Abstract
Purpose of Review
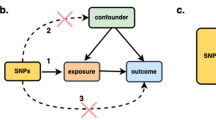
In this paper, we summarize prior studies that have used Mendelian randomization (MR) methods to study the effects of exposures, lifestyle factors, physical traits, and/or biomarkers on cancer risk in humans. Many such risk factors have been associated with cancer risk in observational studies, and the MR approach can be used to provide evidence as to whether these associations represent causal relationships. MR methods require a risk factor of interest to have known genetic determinants that can be used as proxies for the risk factor (i.e., “instrumental variables” or IVs), and these can be used to obtain an effect estimate that, under certain assumptions, is not prone to bias caused by unobserved confounding or reverse causality. This review seeks to describe how MR studies have contributed to our understanding of cancer causation.
Recent Findings
We searched the published literature and identified 76 MR studies of cancer risk published prior to October 31, 2017. Risk factors commonly studied included alcohol consumption, vitamin D, anthropometric traits, telomere length, lipid traits, glycemic traits, and markers of inflammation. Risk factors showing compelling evidence of a causal association with risk for at least one cancer type include alcohol consumption (for head/neck and colorectal), adult body mass index (increases risk for multiple cancers, but decreases risk for breast), height (increases risk for breast, colorectal, and lung; decreases risk for esophageal), telomere length (increases risk for lung adenocarcinoma, melanoma, renal cell carcinoma, glioma, B-cell lymphoma subtypes, chronic lymphocytic leukemia, and neuroblastoma), and hormonal factors (affects risk for sex steroid-sensitive cancers).
Summary
This review highlights alcohol consumption, body mass index, height, telomere length, and the hormonal exposures as factors likely to contribute to cancer causation. This review also highlights the need to study specific cancer types, ideally subtypes, as the effects of risk factors can be heterogeneous across cancer types. As consortia-based genome-wide association studies increase in sample size and analytical methods for MR continue to become more sophisticated, MR will become an increasingly powerful tool for understanding cancer causation.

Similar content being viewed by others
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Didelez V, Sheehan NA. Mendelian randomization as an instrumental variable approach to causal inference. Stat Methods Med Res. 2007;16(4):309–30.
VanderWeele TJ, Tchetgen Tchetgen EJ, Cornelis M, Kraft P. Methodological challenges in mendelian randomization. Epidemiol (Cambridge, Mass). 2014;25:427–35. https://doi.org/10.1097/EDE.0000000000000081.
Zheng J, Baird D, Borges M-C, Bowden J, Hemani G, Haycock P, et al. Recent developments in Mendelian randomization studies. Curr Epidemiol Rep. 2017;4:330–45. https://doi.org/10.1007/s40471-017-0128-6.
Pierce B, Burgess S. Efficient design for Mendelian randomization studies: subsample and two-sample instrumental variable estimators. Am J Epidemiol. 2013;177:S117-S.
Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013;37:658–65. https://doi.org/10.1002/gepi.21758.
Lewis SJ, Davey Smith G. Alcohol, ALDH2, and esophageal cancer: a meta-analysis which illustrates the potentials and limitations of a Mendelian randomization approach. Cancer Epidemiol Biomark Prev. 2005;14:1967–71. https://doi.org/10.1158/1055-9965.EPI-05-0196.
Boccia S, Hashibe M, Galli P, De Feo E, Asakage T, Hashimoto T, et al. Aldehyde dehydrogenase 2 and head and neck cancer: a meta-analysis implementing a Mendelian randomization approach. Cancer Epidemiol Biomark Prev. 2009;18(1):248–54.
Wang J, Wang H, Chen Y, Hao P, Zhang Y. Alcohol ingestion and colorectal neoplasia: a meta-analysis based on a Mendelian randomization approach. Color Dis. 2011;13:e71–e8. https://doi.org/10.1111/j.1463-1318.2010.02530.x.
Bonilla C, Gilbert R, Kemp JP, Timpson NJ, Evans DM, Donovan JL, et al. Using genetic proxies for lifecourse sun exposure to assess the causal relationship of sun exposure with circulating vitamin D and prostate cancer risk. Cancer Epidemiol Biomark Prev. 2013;22:597–606. https://doi.org/10.1158/1055-9965.EPI-12-1248.
