Abstract
The Korean or hard-shelled mussel, Mytilus unguiculatus, previously known as Mytilus coruscus, is one of the most economically and ecologically important bivalves in South Korea. However, the population size of this species has drastically reduced owing to overharvesting and habitat shrinkage. Because its genetic information is poorly documented, we contributed, in this study, the genetic diversity and structural analyses of 246 adult samples of M. unguiculatus from seven populations along the coastal areas of the mainland and islands of South Korea using a microsatellite multiplex assay. Genetic diversity analyzed from eleven polymorphic microsatellite loci was consistently moderate (0.50–0.57) in all populations. No recent bottleneck was found, indicating that the number of the studied populations did not decrease to an extent that resulted in a reduction of genetic diversity. Additional tests did not reveal any genetic structure across them, possibly resulting from constant gene flow, strong dispersal of planktonic larvae, and genetic admixture between wild populations. These results suggest that M. unguiculatus populations along the coastal areas of South Korea should be managed as a single unit. Our study provides crucial information for future genetic monitoring, conservation management, and population restoration plan in preparation for the rapid decline in bivalve resources.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Korean or hard-shelled mussel, Mytilus unguiculatus (Valenciennes 1858), previously known as Mytilus coruscus, is a species of marine bivalve mollusks widely distributed in the coastal waters of China, Japan, and Korea. This species is not only one of the most economically important species among the favored bivalves owing to its high nutritional value and large size (An and Lee 2012; Kang et al. 2013b), but also ecologically important in intertidal and subtidal areas (Li et al. 2020). Recently, there has been a drastic decrease of its populations because of overharvesting for food stock, climate change, and habitat competition with an invasive species, M. galloprovincialis (Yi et al. 2021). To sustain and recover its natural populations through effective conservation and management strategies, it is necessary to carry out a comprehensive investigation on population genetics of M. unguiculatus (i.e., genetic diversity and structure).
Among several available molecular markers, microsatellite or simple sequence repeats (SSR) DNA markers (unique sequences of flanking microsatellite regions) have been extensively used for studying population genetics due to their advantageous features such as co-dominant transmission, high level of mutation rate and polymorphism, abundance in various eukaryotic genomes, and ease of use and application (Ellegren 2004; Karhu 2001; Vieira et al. 2016). Commonly, several microsatellite markers are fluorescently labeled and combined with multiplex PCR, allowing multiple microsatellite loci to be simultaneously amplified in a single reaction. Subsequently, the multiplex products are analyzed using gel capillary electrophoresis and then genotyped. The multiplex PCR assay using microsatellite markers, a time and cost-effective technique, has become a powerful tool to study the genetic diversity and structure of natural populations of bivalves including oysters (An et al. 2013; Galvão and Hilsdorf 2015), mussels (Wenne et al. 2022), clams (Hargrove et al. 2015), and scallops (Morvezen et al. 2015). However, there have been limited studies on population genetics of M. unguiculatus.
In this study, we conducted comprehensive genetic diversity and structure analyses of seven M. unguiculatus populations along the coastal areas of the mainland and islands of South Korea with two sets of a multiplex PCR assay (consisting of eight polymorphic microsatellite markers each) from previous studies (Xu et al. 2010; An and Lee 2012; Kang et al. 2013a; Guo et al. 2013; Fu et al. 2018). The findings of this study will provide the crucial basis for developing effective conservation and management strategies for this economically and ecologically important species in South Korea.
Materials and Methods
Sample Collection and Genomic DNA Extraction
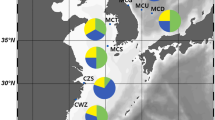
A total of 246 adult samples of M. unguiculatus were collected from seven areas along the coastal coasts of the mainland and islands of South Korea where its major distributions were documented, including the southeast: Hansan-do (HSA), Geomun-do (KOM), Goseong-do (KOS); the west: Sapsi-do (SAB), Socheong-do (SOC); and the east: Uljin (UJI), Dok-do (TOK) from 2021 to 2022 (Table 1, Fig. 1). These sampling sites were determined to postulate the initial hypothesis of geographic isolation by distance. The adductor muscle of each sample was excised with sterile scissors and preserved in absolute alcohol, until its total genomic DNA (gDNA) was extracted following the protocol established by Asahida et al. (1996). Its amount and quality were checked using NanoDrop One Microvolume UV–Vis Spectrophotometer (Thermo Fisher Scientific Inc., Wilmington, DE, USA).
