Abstract
Purpose
Numerous strategies and diagnostic tests were proposed in patients suspected of clinically significant (cs) prostate cancer (PCa) after an initial negative prostate biopsy. The study aimed to create a Random Forest (RF) classifier for predicting the probability of csPCa in specimens taken by the repeated systematic prostate biopsy (SBx), and to determine its diagnostic accuracy and clinical utility.
Methods
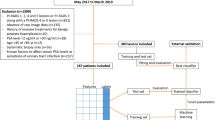
This retrospective, single-center study included patients who underwent repeated SBx due to clinical suspicion of cancer. Data on patient age, serum prostate-specific antigen (PSA) levels, prostate volume, digital rectal examination, first-degree family history, and histology findings from the SBx were collected for all patients. The area under the curve (AUC), and secondary metrics of clinical prediction models were used to assess their discriminative abilities. Clinical usefulness of final model was tested by the decision curve analysis (DCA). The explainability and website placement of the ML model were also performed.
Results
In total, 204 patients were eligible for analysis. The csPCa was detected in 26% (n = 53) patients. The AUC, accuracy, sensitivity, and specificity for detection of csPCa were 0.94, 0.91, 0.84, and 0.98, respectively. With an optimal threshold of 0.8, about 34% of unnecessary biopsies would be avoided, but correct diagnosis would be delayed in 4.4% csPC cases. PSA level, prostate volume, and age were the top-ranked variables in the RF model.
Conclusion
The RF classifier predicts csPCa with good accuracy and may help urologists when deciding whether the repeated biopsy is necessary to avoid being too invasive.





Similar content being viewed by others
References
Rawla, P. (2019). Epidemiology of prostate Cancer. World J Oncol, 10(2), 63–89.
Loeb, S. (2017). When is a negative prostate biopsy really negative? Repeat biopsies in detection and active surveillance. Journal Of Urology, 197(4), 973–974.
Kohaar, I., Chen, Y., Banerjee, S., et al. (2021). A urine exosome gene expression panel distinguishes between indolent and aggressive prostate cancers at Biopsy. Journal Of Urology, 205(2), 420–425.
Cheung, D. C., Li, J., & Finelli, A. (2018). A narrative review and update on management following negative prostate biopsy. CurrOpinUrol, 28(4), 398–402.
Scattoni, V., Russo, A., Di Trapani, E., et al. (2014). Repeated biopsy in the detection of prostate cancer: when and how many cores. Arch Ital UrolAndrol, 30(4), 311–313.
Long, X., Wu, L., Zeng, X., et al. (2020). Biomarkers in previous histologically negative prostate biopsies can be helpful in repeat biopsy decision-making processes. Cancer Medicine, 9(20), 7524–7536.
Uhr, A., Glick, L., & Gomella, L. G. (2020). An overview of biomarkers in the diagnosis and management of prostate cancer. The Canadian Journal Of Urology, 27(S3), 24–27.
Giganti, F., & Moore, C. M. (2017). A critical comparison of techniques for MRI-targeted biopsy of the prostate. TranslAndrolUrol, 6(3), 432–443.
Radtke, J. P., Wiesenfarth, M., Kesch, C., et al. (2017). Combined clinical parameters and multiparametric magnetic resonance imaging for Advanced Risk modeling of prostate Cancer-patient-tailored risk stratification can reduce unnecessary biopsies. EurUrol, 72(6), 888–896.
Truong, M., Wang, B., Gordetsky, J. B., et al. (2018). Multi-institutional nomogram predicting benign prostate pathology on magnetic resonance/ultrasound fusion biopsy in men with a prior negative 12-core systematic biopsy. Cancer, 124(2), 278–285.
Huang, C., Song, G., Wang, H., et al. (2018). MultiParametric Magnetic Resonance Imaging-Based Nomogram for Predicting Prostate Cancer and Clinically Significant Prostate Cancer in Men Undergoing Repeat Prostate Biopsy, Biomed Res Int, 2018:6368309.
Alberts, A. R., Roobol, M. J., Verbeek, J. F. M., et al. (2019). Prediction of High-grade Prostate Cancer Following Multiparametric Magnetic Resonance Imaging: Improving the Rotterdam European Randomized Study of Screening for Prostate Cancer Risk Calculators, EurUrol,75(2):310–318.
Oishi, M., Shin, T., Ohe, C., et al. (2019). Which patients with negative magnetic resonance imaging can safely avoid biopsy for prostate Cancer? Journal Of Urology, 201(2), 268–276.
Schoots, I. G., & Roobol, M. J. (2020). Multivariate risk prediction tools including MRI for individualized biopsy decision in prostate cancer diagnosis: current status and future directions. World Journal Of Urology, 38(3), 517–529.