Theodoratou E, Palmer T, Zgaga L, Farrington SM, McKeigue P, Din FVN, et al. Instrumental variable estimation of the causal effect of plasma 25-hydroxy-vitamin D on colorectal cancer risk: a Mendelian randomization analysis. PLoS One. 2012;7:e37662. https://doi.org/10.1371/journal.pone.0037662.
Wang S, Huo D, Kupfer S, Alleyne D, Ogundiran TO, Ojengbede O, et al. Genetic variation in the vitamin D related pathway and breast cancer risk in women of African ancestry in the root consortium. Int J Cancer. 2017;142:36–43. https://doi.org/10.1002/ijc.31038.
Dimitrakopoulou VI, Tsilidis KK, Haycock PC, Dimou NL, Al-Dabhani K, Martin RM, et al. Circulating vitamin D concentration and risk of seven cancers: Mendelian randomisation study. BMJ. 2017;359:j4761. https://doi.org/10.1136/bmj.j4761.
Ong J-S, Cuellar-Partida G, Lu Y, Fasching PA, Hein A, Burghaus S, et al. Association of vitamin D levels and risk of ovarian cancer: a Mendelian randomization study. Int J Epidemiol. 2016;45:1619–30. https://doi.org/10.1093/ije/dyw207.
Collin SM, Metcalfe C, Palmer TM, Refsum H, Lewis SJ, Smith GD, et al. The causal roles of vitamin B(12) and transcobalamin in prostate cancer: can Mendelian randomization analysis provide definitive answers? Int J Mol Epidemiol Genet. 2011;2:316–27.
Khankari NK, Murff HJ, Zeng C, Wen W, Eeles RA, Easton DF, et al. Polyunsaturated fatty acids and prostate cancer risk: a Mendelian randomisation analysis from the PRACTICAL consortium. Br J Cancer. 2016;115:624–31. https://doi.org/10.1038/bjc.2016.228.
Wang C, Qin N, Zhu M, Chen M, Xie K, Cheng Y, et al. Metabolome-wide association study identified the association between a circulating polyunsaturated fatty acids variant rs174548 and lung cancer. Carcinogenesis. 2017;38:1147–54. https://doi.org/10.1093/carcin/bgx084.
May-Wilson S, Sud A, Law PJ, Palin K, Tuupanen S, Gylfe A, et al. Pro-inflammatory fatty acid profile and colorectal cancer risk: a Mendelian randomisation analysis. Eur J Cancer (Oxford, England: 1990). 2017;84:228–38. https://doi.org/10.1016/j.ejca.2017.07.034.
Taylor AE, Martin RM, Geybels MS, Stanford JL, Shui I, Eeles R, et al. Investigating the possible causal role of coffee consumption with prostate cancer risk and progression using Mendelian randomization analysis. Int J Cancer. 2017;140:322–8. https://doi.org/10.1002/ijc.30462.
Timpson NJ, Brennan P, Gaborieau V, Moore L, Zaridze D, Matveev V, et al. Can lactase persistence genotype be used to reassess the relationship between renal cell carcinoma and milk drinking? Potentials and problems in the application of Mendelian randomization. Cancer Epidemiol Biomark Prev. 2010;19:1341–8. https://doi.org/10.1158/1055-9965.EPI-09-1019.
Bergholdt HKM, Nordestgaard BG, Varbo A, Ellervik C. Lactase persistence, milk intake, and mortality in the Danish general population: a Mendelian randomization study. Eur J Epidemiol. 2017; https://doi.org/10.1007/s10654-017-0328-x.
Thrift AP, Gong J, Peters U, Chang-Claude J, Rudolph A, Slattery ML, et al. Mendelian randomization study of body mass index and colorectal cancer risk. Cancer Epidemiol Biomark Prev. 2015;24:1024–31. https://doi.org/10.1158/1055-9965.EPI-14-1309.
Gao C, Patel CJ, Michailidou K, Peters U, Gong J, Schildkraut J, et al. Mendelian randomization study of adiposity-related traits and risk of breast, ovarian, prostate, lung and colorectal cancer. Int J Epidemiol. 2016;45:896–908. https://doi.org/10.1093/ije/dyw129.