Microsatellite Marker Selection and Genotyping
A total of 65 polymorphic microsatellite markers that had previously been characterized in M. unguiculatus were tested on three random samples from each population. The optimization of PCR settings in multiplex panels was applied to the markers with consistent PCR amplification and obvious allelic size variation. Finally, two multiplex PCR panels (set A and set B) were obtained, each with eight microsatellite markers, generated by Xu et al. (2010), An and Lee (2012), Kang et al. (2013a), Guo et al. (2013), and Fu et al. (2018) (Table S1). The PCR amplification of sixteen microsatellite loci was carried out in a 20-µl volume using AccuPower® PCR PreMix (Bioneer Inc., Daejeon, South Korea) including 1 µl gDNA and each 1 µl forward and reverse primers (the forward primer was fluorescently labeled with FAM, HEX, TAMRA, and ATTO). Amplification was performed in a ProFlex PCR System (Thermo Fisher Scientific Inc.) under the following reaction conditions: initial denaturation at 94 °C for 5 min, 35 cycles of denaturation at 95 °C for 20 s, annealing at 55 °C (Set A) and 50 °C (set B) for 20 s, extension at 72 °C for 30 s, and final extension at 72 °C for 5 min. The fluorescent PCR products were sized and separated on an ABI 3130xl DNA Analyzer (Applied Biosystems, Foster City, CA, USA). Allele sizes were scored against GeneScan 500LIZ Size Standard using GeneMapper v.4.0 (Applied Biosystems).
Statistical Analysis
Genetic Diversity Analysis
Genetic diversity indices at locus and population levels, i.e., number of alleles per locus (Na), effective number of alleles (Ne), observed heterozygosity (Ho), expected heterozygosity (He), polymorphic information content (PIC), and Wright’s inbreeding coefficient (Fis = 1 – Ho / He) were calculated using Cervus v.3.0.7 (Marshall et al. 1998) and GenAlEx v.6.5 (Peakall and Smouse 2006, 2012). Null allele frequencies (Fnull) at each locus and in each population were estimated using the Expectation Maximization (EM) algorithm (Dempster et al. 1977) with 1,000 bootstrap replicates using FreeNA (Chapuis and Estoup 2007). Micro-Checker v.2.2.3 (Oosterhout et al. 2004) was used to evaluate the microsatellite data by identifying genotyping errors, allelic dropouts, and null allele presence. Deviation from Hardy–Weinberg equilibrium (HWE) of each locus across populations was computed using an exact HW test (Guo and Thompson 1992) in Genepop v.4.7.5 (Raymond and Rousset 1995; Rousset 2008). This software was also used for testing the heterozygote excess and deficiency of each locus per population and the linkage disequilibrium (LD) between locus pairs.
Genetic Differentiation and Structure Analysis
Determination of appropriate statistics (allele identity-based FST or its analog, allele size-identity RST) for assessing the genetic differentiation of microsatellite loci was performed using the allele size randomization test (Hardy et al. 2003) implemented in SPAGeDi v. 1.5 (Hardy and Vekemans 2002) with 10,000 permutations. The results showed a nonsignificant mutation effect (RST = 0.001, p = 0.225) on eleven loci, indicating that FST is more reliable than RST to investigate the genetic differentiation among the seven populations. Hence, pairwise FST calculated in Arlequin v. 5.3.2 (Excoffier and Lischer 2010) with 1,000 permutations was used for subsequent analysis. Analysis of molecular variance (AMOVA) to evaluate the genetic differences among three different groups of the seven populations including the southeast: HSA, KOM, KOS; the west: SAB, SOC; and the east: UJI, TOK was implemented in Arlequin v. 5.3.2 with 1,000 permutations.
Structure v. 2.3.4 (Pritchard et al. 2000) was used to identify genetic population structure (K genetic clusters) and assign individuals to those clusters using a Bayesian model-based cluster method. Determination of K (from 2 to 10) was carried out with 10 independent replicates for each K in 100,000 Markov Chain Monte Carlo steps and a burn-in period of 1,000,000. The results of Structure v. 2.3.4 with the most likely K value were verified by an online program, STRUCTURE HARVESTER (Earl and VonHoldt 2012) using the delta K method established by Evanno et al. (2005).
Principal component analysis (PCA) was conducted to identify the major patterns of microsatellite loci across populations using the packages FactoMineR v. 1.3.4 (Le et al. 2008) and Factoextra v.1.0.7 (Kassambara and Mundt 2020) in R software v. 4.2.1 (R Core Team 2022). In addition, GenAlEx v.6.5 was used to calculate the genetic distance matrix according to Nei (1972) and it was subsequently imported to MEGA v. 10.2.2 (Tamura et al. 2021) for constructing an unweighted pair group method with arithmetic mean (UPGMA) dendrogram.