Xiao, L. H., Chen, P., Gou, Z. P., et al. (2017). Prostate cancer prediction using the random forest algorithm that takes into account transrectal ultrasound findings, age, and serum levels of prostate-specific antigen. Asian Journal Of Andrology, 19(5), 586–590.
Wang, G., Teoh, J. Y., & Choi, K. S. (2018). Diagnosis of prostate cancer in a Chinese population by using machine learning methods. AnnuIntConf IEEE Eng Med BiolSoc, 2018:1–4.
Chiu, P. K., Shen, X., Wang, G., et al. (2021). Enhancement of prostate cancer diagnosis by machine learning techniques: an algorithm development and validation study. Prostate Cancer And Prostatic Diseases. doi: https://doi.org/10.1038/s41391-021-00429-x Epub ahead of print.
Bernatz, S., Ackermann, J., Mandel, P., et al. (2020). Comparison of machine learning algorithms to predict clinically significant prostate cancer of the peripheral zone with multiparametric MRI using clinical assessment categories and radiomic features. EurRadiol, 30(12), 6757–6769.
Toth, R., Schiffmann, H., Hube-Magg, C., et al. (2019). Random forest-based modelling to detect biomarkers for prostate cancer progression. Clin Epigenetics, 11(1), 148.
Roobol, M. J., van Vugt, H. A., Loeb, S., et al. (2012). Prediction of prostate cancer risk: the role of prostate volume and digital rectal examination in the ERSPC risk calculators. EurUrol, 61(3), 577–583.
September, H. O., & .ai, Distributed Random Forest (DRF). (2021). URLhttps://docs.h2o.ai/h2o/latest-stable/h2o-docs/data-science/drf.html H2O version 3.34.0.1.
Huang, Y., Li, W., Macheret, F., et al. (2020). A tutorial on calibration measurements and calibration models for clinical prediction models. Journal Of The American Medical Informatics Association, 27(4), 621–633.
Vickers, A. J., Van Calster, B., & Steyerberg, E. W. (2016). Net benefit approaches to the evaluation of prediction models, molecular markers, and diagnostic tests. Bmj, 352, i6.
Kerr, K. F., Brown, M. D., Zhu, K., et al. (2016). Assessing the clinical impact of risk prediction models with decision curves: Guidance for correct interpretation and appropriate use. J ClinOncol, 34(21), 2534–2540.
Lundberg, S. M., & Lee, S. I. (2017). A unified approach to interpreting model predictions,Adv Neural Inf Process Syst,https://arxiv.org/pdf/1705.07874.pdf
Kirby, R., & Fitzpatrick, J. M. (2012). Optimising repeat prostate biopsy decisions and procedures. Bju International, 109(12), 1750–1754.
Naji, L., Randhawa, H., Sohani, Z., et al. (2018). Digital rectal examination for prostate Cancer screening in primary care: a systematic review and Meta-analysis. Annals Of Family Medicine, 16(2), 149–154.
Barber, L., Gerke, T., Markt, S. C., et al. (2018). Family history of breast or prostate Cancer and prostate Cancer risk. Clinical Cancer Research, 24(23), 5910–5917.
Parikh, R. B., Manz, C., Chivers, C., et al. (2019). Machine learning approaches to predict 6-Month Mortality among patients with Cancer. JAMA Netw Open, 2(10), e1915997.
Le Dell, E. (2018). useR! Machine Learning Tutorial, URL, https://koalaverse.github.io/machine-learning-in-R/
Brownlee, J. (2017). What is the Difference Between Test and Validation Datasets? URL, https://machinelearningmastery.com/difference-test-validation-datasets/
Oleszak, M. (2020). Calibrating classifiers Are you sure your model returns probabilities?URL, https://towardsdatascience.com/calibrating-classifiers-559abc30711a
Dankowski, T., & Ziegler, A. (2016). Calibrating random forests for probability estimation. Statistics In Medicine, 35(22), 3949–3960.
Acknowledgements
The authors were financially supported through a research grant N0175014, N175007and III 41007of the Ministry of Science and Technological Development of Serbia and Grants OI174028 from the City of Kragujevac. The authors thank the Ministry for this support.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by [Miroslav Stojadinovic] and [Bogdan Milicevic]. The first draft of the manuscript was written by [Miroslav Stojadinovic]. Supervision: [Slobodan Jankovic].All authors commented on the previous versions of the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stojadinovic, M., Milicevic, B. & Jankovic, S. Improved Prediction of Significant Prostate Cancer Following Repeated Prostate Biopsy by the Random Forest Classifier. J. Med. Biol. Eng. 43, 83–92 (2023). https://doi.org/10.1007/s40846-022-00768-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40846-022-00768-7