Jarvis D, Mitchell JS, Law PJ, Palin K, Tuupanen S, Gylfe A, et al. Mendelian randomisation analysis strongly implicates adiposity with risk of developing colorectal cancer. Br J Cancer. 2016;115:266–72. https://doi.org/10.1038/bjc.2016.188.
Dixon SC, Nagle CM, Thrift AP, Pharoah PD, Pearce CL, Zheng W, et al. Adult body mass index and risk of ovarian cancer by subtype: a Mendelian randomization study. Int J Epidemiol. 2016;45:884–95. https://doi.org/10.1093/ije/dyw158.
Carreras-Torres R, Haycock PC, Relton CL, Martin RM, Smith GD, Kraft P, et al. The causal relevance of body mass index in different histological types of lung cancer: a Mendelian randomization study. Sci Rep. 2016;6:31121. https://doi.org/10.1038/srep31121.
Carreras-Torres R, Johansson M, Haycock PC, Wade KH, Relton CL, Martin RM, et al. Obesity, metabolic factors and risk of different histological types of lung cancer: a Mendelian randomization study. PLoS One. 2017;12:e0177875. https://doi.org/10.1371/journal.pone.0177875.
Painter JN, O'Mara TA, Marquart L, Webb PM, Attia J, Medland SE, et al. Genetic risk score Mendelian randomization shows that obesity measured as body mass index, but not waist:hip ratio, is causal for endometrial cancer. Cancer Epidemiol Biomark Prev. 2016;25:1503–10. https://doi.org/10.1158/1055-9965.EPI-16-0147.
Nead KT, Sharp SJ, Thompson DJ, Painter JN, Savage DB, Semple RK, et al. Evidence of a causal association between insulinemia and endometrial cancer: a Mendelian randomization analysis. J Natl Cancer Inst. 2015;107 https://doi.org/10.1093/jnci/djv178.
Brennan P, McKay J, Moore L, Zaridze D, Mukeria A, Szeszenia-Dabrowska N, et al. Obesity and cancer: Mendelian randomization approach utilizing the FTO genotype. Int J Epidemiol. 2009;38:971–5. https://doi.org/10.1093/ije/dyp162.
Mao Y, Yan C, Lu Q, Zhu M, Yu F, Wang C, et al. Genetically predicted high body mass index is associated with increased gastric cancer risk. Eur J Hum Genet. 2017;25:1061–6. https://doi.org/10.1038/ejhg.2017.103.
Carreras-Torres R, Johansson M, Gaborieau V, Haycock PC, Wade KH, Relton CL, et al. The role of obesity, type 2 diabetes, and metabolic factors in pancreatic cancer: a Mendelian randomization study. J Natl Cancer Inst. 2017;109 https://doi.org/10.1093/jnci/djx012.
Thrift AP, Shaheen NJ, Gammon MD, Bernstein L, Reid BJ, Onstad L, et al. Obesity and risk of esophageal adenocarcinoma and Barrett’s esophagus: a Mendelian randomization study. J Natl Cancer Inst. 2014;106 https://doi.org/10.1093/jnci/dju252.
Guo Y, Warren Andersen S, Shu X-O, Michailidou K, Bolla MK, Wang Q, et al. Genetically predicted body mass index and breast cancer risk: Mendelian randomization analyses of data from 145,000 women of European descent. PLoS Med. 2016;13:e1002105. https://doi.org/10.1371/journal.pmed.1002105.
Davies NM, Gaunt TR, Lewis SJ, Holly J, Donovan JL, Hamdy FC, et al. The effects of height and BMI on prostate cancer incidence and mortality: a Mendelian randomization study in 20,848 cases and 20,214 controls from the PRACTICAL consortium. Cancer Causes Control. 2015;26:1603–16. https://doi.org/10.1007/s10552-015-0654-9.
Benn M, Tybjaerg-Hansen A, Smith GD, Nordestgaard BG. High body mass index and cancer risk—a Mendelian randomisation study. Eur J Epidemiol. 2016;31:879–92. https://doi.org/10.1007/s10654-016-0147-5.
Zhang B, Shu X-O, Delahanty RJ, Zeng C, Michailidou K, Bolla MK, et al. Height and breast cancer risk: evidence from prospective studies and Mendelian randomization. J Natl Cancer Inst. 2015;107:djv219. https://doi.org/10.1093/jnci/djv219.