Isolation by Distance
Isolation by distance (IBD) among populations was assessed by regressing the pairwise estimates of FST / (1 – FST) against the logarithm of the pairwise geographic distance (km) (Rousset 1997) using the nonparametric Mantel test with 9,999 permutations implemented in GenAlEx v.6.5.
Bottleneck Analysis
To observe the recent bottleneck events among populations, Wilcoxon signed-rank test established by Luikart (1997) was used with 1,000 iterations in the software package BOTTLENECK v. 1.2.02 (Piry et al. 1999) under three different mutation models: infinite allele model (IAM), two-phased model of mutation (TPM), and stepwise-mutation model (SMM). According to Piry et al. (1999), the recommended parameters for our microsatellite dataset are TPM = 95% SMM and variance among multiple steps of 12. Mode-shift indicators demonstrating the allele frequency distribution could detect any population under mutation-drift equilibrium (approximately L-shape) or recent bottleneck (mode shift or others).
Results
Polymorphic Microsatellite Verification
Sixteen microsatellite markers available in previous studies (Xu et al. 2010; An et al. 2013; Kang et al. 2013a; Fu et al. 2018) tested in this study were characterized across the seven M. unguiculatus populations and summarized in Table S1. All of them successfully produced PCR products for all samples (size ranging from 64 to 376 bp) of the seven populations with no evidence of scoring errors or large allelic dropout indicated by Micro-Checker v.2.2.3. The microsatellite markers were polymorphic, ranging from 3 (P31) to 42 (P05 and P39) in the number of alleles. The greatest Ho and He were found in the locus P05 (0.742 and 0.963, respectively), while these values were lowest in P32 (0.153 and 0.186, respectively). The PIC results showed that eleven loci (P26, P36, P05, P30, P25, P60, P39, P50, P52, P21, and P45) were highly informative (PIC > 0.5), three (P57, P07, and P34) moderately informative (0.25 < PIC < 0.5), and two (P31 and P32) slightly informative (PIC < 0.25). Among them, two loci had negative Fis values (P31 and P34), most likely resulting from their heterozygote deficiency. Seven loci showed a significant excess of homozygosity and were flagged with null allele presence. Among them, two loci (P05 and P30) with null allele frequency (Fnull) lower than 0.2 were retained due to their little influence on the average genetic diversity and structure, whereas five loci (P26, P25, P60, P39, and P21) with Fnull values greater than 0.2 were excluded in the further analysis. There was no genotypic LD for any locus pair across all populations, indicating that each could be treated as an independent genetic marker (data not shown). Nine loci (P26, P05, P57, P30, P25, P60, P39, P21, and P45) significantly deviated from HWE (p < 0.01).
Genetic Diversity
After the polymorphic microsatellite verification, eleven loci (P36, P05, P57, P30, P31, P07, P50, P52, P34, P45, and P32) were finally selected to study the genetic diversity of 246 individuals from the seven M. unguiculatus populations in South Korea. Table 2 summarizes the genetic variation for each locus in each population. The mean number of alleles of all loci (Na) at the population level varied from 7.45 (SOC) to 10.27 (KOM). The mean effective number of loci (Ne) was consistently lower than the mean number of alleles in all populations ranging from 4.12 (SAB) to 4.62 (KOM). There was little discrepancy between the mean Ho and He values among the seven populations. In addition, Ho (0.50–0.57) was found to be constantly lower than He (0.56–0.59) in all populations. As a result, the inbreeding coefficient Fis positively approximated to 0.00 in all populations. The mean value of Ne (5.10) was also lower than that of Na (13.27) in total (grouping all populations into one). The mean values of Ho, He, and Fis in total was 0.52, 0.58, and 0.08, respectively.
Five loci (P05, P57, P30, P52, and P45) out of eleven significantly deviated from HWE across the seven populations. Among them, P05, P30, and P45 showed significant HWE deviation in all populations (except TOK and UJI at locus P30), while the loci P57 and P52 were only significant in two populations (KOM and SAB) and one population (KOM), respectively. Estimation of significant HWE deviation in total was detected at the seven loci (P05, P57, P30, P50, P52, P45, and P32). Most deviations were associated with the statistically significant result of the heterozygote deficit test (p < 0.05) (except for locus P45).
