Abstract
Background
Data on molecular alterations harbored by melanoma brain metastases (MBMs) are limited, and this has hampered the development of more effective therapeutic strategies. We conducted a systematic review and meta-analysis of all the studies reporting DNA sequencing data of MBMs, in order to identify recurrently mutated genes and molecular pathways significantly enriched for genetic alterations.
Methods
We searched PubMed, Embase and Scopus for articles published from the inception of each database to June 30, 2021. We included in the analysis all the studies that reported individual patient data on DNA sequencing of MBMs, assessing single nucleotide variants (SNVs) and/or gene copy number variations (CNVs) in at least five tumor samples. Meta-analysis was performed for genes evaluated for SNVs and/or CNVs in at least two studies. Pooled proportions of samples with SNVs and/or CNVs was calculated by applying random-effect models based on the DerSimonian–Laird method. Gene-set enrichment analysis (GSEA) was performed to identify molecular pathways significantly enriched for mutated genes.
Results
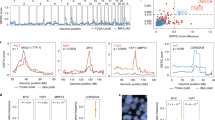
Ten studies fulfilled the inclusion criteria and were included in the analysis, for a total of 531 samples of MBMs evaluated. Twenty-seven genes were found recurrently mutated with a meta-analytic rate of SNVs higher than 5%. GSEA conducted on the list of these 27 recurrently mutated genes revealed vascular endothelial growth factor-activated receptor activity and transmembrane receptor protein tyrosine kinase activity to be among the top 10 gene ontology (GO) molecular functions significantly enriched for mutated genes, while regulation of apoptosis and cell proliferation were among the top 10 significantly enriched GO biological processes. Notably, a high meta-analytic rate of SNVs was found in several actionable cancer-associated genes, such as all the vascular endothelial growth factor (VEGF) receptor isoforms (i.e., Flt1 and Flt2 genes, for both SNV rate: 0.22, 95% CI 0.04–0.49; KDR gene, SNV rate: 0.1, 95% CI 0.05–0.16). Finally, two tumor suppressor genes were characterized by a high meta-analytic rate of CNVs: CDKN2A/B (CNV rate: 0.59, 95% CI 0.23–0.90) and PTEN (CNV rate: 0.31, 95% CI 0.02–0.95).
Conclusion
MBMs harbored actionable molecular alterations that could be exploited as therapeutic targets to improve the poor prognosis of patients.



Similar content being viewed by others
References
Cagney DN, Martin AM, Catalano PJ, et al. Incidence and prognosis of patients with brain metastases at diagnosis of systemic malignancy: a population-based study. Neuro Oncol. 2017;19:1511–21.
Davies MA, Saiag P, Robert C, et al. Dabrafenib plus trametinib in patients with BRAFV600-mutant melanoma brain metastases (COMBI-MB): a multicentre, multicohort, open-label, phase 2 trial. Lancet Oncol. 2017;18:863–73.
Tawbi HA, Forsyth PA, Algazi A, et al. Combined nivolumab and ipilimumab in melanoma metastatic to the Brain. N Engl J Med. 2018;379(8):722–30.
Davies MA, Saiag P, Robert C, et al. Dabrafenib plus trametinib in patients with BRAFV600-mutant melanoma brain metastases (COMBI-MB): a multicentre, multicohort, open-label, phase 2 trial. Lancet Oncol. 2017;18(7):863–73.
Kluger HM, Chiang V, Mahajan A, et al. Long-term survival of patients with melanoma with active brain metastases treated with pembrolizumab on a phase II trial. J Clin Oncol. 2019;37(1):52–60.
Srinivasan ES, Tan AC, Anders CK, et al. Salting the soil: targeting the microenvironment of brain metastases. Mol Cancer Ther. 2021;20(3):455–66.
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372: n71.
Van Allen EM, Miao D, Schilling B, et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science. 2015;350:207–11.
Network CGA. Genomic classification of cutaneous melanoma. Cell. 2015;161(7):1681–96.
Liao Y, Wang J, Jaehnig E, et al. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019;47:W199–205. https://doi.org/10.1093/nar/gkz401.
Dono A, Takayasu T, Yan Y, et al. Differences in genomic alterations between brain metastases and primary tumors. Neurosurgery. 2021;88(3):592–602.
Ferguson SD, Zheng S, Xiu J, et al. Profiles of brain metastases: prioritization of therapeutic targets. Int J Cancer. 2018;143(11):3019–26.
Fischer GM, Jalali A, Kircher DA, et al. Molecular profiling reveals unique immune and metabolic features of melanoma brain metastases. Cancer Discov. 2019;9(5):628–45.
Gamsizkan M, Yilmaz I, Simsek HA, Onguru O, Griffin A, Tihan T. Mutation analysis of metastatic melanomas in the central nervous system: results of a panel of 5 genes in 48 cases. Clin Neuropathol. 2016;35(4):178–85.
In GK, Poorman K, Saul M, et al. Molecular profiling of melanoma brain metastases compared to primary cutaneous melanoma and to extracranial metastases. Oncotarget. 2020;11(33):3118–28.
Chen G, Chakravarti N, Aardalen K, et al. Molecular profiling of patient-matched brain and extracranial melanoma metastases implicates the PI3K pathway as a therapeutic target. Clin Cancer Res. 2014;20(21):5537–46.
Váraljai R, Horn S, Sucker A, et al. Integrative genomic analyses of patient-matched intracranial and extracranial metastases reveal a novel brain-specific landscape of genetic variants in driver genes of malignant melanoma. Cancers. 2021;13(4):731.
