Abstract
Background and Objectives
Monomethyl auristatin E (MMAE, a cytotoxic agent), upon releasing from valine-citrulline-MMAE (vc-MMAE) antibody-drug conjugates (ADCs), is expected to behave like small molecules. Therefore, evaluating the drug–drug interaction (DDI) potential associated with MMAE is important in the clinical development of ADCs. The objective of this work was to build a physiologically based pharmacokinetic (PBPK) model to assess MMAE–drug interactions for vc-MMAE ADCs.
Methods
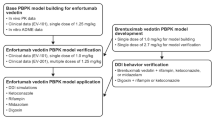
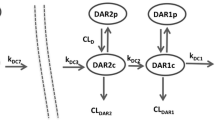
A PBPK model linking antibody-conjugated MMAE (acMMAE) to its catabolite unconjugated MMAE associated with vc-MMAE ADCs was developed using a mixed ‘bottom-up’ and ‘top-down’ approach. The model was developed using in silico and in vitro data and in vivo pharmacokinetic data from anti-CD22-vc-MMAE ADC. Subsequently, the model was validated using clinical pharmacokinetic data from another vc-MMAE ADC, brentuximab vedotin. Finally, the verified model was used to simulate the results of clinical DDI studies between brentuximab vedotin and midazolam, ketoconazole, and rifampicin.
Results
The pharmacokinetic profile of acMMAE and unconjugated MMAE following administration of anti-CD22-vc-MMAE was well described by simulations using the developed PBPK model. The model’s performance in predicting unconjugated MMAE pharmacokinetics was verified by successful simulation of the pharmacokinetic profile following brentuximab vedotin administration. The model simulated DDIs, expressed as area under the concentration-time curve (AUC) and maximum concentration (C max) ratios, were well within the two-fold of the observed data from clinical DDI studies.
Conclusions
This work is the first demonstration of the use of PBPK modelling to predict MMAE-based DDI potential. The described model can be extended to assess the DDI potential of other vc-MMAE ADCs.







Similar content being viewed by others
References
Sasoon I, Blanc V. Antibody-drug conjugate (ADC) clinical pipeline: a review. Methods Mol Biol. 2013;1045:1–27.
Trail PA. Antibody drug conjugates as cancer therapeutics. Antibodies. 2013;2:113–29.
Gorovits BA, Alley SC, Bilic S, Booth B, Kaur S, Oldfield P, Purushothama S, Rao C, Shord S, Siguenza P. Bioanalysis of antibody–drug conjugates: American Association of Pharmaceutical Scientists Antibody–Drug Conjugate Working Group position paper. Bioanalysis. 2013;5:997–1006.
Kaur S, Xu K, Saad OM, Dere RC, Carrasco-Triguera M. Bioanalytical assay strategies for the development of antibody–drug conjugate biotherapeutics. Bioanalysis. 2013;5:201–26.
US FDA, Center for Drug Evaluation and Research. Clinical Pharmacology and Biopharmaceutics Review(s). Application no. 125388 Orig1s000; BLA 125388 and 125399 brentuximab vedotin.
Lu D, Sahasranaman S, Zhang Y, Girish S. Strategies to address drug interaction potential for antibody–drug conjugates in clinical development. Bioanalysis. 2013;5:1115–30.
Han TH, Gopal AK, Ramchandren R, Goy A, Chen R, Matous JV, Cooper M, Grove LE, Alley SC, Lynch CM, O’Connor OA. CYP3A-mediated drug–drug interaction potential and excretion of brentuximab vedotin, an antibody–drug conjugate, in patients with CD30-positive hematologic malignancies. J Clin Pharmacol. 2013;53:866–77.
Houston JB. Utility of in vitro drug metabolism data in predicting in vivo metabolic clearance. Biochem Pharmacol. 1994;47:1469–79.