Genetic Differentiation and Structure
The pairwise FST values (Table 3) of the finally selected eleven microsatellite loci showed a very low degree of genetic differentiation among the seven M. unguiculatus populations (ranging from 0.000 to 0.009). Most of them were not statistically significant except for the pair of SOC and HSA (FST = 0.001) and those of TOK and HSA (FST = 0.000).
The AMOVA results performed on the three putative groups of localities, including the southeast: HSA, KOM, KOS; the west: SAB, SOC; and the east: UJI, TOK, were displayed in Table 4. The majority of genetic variation (99.86%) was distributed within populations, whereas genetic variation among the three geographic groups was asymptotic to 0.000. All fixation indices (FST, FSC, and FCT) were not significant and low, indicating that none of the three groups were structured. This result was supported by those from the PCA analysis (Fig. 2).
Principal Component Analysis (PCA) of the seven Mytilus unguiculatus populations based on the finally selected eleven microsatellite loci. Each population is marked by different colors. Each axis corresponds to the principal component (Dim1) factor 1 with 13% total variance and Dim2 with 11% total variance. HSA, Hansan-do; KOM, Geomun-do; KOS, Goseong-do; SAB, Sapsi-do; SOC, Socheong-do; TOK, Dok-do; UJI, Uljin
The Evanno method from STRUCTURE HARVESTER based on the finally selected eleven microsatellite loci across the seven populations suggested that the most likely number of groups for the dataset was K = 3 corresponding to a delta K value of 7.790 (Figure S1). However, the bar plot of K = 3, 4, and 5 similarly showed that all individuals clustered as a mixture, regardless of populations (Figure S2). The Bayesian clustering analysis found no significant structures among the populations.
The genetic distance among the seven populations according to Nei’s method was substantially low (ranging from 0.012 to 0.019) (data not shown). The UPGMA dendrogram based on the Nei’s standard genetic distance of the seven populations (Fig. 3) showed three distinct clusters according to three sampling areas including the southeast: HSA, KOM, KOS; the west: SAB, SOC; and the east: UJI, TOK.
IBD
The regression between pairwise population estimates of FST / (1 – FST) and the logarithms of geographic distance in the seven M. unguiculatus populations (Fig. 4) slightly showed a positive correlation (R2 = 0.0132). However, no statistical significance between them (p = 0.349) was detected from the Mantel test indicating that there was no IBD among them.
Bottleneck Analysis
Under three mutation models (IAM, TPM, and SMM), no significant heterozygosity excess was detected in any of the seven M. unguiculatus populations and in total (p > 0.05) based on the finally selected eleven microsatellite loci. Moreover, the graphical descriptor of the allele distribution exhibited an L-shape, as expected under mutation-drift equilibrium, in all populations and in total (Table 5). These results suggest that there had not been a recent genetic bottleneck in these populations.
Discussion
Genetic variation is of considerable interest for population genetic analysis in natural populations. This estimation, measured by Ho and He, serves as an indicator not only for monitoring but also predicting the maximum amount of genetic variability for population breeding, restoration, and management plans (Allendorf 1986). A high degree of genetic diversity increases evolutionary potential and adaptability to environmental changes, which is crucial for the long-term survival of populations (Zheng et al. 2022). Typically, populations experiencing a recent drastic reduction of effective population size due to sudden climate changes, habitat shrinkage, inbreeding, repeated harvesting, and marine pollution are generally characterized by low genetic diversity (Yi et al. 2021).
In this study, the average genetic diversity estimated from the finally selected eleven microsatellite loci was similarly moderate in the seven M. unguiculatus populations in South Korea. Moreover, no recent genetic bottleneck was detected based on three allele mutation models (IAM, TPM, and SMM) from the Wilcoxon signed-rank test, indicating that sufficient genetic variability persisted across the examined populations under mutation-drift equilibrium, as expected. This result is consistent with a previous report by Kang et al. (2013b). Although a good indicator of genetic diversity was revealed in the studied populations, for reparation of the rapid decline in M. unguiculatus populations, continuous genetic research in these populations is necessary for planning an optimal conservation and management strategies.