Saunus JM, Quinn MC, Patch AM, et al. Integrated genomic and transcriptomic analysis of human brain metastases identifies alterations of potential clinical significance. J Pathol. 2015;237(3):363–78.
Marzese DM, Scolyer RA, Roqué M, et al. DNA methylation and gene deletion analysis of brain metastases in melanoma patients identifies mutually exclusive molecular alterations. Neuro Oncol. 2014;16(11):1499–509.
Rabbie R, Ferguson P, Wong K, et al. The mutational landscape of melanoma brain metastases presenting as the first visceral site of recurrence. Br J Cancer. 2021;124(1):156–60.
Brastianos PK, Carter SL, Santagata S, et al. Genomic characterization of brain metastases reveals branched evolution and potential therapeutic targets. Cancer Discov. 2015;5(11):1164–77.
Ali S, Górska Z, Duchnowska R, Jassem J. Molecular profiles of brain metastases: a focus on heterogeneity. Cancers (Basel). 2021;13(11):2645.
Deng Z, Cui L, Li P, et al. Genomic comparison between cerebrospinal fluid and primary tumor revealed the genetic events associated with brain metastasis in lung adenocarcinoma. Cell Death Dis. 2021;12(10):935.
Morgan AJ, Giannoudis A, Palmieri C. The genomic landscape of breast cancer brain metastases: a systematic review. Lancet Oncol. 2021;22(1):e7–17. https://doi.org/10.1016/S1470-2045(20)30556-8.
Atzori MG, Ceci C, Ruffini F, et al. Role of VEGFR-1 in melanoma acquired resistance to the BRAF inhibitor vemurafenib. J Cell Mol Med. 2020;24:465–75.
Tawbi HA, Boutros C, Kok D, Robert C, McArthur G. New era in the management of melanoma brain metastases. Am Soc Clin Oncol Educ Book. 2018;38:741–50.
Trembath DG, Davis ES, Rao S, et al. Brain tumor microenvironment and angiogenesis in melanoma brain metastases. Front Oncol. 2021;10: 604213.
Hodi FS, Lawrence D, Lezcano C, et al. Bevacizumab plus ipilimumab in patients with metastatic melanoma [published correction appears in Cancer Immunol Res. 2014 Sep;2(9):923]. Cancer Immunol Res. 2014;2(7):632–42.
Fukumura D, Kloepper J, Amoozgar Z, Duda DG, Jain RK. Enhancing cancer immunotherapy using antiangiogenics: opportunities and challenges. Nat Rev Clin Oncol. 2018;15(5):325–40. https://doi.org/10.1038/nrclinonc.2018.29.
Zeng H, Jorapur A, Shain AH, et al. Bi-allelic loss of CDKN2A initiates melanoma invasion via BRN2 activation. Cancer Cell. 2018;34(1):56-68.e9.
Skakodub A, Walch HS, Tringale KR, et al. Genomic analysis and clinical correlations of non-small cell lung cancer (NSCLC) brain metastasis (BM). J Clin Oncol. 2022;40(suppl 16):2008. https://doi.org/10.1200/JCO.2022.40.16_suppl.2008.
Young R, Waldeck K, Martin C, et al. Loss of CDKN2A expression is a frequent event in primary invasive melanoma and correlates with sensitivity to the CDK4/6 inhibitor PD0332991 in melanoma cell lines. Pigment Cell Melanoma Res. 2014;27:590–600.
Goel S, DeCristo M, Watt A, et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature. 2017;548:471–5.
Vidotto T, Melo CM, Castelli E, et al. Emerging role of PTEN loss in evasion of the immune response to tumours. Br J Cancer. 2020;122:1732–43.
Catalanotti F, Cheng DT, Shoushtari AN, et al. PTEN loss-of-function alterations are associated with intrinsic resistance to BRAF inhibitors in metastatic melanoma. JCO Precis Oncol. 2017;1:PO.16.00054.
Niessner H, Schmitz J, Tabatabai G, et al. PI3K pathway inhibition achieves potent antitumor activity in melanoma brain metastases in vitro and in vivo [published correction appears in Clin Cancer Res. 2017 Mar 1;23 (5):1361]. Clin Cancer Res. 2016;22(23):5818–28.
Aasen SN, Parajuli H, Hoang T, et al. Effective treatment of metastatic melanoma by combining MAPK and PI3K signaling pathway inhibitors. Int J Mol Sci. 2019;20(17):4235.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Authors’ contributions
LP, VB, FT, and FC were involved in the conception, design, planning, and management of the study, data acquisition, interpretation of results, drafting the manuscript, and critically reviewing or revising the manuscript for important intellectual content. All other authors supervised the data analysis, provided the interpretation of results, and contributed to the drafting and critical review of the manuscript. All authors approved the final draft. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted. The lead author (LP) is the guarantor.
Competing interests
All authors declared no conflicts of interest.
Funding
There was no funding source for this study. The corresponding author had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Data availability statement
Detailed extracted data on all included studies are available upon reasonable request to the corresponding author.
Code availability
Not applicable.
Consent (participation and publication)
Not applicable.
Ethics approval
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pala, L., Bagnardi, V., Tettamanzi, F. et al. Genetic Alterations of Melanoma Brain Metastases: A Systematic Review and Meta-Analysis. Mol Diagn Ther 27, 5–13 (2023). https://doi.org/10.1007/s40291-022-00623-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-022-00623-0