Thrift AP, Gong J, Peters U, Chang-Claude J, Rudolph A, Slattery ML, et al. Mendelian randomization study of height and risk of colorectal cancer. Int J Epidemiol. 2015;44:662–72. https://doi.org/10.1093/ije/dyv082.
Khankari NK, Shu X-O, Wen W, Kraft P, Lindstrom S, Peters U, et al. Association between adult height and risk of colorectal, lung, and prostate cancer: results from meta-analyses of prospective studies and Mendelian randomization analyses. PLoS Med. 2016;13:e1002118. https://doi.org/10.1371/journal.pmed.1002118.
Thrift AP, Risch HA, Onstad L, Shaheen NJ, Casson AG, Bernstein L, et al. Risk of esophageal adenocarcinoma decreases with height, based on consortium analysis and confirmed by Mendelian randomization. Clin Gastroenterol Hepatol. 2014;12:1667–76e1. https://doi.org/10.1016/j.cgh.201401.039.
Zhang C, Doherty JA, Burgess S, Hung RJ, Lindstrom S, Kraft P, et al. Genetic determinants of telomere length and risk of common cancers: a Mendelian randomization study. Hum Mol Genet. 2015;24:5356–66. https://doi.org/10.1093/hmg/ddv252.
Machiela MJ, Hsiung CA, Shu X-O, Seow WJ, Wang Z, Matsuo K, et al. Genetic variants associated with longer telomere length are associated with increased lung cancer risk among never-smoking women in Asia: a report from the female lung cancer consortium in Asia. Int J Cancer. 2015;137:311–9. https://doi.org/10.1002/ijc.29393.
Rode L, Nordestgaard BG, Bojesen SE. Long telomeres and cancer risk among 95 568 individuals from the general population. Int J Epidemiol. 2016;45:1634–43. https://doi.org/10.1093/ije/dyw179.
Iles MM, Bishop DT, Taylor JC, Hayward NK, Brossard M, Cust AE, et al. The effect on melanoma risk of genes previously associated with telomere length. J Natl Cancer Inst. 2014;106 https://doi.org/10.1093/jnci/dju267.
Machiela MJ, Hofmann JN, Carreras-Torres R, Brown KM, Johansson M, Wang Z, et al. Genetic variants related to longer telomere length are associated with increased risk of renal cell carcinoma. Eur Urol. 2017;72:747–54. https://doi.org/10.1016/j.eururo.2017.07.015.
Walsh KM, Codd V, Rice T, Nelson CP, Smirnov IV, McCoy LS, et al. Longer genotypically-estimated leukocyte telomere length is associated with increased adult glioma risk. Oncotarget. 2015;6:42468–77. https://doi.org/10.18632/oncotarget.6468.
Machiela MJ, Lan Q, Slager SL, Vermeulen RCH, Teras LR, Camp NJ, et al. Genetically predicted longer telomere length is associated with increased risk of B-cell lymphoma subtypes. Hum Mol Genet. 2016;25:1663–76. https://doi.org/10.1093/hmg/ddw027.
Ojha J, Codd V, Nelson CP, Samani NJ, Smirnov IV, Madsen NR, et al. Genetic variation associated with longer telomere length increases risk of chronic lymphocytic leukemia. Cancer Epidemiol Biomark Prev. 2016;25:1043–9. https://doi.org/10.1158/1055-9965.EPI-15-1329.
Walsh KM, Whitehead TP, de Smith AJ, Smirnov IV, Park M, Endicott AA, et al. Common genetic variants associated with telomere length confer risk for neuroblastoma and other childhood cancers. Carcinogenesis. 2016;37:576–82. https://doi.org/10.1093/carcin/bgw037.
Luu HN, Long J, Wen W, Zheng Y, Cai Q, Gao Y-T, et al. Association between genetic risk score for telomere length and risk of breast cancer. Cancer Causes Control. 2016;27:1219–28. https://doi.org/10.1007/s10552-016-0800-z.
• Haycock PC, Burgess S, Nounu A, Zheng J, Okoli GN, Bowden J et al. Association between telomere length and risk of cancer and non-neoplastic diseases: a Mendelian randomization study. JAMA Oncol. 2017. https://doi.org/10.1001/jamaoncol.2016.5945. This meta-analysis provides MR estimates for the effect of telomere length on risk for various cancers and other chronic diseases.