Five out of eleven microsatellite loci finally selected in this study showed significant HWE deviations due to heterozygote deficiency commonly detected in aquatic organisms (An et al. 2013; Galvão and Hilsdorf 2015; Gonzalez et al. 2012; Valles-Jimenez et al. 2004; Yu and Li 2007). Many factors could cause these deficit deviations, such as biological factors (inbreeding, Wahlund effect, and gene flow), technical factors (null alleles, technical artifacts, and small sampling size) (Valles-Jimenez et al. 2004). Among them, non-random mating with close relatives (inbreeding) could be ruled out from this study, as there were no genotypic LD among microsatellite loci and a relative frequency of heterozygotes inferred from the quantitative calculation, the confirmation of a lack of bottleneck effect, and mean inbreeding coefficient Fis of 0.000. The presence of null alleles could have caused heterozygote deficits. These deficits are supposedly caused by polymorphic mutations at binding sites of PCR primers resulting from the amplification failure of particular alleles, misscoring due to stutter bands, and slipped-strand mispairing of di-nucleotide repeat loci during PCR (Buttler 2005; Carlsson 2008; Oosterhout et al. 2006; Valles-Jimenez et al. 2004). Although most of these five microsatellite loci carry di-nucleotide repeat motifs, except for P57 (tri-nucleotide motif) and P30 (tetra-nucleotide motif), scoring errors due to stutter bands was not a major cause of heterozygote deficiency, confirmed by our strict procedure of scoring. Alternatively, a random subpopulation structure or small sampling size might be explanations for the HWE deviations.
Absence of geographic barriers in the ocean, large population sizes, and high dispersal potential of the planktonic larvae phase in the life cycle generally leads to a poorly geographical differentiation among populations of marine species, especially bivalves (Launey et al. 2002). This characteristic has been observed in many marine bivalves, such as mussels (Han et al. 2017), pen shells (An et al. 2012), and oysters (An et al. 2014). The pattern of M. unguiculatus population structure varies greatly depending upon hydrological, ecological, and physical barriers, allowing gene flow across populations, which is similar to other bivalves (Palumbi et al. 1997), although its biology and ecology is poorly documented. In this study, the geographic distance between HSA and KOS was short (25.9 km) compared with that between SOC and TOK (630.4 km) among the seven M. unguiculatus populations. However, regardless of distance, low genetic differentiation and weak geographic isolation among them were revealed by the results of the pairwise FST values, PCA, AMOVA, and IBD analyses. These results may be due to persistent gene flow either between wild populations or the passive dispersal of M. unguiculatus planktonic larvae along marine currents, detected through a strong genetic admixture. Such findings are supported by previous studies that used either mitochondrial cytochrome c oxidase I marker (Shen et al. 2009; Yi et al. 2021) or microsatellite markers (Kang et al. 2013b) on M. unguiculatus populations in South Korea and China. Absence of genetic structure detected in this study suggests that different M. unguiculatus populations across South Korea should be managed as a single unit.
In order to investigate genetic differentiation among M. unguiculatus populations, Nei’s standard genetic distance, closely related to IAM, a size-independent mutation model where each mutation is likely to produce an allele of any size, is usually used (Kimura and Crow 1964; Wang et al. 2001). Many previous studies have used this parameter to infer population differentiation based on microsatellite data (Alam and Islam 2005; Ball and Chapman 2003; Barat et al. 2015; Chen et al. 2005; Gonzalez et al. 2012; Kim et al. 2012). However, calculation of genetic differentiation from microsatellite data based on IAM instead of using a size-dependence mutation model such as SMM is supposedly not appropriate (Slatkin 1995). Hence, selection of a suitable mutation model for a microsatellite dataset remains controversial. In this study, Nei’s standard genetic distance was significantly low, but showed that the seven M. unguiculatus populations were clustered into the three groups according to the three geographic areas, fitting the initial hypothesis of geographic separation. This conclusion should be considered carefully because of a significantly low genetic variation among populations or unfit selection of a mutation model. However, our results may be a valuable reference for a future sampling plan to explore the genetic structure of M. unguiculatus populations in South Korea.
Conclusions
The seven M. unguiculatus populations from the three geographic groups, i.e., the southeast, the west, and the east of South Korea consistently maintained a moderate genetic diversity. Moreover, the low genetic differentiation and weak geographic separation were detected across them, suggesting that different populations along the coastal areas of the mainland and islands of South Korea should be managed as a single unit. Such findings contribute essential information for future resource management and population restoration, thus aiding optimizing inbreeding programs for this economically and ecologically important marine bivalve species.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon requests.
