. Environ Sci Technol 41(20):7126–7130. https://doi.org/10.1021/es071002n
Tu YJ, Schuler AJ (2013) Low acetate concentrations favour polyphosphate-accumulating organisms over glycogen accumulating organisms in enhanced biological phosphorus removal from wastewater. Environ Sci Technol 47(8):3816–3824. https://doi.org/10.1021/es304846s
Zheng X, Sun P, Han J, Song Y, Hu Z, Fan H, Lv S (2014) Inhibitory factors affecting the process of enhanced biological phosphorus removal (EBPR)—a mini-review. Process Biochem. https://doi.org/10.1016/j.procbio.2014.10.008
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Šikić, T., Welles, L., Rubio-Rincón, F.J. et al. Assessment of Enhanced Biological Phosphorus Removal Implementation Potential in a Full-Scale Wastewater Treatment Plant in Croatia. Int J Environ Res 13, 1005–1013 (2019). https://doi.org/10.1007/s41742-019-00234-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41742-019-00234-4