Cheng Y, Yu C, Huang M, Du F, Song C, Ma Z, et al. Genetic association of telomere length with hepatocellular carcinoma risk: a Mendelian randomization analysis. Cancer Epidemiol. 2017;50:39–45. https://doi.org/10.1016/j.canep.2017.07.011.
Du J, Zhu X, Xie C, Dai N, Gu Y, Zhu M, et al. Telomere length, genetic variants and gastric cancer risk in a Chinese population. Carcinogenesis. 2015;36:963–70. https://doi.org/10.1093/carcin/bgv075.
Trompet S, Jukema JW, Katan MB, Blauw GJ, Sattar N, Buckley B, et al. Apolipoprotein E genotype, plasma cholesterol, and cancer: a Mendelian randomization study. Am J Epidemiol. 2009;170:1415–21. https://doi.org/10.1093/aje/kwp294.
Benn M, Tybjærg-Hansen A, Stender S, Frikke-Schmidt R, Nordestgaard BG. Low-density lipoprotein cholesterol and the risk of cancer: a Mendelian randomization study. J Natl Cancer Inst. 2011;103:508–19. https://doi.org/10.1093/jnci/djr008.
Yang C, Tian G, Mi J, Wei X, Li X, Li X, et al. Causal relevance of circulating high-density lipoprotein cholesterol with cancer: a Mendelian randomization meta-analysis. Sci Rep. 2015;5:9495.
He L, Culminskaya I, Loika Y, Arbeev KG, Bagley O, Duan M, et al. Causal effects of cardiovascular risk factors on onset of major age-related diseases: a time-to-event Mendelian randomization study. Exp Gerontol. 2017; https://doi.org/10.1016/j.exger.2017.09.019.
Rodriguez-Broadbent H, Law PJ, Sud A, Palin K, Tuupanen S, Gylfe A, et al. Mendelian randomisation implicates hyperlipidaemia as a risk factor for colorectal cancer. Int J Cancer. 2017;140:2701–8. https://doi.org/10.1002/ijc.30709.
Bull CJ, Bonilla C, Holly JMP, Perks CM, Davies N, Haycock P, et al. Blood lipids and prostate cancer: a Mendelian randomization analysis. Cancer Med. 2016;5:1125–36. https://doi.org/10.1002/cam4.695.
Song M, Gong J, Giovannucci EL, Berndt SI, Brenner H, Chang-Claude J, et al. Genetic variants of adiponectin and risk of colorectal cancer. Int J Cancer. 2015;137:154–64. https://doi.org/10.1002/ijc.29360.
Nimptsch K, Song M, Aleksandrova K, Katsoulis M, Freisling H, Jenab M, et al. Genetic variation in the ADIPOQ gene, adiponectin concentrations and risk of colorectal cancer: a Mendelian randomization analysis using data from three large cohort studies. Eur J Epidemiol. 2017;32:419–30. https://doi.org/10.1007/s10654-017-0262-y.
Pei Y, Xu Y, Niu W. Causal relevance of circulating adiponectin with cancer: a meta-analysis implementing Mendelian randomization. Tumor Biol. 2015;36:585–94. https://doi.org/10.1007/s13277-014-2654-x.
Qu K, Pang Q, Lin T, Zhang L, Gu M, Niu W, et al. Circulating interleukin-10 levels and human papilloma virus and Epstein-Barr virus-associated cancers: evidence from a Mendelian randomization meta-analysis based on 11,170 subjects. OncoTargets Ther. 2016;9:1251–67. https://doi.org/10.2147/OTT.S96772.
Niu W, Pang Q, Lin T, Wang Z, Zhang J, Tai M, et al. A causal role of genetically elevated circulating interleukin-10 in the development of digestive cancers: evidence from Mendelian randomization analysis based on 29,307 subjects. Medicine. 2016;95:e2799. https://doi.org/10.1097/MD.0000000000002799.
Tian G, Mi J, Wei X, Zhao D, Qiao L, Yang C, et al. Circulating interleukin-6 and cancer: a meta-analysis using Mendelian randomization. Sci Rep. 2015;5:11394.
Nimptsch K, Aleksandrova K, Boeing H, Janke J, Lee Y-A, Jenab M, et al. Association of CRP genetic variants with blood concentrations of C-reactive protein and colorectal cancer risk. Int J Cancer. 2015;136:1181–92. https://doi.org/10.1002/ijc.29086.