References
Alam MS, Islam MS (2005) Population genetic structure of Catla catla (Hamilton) revealed by microsatellite DNA markers. Aquaculture 246(1–4):151–160. https://doi.org/10.1016/j.aquaculture.2005.02.012
Allendorf FW (1986) Genetic drift and the loss of alleles versus heterozygosity. Zoo Biol 5(2):181–190. https://doi.org/10.1002/zoo.1430050212
An HS, Lee JW (2012) Development of microsatellite markers for the Korean mussel, Mytilus coruscus (Mytilidae) using next-generation sequencing. Int J Mol Sci 13(8):10583–10593. https://doi.org/10.3390/ijms130810583
An HS, Lee JW, Dong CM (2012) Population genetic structure of Korean pen shell (Atrina pectinata) in Korea inferred from microsatellite marker analysis. Genes Genom 34(6):681–688. https://doi.org/10.1007/s13258-012-0091-z
An HS, Lee JW, Kim WJ, Lim HJ, Kim EM, Byun SG, Hur YB, Park JY, Myeong JI, An CM (2013) Comparative genetic diversity of wild and hatchery-produced Pacific oyster (Crassostrea gigas) populations in Korea using multiplex PCR assays with nine polymorphic microsatellite markers. Genes Genom 35(6):805–815. https://doi.org/10.1007/s13258-013-0140-2
Asahida T, Kobayashi T, Saitoh K, Nakayama I (1996) Tissue preservation and total DNA extraction form fish stored at ambient temperature using buffers containing high concentration of urea. Fish Sci 62(5):727–730. https://doi.org/10.2331/fishsci.62.727
Ball AO, Chapman RW (2003) Population genetic analysis of white shrimp, Litopenaeus setiferus, using microsatellite genetic markers. Mol Ecol 12(9):2319–2330. https://doi.org/10.1046/j.1365-294X.2003.01922.x
Barat A, Sahoo PK, Kumar R, Mir JI, Ali S, Patiyal RS, Singh AK (2015) Molecular characterization of rainbow trout, Oncorhynchus mykiss (Walbaum, 1792) stocks in India. J Genet 94(2):13–18. https://doi.org/10.1007/s12041-015-0483-3
Butler JM (2005) Forensic DNA typing: biology, technology, and genetics of STR markers. Elsevier Academic Press, Massachusetts
Carlsson J (2008) Effects of microsatellite null alleles on assignment testing. J Hered 99(6):616–623. https://doi.org/10.1093/jhered/esn048
Chapuis MP, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24(3):621–631. https://doi.org/10.1093/molbev/msl191
Chen JP, Sun DJ, Dong CZ, Liang B, Wu WH, Zhang SY (2005) Genetic analysis of four wild chum salmon Oncorhynchus keta populations in China based on microsatellite markers. Environ Biol Fishes 73(2):181–188. https://doi.org/10.1007/s10641-004-6114-0
Dempster AP, Laird NM, Rubin DB (1977) Maximum likelihood from incomplete data via the EM algorithm. J R Stat Soc Ser B Methodol 39(1):1–22. https://doi.org/10.1111/j.2517-6161.1977.tb01600.x
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361. https://doi.org/10.1007/s12686-011-9548-7
Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5(6):435–445. https://doi.org/10.1038/nrg1348
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10(3):564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fu Z, Tian Y, Ye Y, Qi P, Wu C (2018) Development and characterization of twenty-three novel polymorphic microsatellite markers for mussel, Mytilus coruscus. Pak J Zool 50(4):1541–1543. https://doi.org/10.17582/journal.pjz/2018.50.4.sc3
Galvão MSN, Hilsdorf AWS (2015) Assessing the genetic diversity of the mangrove oyster Crassostrea rhizophorae (Bivalvia, Ostreidae) by microsatellite markers in southeastern Brazil. Mar Biol Res 11(9):944–954. https://doi.org/10.1080/17451000.2015.1047381
Gonzalez EB, Aritaki M, Taniguchi N (2012) Microsatellite multiplex panels for population genetic analysis of red sea bream Pagrus major. Fish Sci 78(3):603–611. https://doi.org/10.1007/s12562-012-0469-5
Guo BY, Wu CW, Liu CL, Zhu AY, Ye YY (2013) Development of microsatellite markers for Mytilus coruscus (Mytilidae), an economically important mussel in the East China Sea. Genet Mol Res 12(3):2670–2673. https://doi.org/10.3390/ijms130810583
Guo SW, Thompson EA (1992) Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics 48(2):361–372. https://doi.org/10.2307/2532296
Han Z, Mao Y, Shui B, Yanagimoto T, Gao T (2017) Genetic structure and unique origin of the introduced blue mussel Mytilus galloprovincialis in the north-western Pacific: clues from mitochondrial cytochrome c oxidase I (COI) sequences. Mar Freshw Res 68(2):263–269. https://doi.org/10.1071/MF15186