Obach RS. Prediction of human clearance of twenty-nine drugs from hepatic microsomal intrinsic clearance data: an examination of in vitro half-life approach and nonspecific binding to microsomes. Drug Metab Dispos. 1999;27:1350–9.
Poulin P, Theil F-P. Prediction of pharmacokinetics prior to in vivo studies 1. Mechanistic-based prediction of volume of distribution. J Pharm Sci. 2002;91(1):129–56.
Berezhkovskiy LM. Determination of volume of distribution at steady state with complete consideration of the kinetics of protein and tissue binding in linear pharmacokinetics. J Pharm Sci. 2004;93:364–74.
Rodgers T, Leahy D, Rowland M. Physiologically based pharmacokinetic modeling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci. 2005;95:1238–57.
Lin K, Tibbits J. Pharmacokinetic considerations for antibody drug conjugates. Pharm Res. 2012;92:2354–66.
Li D, Poon KA, Yu SF, Dere R, Go M, Lau J, Zheng B, Elkins K, Danilenko D, Kozak KR, Chan P, Chuh J, Shi X, Nazzal D, Fuh F, McBride J, Ramakrishnan V, de Tute R, Rawstron A, Jack AS, Deng R, Chu YW, Dornan D, Williams M, Ho W, Ebens A, Prabhu S, Polson AG. DCDT2980S, an anti-CD22-monomethyl auristatin E antibody-drug conjugate, is a potential treatment for non-Hodgkin lymphoma. Mol Cancer Ther. 2013;12:1255–65.
Cao Y, Jusko WJ. Applications of minimal physiologically-based pharmacokinetic models. J Pharmacokinet Pharmacodyn. 2012;39:711–23.
European Medicines Agency. Assessment report: Adcetris. EMA/702390/2012. Procedure number EMEA/H/C/002455. 19 Jul 2012. Available from: http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Public_assessment_report/human/002455/WC500135054.pdf. Accessed 12 Sep 2014.
Lu D, Agarwal P, Prabhu S, Nazzal D, Dere R, Saad O, et al. Pharmacokinetics of anti-CD22 and anti-CD79b antibody drug conjugates in relapsed or refractory B-cell non-Hodgkin’s lymphoma patients: results from phase 1 dose escalation studies. American Society for Clinical Pharmacology and Therapeutics Annual Meeting; 5–9 March 2013; Indianapolis, IN.
Oncologic Drugs Advisory Committee. BLA number 125399 (relapsed or refractory systemic anaplastic large cell lymphoma).
Acknowledgments
This study was funded by Genentech (a member of the Roche group). All authors were employees of Genentech when this work was carried out. They have no other conflicts of interest to declare. We would like to thank the DMPK ADME group for generating the in vitro data, Priya Agarwal for processing the CD22 pharmacokinetic data, Amita Joshi, John Prescott, and Yu-Waye Chu for reviewing the manuscript, and Anshin BioSolutions for editorial support.
Author contributions
Yuan Chen, Divya Samineni, Sophie Mukadam, Jin Yan Jin, and Chunze Li participated in the model design; Yuan Chen, Divya Samineni, Sophie Mukadam, Ben-Quan Shen, Dan Lu, and Chunze Li collected data and ran simulations; Yuan Chen, Divya Samineni, Sophie Mukadam, Harvey Wong, and Chunze Li performed the data analysis and wrote the manuscript; and Yuan Chen, Divya Samineni, Sophie Mukadam, Ben-Quan Shen, Harvey Wong, Jin Yan Jin, Dan Lu, Sandhya Girish, Cornelis Hop, and Chunze Li reviewed the manuscript and approved it for submission.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chen, Y., Samineni, D., Mukadam, S. et al. Physiologically Based Pharmacokinetic Modeling as a Tool to Predict Drug Interactions for Antibody-Drug Conjugates. Clin Pharmacokinet 54, 81–93 (2015). https://doi.org/10.1007/s40262-014-0182-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-014-0182-x