Allin KH, Nordestgaard BG, Zacho J, Tybjærg-Hansen A, Bojesen SE. C-reactive protein and the risk of cancer: a Mendelian randomization study. J Natl Cancer Inst. 2010;102:202–6. https://doi.org/10.1093/jnci/djp459.
Legason ID, Pfeiffer RM, Udquim K-I, Bergen AW, Gouveia MH, Kirimunda S, et al. Evaluating the causal link between malaria infection and endemic Burkitt lymphoma in Northern Uganda: a Mendelian randomization study. EBioMedicine. 2017;25:58–65. https://doi.org/10.1016/j.ebiom.2017.09.037.
Day FR, Ruth KS, Thompson DJ, Lunetta KL, Pervjakova N, Chasman DI, et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat Genet. 2015;47:1294–303.
• Day FR, Thompson DJ, Helgason H, Chasman DI, Finucane H, Sulem P, et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat gGenet. 2017;49:834–41. https://doi.org/10.1038/ng.3841. This study provides evidence of causal links between timing of puberty and risk for breast, endometrial, and prostate cancers.
Bonilla C, Lewis SJ, Martin RM, Donovan JL, Hamdy FC, Neal DE, et al. Pubertal development and prostate cancer risk: Mendelian randomization study in a population-based cohort. BMC Med. 2016;14:66. https://doi.org/10.1186/s12916-016-0602-x.
Thompson DJ, O'Mara TA, Glubb DM, Painter JN, Cheng T, Folkerd E, et al. CYP19A1 fine-mapping and Mendelian randomization: estradiol is causal for endometrial cancer. Endocr Relat Cancer. 2016;23:77–91. https://doi.org/10.1530/ERC-15-0386.
Tian Q, Jia J, Ling S, Liu Y, Yang S, Shao Z. A causal role for circulating miR-34b in osteosarcoma. Eur J Surg Oncol. 2014;40:67–72. https://doi.org/10.1016/j.ejso.2013.08.024.
Nimptsch K, Aleksandrova K, Boeing H, Janke J, Lee Y-A, Jenab M, et al. Plasma fetuin-A concentration, genetic variation in the AHSG gene and risk of colorectal cancer. Int J Cancer. 2015;137:911–20. https://doi.org/10.1002/ijc.29448.
Kjaergaard AD, Nordestgaard BG, Johansen JS, Bojesen SE. Observational and genetic plasma YKL-40 and cancer in 96,099 individuals from the general population. Int J Cancer. 2015;137:2696–704. https://doi.org/10.1002/ijc.29638.
Bonilla C, Lewis SJ, Rowlands M-A, Gaunt TR, Davey Smith G, Gunnell D, et al. Assessing the role of insulin-like growth factors and binding proteins in prostate cancer using Mendelian randomization: genetic variants as instruments for circulating levels. Int J Cancer. 2016;139:1520–33. https://doi.org/10.1002/ijc.30206.
Lu W-Q, Qiu J-L, Huang Z-L, Liu H-Y. Enhanced circulating transforming growth factor beta 1 is causally associated with an increased risk of hepatocellular carcinoma: a Mendelian randomization meta-analysis. Oncotarget. 2016;7:84695–704. https://doi.org/10.18632/oncotarget.13218.
Xu W, Cheng Y, Zhu H. Evaluation of an association of blood homocysteine levels with Gastric cancer risk from 27 case-control studies. Medicine. 2016;95:e3700. https://doi.org/10.1097/MD.0000000000003700.
Huang Q, Mi J, Wang X, Liu F, Wang D, Yan D, et al. Genetically lowered concentrations of circulating sRAGE might cause an increased risk of cancer: meta-analysis using Mendelian randomization. J Int Med Res. 2016;44:179–91. https://doi.org/10.1177/0300060515617869.
Kobylecki CJ, Afzal S, Nordestgaard BG. Plasma urate, cancer incidence, and all-cause mortality: a Mendelian randomization study. Clin Chem. 2017;63:1151–60. https://doi.org/10.1373/clinchem.2016.268185.
Garland CF, Garland FC, Gorham ED, Lipkin M, Newmark H, Mohr SB, et al. The role of vitamin D in cancer prevention. Am J Public Health. 2006;96(2):252–61. https://doi.org/10.2105/AJPH.2004.045260.