Hardy OJ, Charbonnel N, Fréville H, Heuertz M (2003) Microsatellite allele sizes: a simple test to assess their significance on genetic differentiation. Genetics 163(4):1467–1482. https://doi.org/10.1093/genetics/163.4.1467
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2(4):618–620. https://doi.org/10.1046/j.1471-8286.2002.00305.x
Hargrove JS, Sturmer L, Scarpa J, Austin JD (2015) Assessment of genetic diversity in wild and aquaculture stocks of Mercenaria mercenaria in Florida. J Shellfish Res 34(2):55–365. https://doi.org/10.2983/035.034.0218
Kang JH, Kim YK, Park JY, Noh ES, Jeong JE, Lee YS, Choi TJ (2013a) Development of microsatellite markers for a hard-shelled mussel, Mytilus coruscus, and cross-species transfer. Genet Mol Res 12(3):4009–4017. https://doi.org/10.4238/2013.September.27.2
Kang JH, Lee JM, Noh ES, Park JY, An CM (2013b) Genetic characterization of Mytilus coruscus and M. galloprovincialis using microsatellite markers. Genet Mol Res 12(4):5494–5505. https://doi.org/10.4238/2013.November.13.2
Karhu A (2001) Evolution and applications of pine microsatellites. Dissertation. University of Oulu
Kassambara A, Mundt F (2020) Factoextra: extract and visualize the results of multivariate data analyses. R Package version 1.0.7. The Comprehensive R Archive Network. https://CRAN.R-project.org/package=factoextra. Accessed 12 Jan 2023
Kim JK, Min GS, Yoon M, Kim Y, Choi JH, Oh TY, Ni Y (2012) Genetic structure of Larimichthys polyactis (Pisces: Sciaenidae) in the Yellow and East China Seas inferred from microsatellite and mitochondrial DNA analyses. Anim Cells Syst 16(4):313–320. https://doi.org/10.1080/19768354.2011.652668
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics 49(4):725. https://doi.org/10.1093/genetics/49.4.725
Launey S, Ledu C, Boudry P, Bonhomme F, Naciri-Graven Y (2002) Geographic structure in the European flat oyster (Ostrea edulis L.) as revealed by microsatellite polymorphism. J Hered 93(5):331–351. https://doi.org/10.1093/jhered/93.5.331
Le S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18. https://doi.org/10.18637/jss.v025.i01
Li R, Zhang W, Lu J, Zhang Z, Mu C, Song W, Migaud H, Wang C, Bekaert M (2020) The whole-genome sequencing and hybrid assembly of Mytilus coruscus. Front Genet 11:440. https://doi.org/10.3389/fgene.2020.00440
Luikart GH (1997) Usefulness of molecular markers for detecting population bottlenecks and monitoring genetic change. Dissertation. University of Montana
Marshall TC, Slate JEBK, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7(5):639–655. https://doi.org/10.1046/j.1365-294x.1998.00374.x
Morvezen R, Charrier G, Boudry P, Chauvaud L, Breton F, Strand Ø, Laroche J (2015) Genetic structure of a commercially exploited bivalve, the great scallop Pecten maximus, along the European coasts. Conserv Genet 17(1):57–67. https://doi.org/10.1007/s10592-015-0760-y
Nei M (1972) Genetic distance between populations. Am Nat 106(949):283–292
Oosterhout CV, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Oosterhout CV, Weetman D, Hutchinson WF (2006) Estimation and adjustment of microsatellite null alleles in nonequilibrium populations. Mol Ecol Notes 6(1):255–256. https://doi.org/10.1111/j.1471-8286.2005.01082.x
Palumbi SR, Grabowsky G, Duda T, Geyer L, Tachino N (1997) Speciation and population genetic structure in tropical Pacific sea urchins. Evolution 51(5):1506–1517. https://doi.org/10.1111/j.1558-5646.1997.tb01474.x
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective size using allele frequency data. J Hered 90(4):502–503. https://doi.org/10.1093/jhered/90.4.502
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org. Accessed 12 Jan 2023
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86(3):248–249. https://doi.org/10.1093/oxfordjournals.jhered.a111573
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145(4):1219–1228. https://doi.org/10.1093/genetics/145.4.1219
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8(1):103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Shen YB, Li JL, Feng BB (2009) Genetic analysis of cultured and wild populations of Mytilus coruscus based on mitochondrial DNA. Zool Res 30(3):240–246. https://doi.org/10.3724/SP.J.1141.2009.03240