Ma H, Zhou Z, Wei S, Liu Z, Pooley KA, Dunning AM, et al. Shortened telomere length is associated with increased risk of cancer: a meta-analysis. PLoS One. 2011;6:e20466. https://doi.org/10.1371/journal.pone.0020466.
Wentzensen IM, Mirabello L, Pfeiffer RM, Savage SA. The Association of telomere length and cancer: a meta-analysis. Cancer Epidemiol Biomark Prev. 2011;20:1238–50. https://doi.org/10.1158/1055-9965.EPI-11-0005.
Pierce BL, Ahsan H, Vanderweele TJ. Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int J Epidemiol. 2011;40(3):740–52. https://doi.org/10.1093/ije/dyq151.
Freeman G, Cowling BJ, Schooling CM. Power sample size calculations for Mendelian randomization studies using one genetic instrument. Int J Epidemiol. 2013;42(4):1157–63. https://doi.org/10.1093/ije/dyt110.
Brion M-JA, Shakhbazov K, Visscher PM. Calculating statistical power in Mendelian randomization studies. Int J Epidemiol. 2013;42:1497–501. https://doi.org/10.1093/ije/dyt179.
Burgess S, Thompson SG. Use of allele scores as instrumental variables for Mendelian randomization. Int J Epidemiol. 2013;42(4):1134–44. https://doi.org/10.1093/Ije/Dyt093.
Palmer TM, Lawlor DA, Harbord RM, Sheehan NA, Tobias JH, Timpson NJ, et al. Using multiple genetic variants as instrumental variables for modifiable risk factors. Stat Methods Med Res. 2012;21:223–42. https://doi.org/10.1177/0962280210394459.
Burgess S, Davies NM, Thompson SG. Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol. 2016;40(7):597–608. https://doi.org/10.1002/gepi.21998.
• Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol. 2015;44:512–25. https://doi.org/10.1093/ije/dyv080. The method described in this paper, MR-Egger, is one of the several methods available for generating MR estimates that can be robust to violations of the MR assumptions caused by pleiotropic SNPs.
Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016;40(4):304–14. https://doi.org/10.1002/gepi.21965.
Burgess S, Daniel RM, Butterworth AS, Thompson SG, the E-IC. Network Mendelian randomization: using genetic variants as instrumental variables to investigate mediation in causal pathways. Int J Epidemiol. 2014;44:484–95. https://doi.org/10.1093/ije/dyu176.
Burgess S, Thompson SG. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am J Epidemiol. 2015;181(4):251–60. https://doi.org/10.1093/aje/kwu283.
Glymour MM, Tchetgen EJ, Robins JM. Credible Mendelian randomization studies: approaches for evaluating the instrumental variable assumptions. Am J Epidemiol. 2012;175(4):332–9. https://doi.org/10.1093/aje/kwr323.
Gamazon ER, Wheeler HE, Shah KP, Mozaffari SV, Aquino-Michaels K, Carroll RJ, et al. A gene-based association method for mapping traits using reference transcriptome data. Nat Genet. 2015;47(9):1091–8. https://doi.org/10.1038/ng.3367.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48(5):481–7. https://doi.org/10.1038/ng.3538.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48(3):245–52. https://doi.org/10.1038/ng.3506.
Grubert F, Zaugg JB, Kasowski M, Ursu O, Spacek DV, Martin AR, et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell. 2015;162(5):1051–65. https://doi.org/10.1016/j.cell.2015.07.048.
Calderon D, Bhaskar A, Knowles DA, Golan D, Raj T, Fu AQ, et al. Inferring relevant cell types for complex traits by using single-cell gene expression. Am J Hum Genet. 2017;101(5):686–99. https://doi.org/10.1016/j.ajhg.2017.09.009.
Funding
U01HG007601 (BLP), R01 ES020506 (BLP), and P01CA134294 (PK)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Cancer Epidemiology
Rights and permissions
About this article
Cite this article
Pierce, B.L., Kraft, P. & Zhang, C. Mendelian Randomization Studies of Cancer Risk: a Literature Review. Curr Epidemiol Rep 5, 184–196 (2018). https://doi.org/10.1007/s40471-018-0144-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40471-018-0144-1