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequencies. Genetics 139(1):457–462. https://doi.org/10.1093/genetics/139.1.457
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38(7):3022–3027. https://doi.org/10.1093/molbev/msab120
Valles-Jimenez R, Cruz P, Perez-Enriquez R (2004) Population genetic structure of Pacific white shrimp (Litopenaeus vannamei) from Mexico to Panama: microsatellite DNA variation. Mar Biotechnol 6(5):475–484. https://doi.org/10.1007/s10126-004-3138-6
Vieira MLC, Santini L, Diniz AL, Munhoz CDF (2016) Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol 39:312–328. https://doi.org/10.1590/1678-4685-GMB-2016-0027
Wang R, Zheng L, Touré YT, Dandekar T, Kafatos FC (2001) When genetic distance matters: measuring genetic differentiation at microsatellite loci in whole-genome scans of recent and incipient mosquito species. Proc Natl Acad Sci 98(19):10769–10774. https://doi.org/10.1073/pnas.191003598
Wenne R, Zbawicka M, Prądzińska A, Kotta J, Herkül K, Gardner JPA, Apostolidis AP, Poćwierz-Kotus A, Rouane-Hacene O, Korrida A, Dondero F, Baptista M, Reizopoulou S, Hamer B, Sundsaasen KK, Árnyasi M, Kent MP (2022) Molecular genetic differentiation of native populations of Mediterranean blue mussels, Mytilus galloprovincialis Lamarck, 1819, and the relationship with environmental variables. Eur Zool J 89(1):755–784. https://doi.org/10.1080/24750263.2022.2086306
Xu TJ, Sun YN, Yuan YT, Liao Z, Wang RX (2010) Isolation and characterization of polymorphic microsatellite loci in the hard-shelled mussel, Mytilus coruscus (Mytilidae). Genet Mol Res 9(3):1388–1391. https://doi.org/10.4238/vol9-3gmr856
Yi CH, Yoon M, Kim JM, Kim IH, Cho IY, An HS (2021) Genetic analysis and population genetic structure of hard-shelled mussel, Mytilus coruscus Gould 1861 (Mytiloida: Mytilidae) from the coasts of South Korea based on mitochondrial cytochrome oxidase (COI) gene sequences. Genes Genom 43(6):577–585. https://doi.org/10.1007/s13258-021-01073-4
Yu H, Li Q (2007) Genetic variation of wild and hatchery populations of the Pacific oyster Crassostrea gigas assessed by microsatellite markers. J Genet Genom 34(12):1114–1122. https://doi.org/10.1016/S1673-8527(07)60127-1
Zheng J, Yan Y, Li Z, Song N (2022) Genetic structure of the small yellow croaker (Larimichthys polyactis) across the Yellow Sea and the East China Sea by microsatellite DNA variation: implications for the division of management units. PeerJ 10:e13789. https://doi.org/10.7717/peerj.13789
Funding
This work was funded by National Marine Biodiversity Institute of Korea (MABIK) under in-house Research Programs [2023M00400].
Author information
Authors and Affiliations
Contributions
Conceptualization: Moongeun Yoon, Keun-Yong Kim Methodology: Moongeun Yoon, Jung Soo Heo, Jiyoung Woo, Biet Thanh Tran Formal analysis and investigation: Jiyoung Woo, Jung Soo Heo, Biet Thanh Tran Writing-original draft preparation: Jiyoung Woo, Biet Thanh Tran Writing-review and editing: Hyun-Ju Hwang, Keun-Yong Kim, Keun-Sik Kim, Insong Koh Funding acquisition: Moongeun Yoon Resources: Kook Hee Kang, Jeong Su Park, Ki-Woong Nam Supervision: Insong Koh, Biet Thanh Tran.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent to Publish
Not applicable.
Research Involving Human Participants and/or Animals
This article does not contain any studies involving human participants performed by any of the authors. All experiments were performed following the guidelines of the Korean Association for Laboratory Animals (Approval no. 18–0680, 5 October 2018).
Conflicts of Interest
The authors have no conflicts of interest to declare.
Competing Interests
The authors declare that they have no competing of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Woo, J., Heo, J.S., Kim, KY. et al. Population Genetic Analysis of the Wild Hard-shelled Mussel, Mytilus Unguiculatus (Valenciennes 1858) in South Korea Using a Microsatellite Multiplex Assay. Thalassas 39, 931–942 (2023). https://doi.org/10.1007/s41208-023-00565-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41208-023-00565-2