Abstract
Background
COVID-19 is an infectious disease caused by SARS-CoV-2, a close relative of SARS-CoV. Several studies have searched for COVID-19 therapies. The topics of these works ranged from vaccine discovery to natural products targeting the SARS-CoV-2 main protease (Mpro), a potential therapeutic target due to its essential role in replication and conserved sequences. However, published research on this target is limited, presenting an opportunity for drug discovery and development.
Method
This study aims to repurpose 10692 drugs in DrugBank by using ligand-based virtual screening (LBVS) machine learning (ML) with Konstanz Information Miner (KNIME) to seek potential therapeutics based on Mpro inhibitors. The top candidate compounds, the native ligand (GC-376) of the Mpro inhibitor, and the positive control boceprevir were then subjected to absorption, distribution, metabolism, excretion, and toxicity (ADMET) characterization, drug-likeness prediction, and molecular docking (MD). Protein–protein interaction (PPI) network analysis was added to provide accurate information about the Mpro regulatory network.
Results
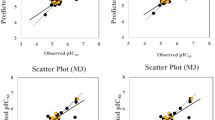
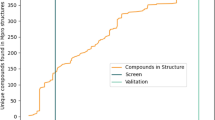
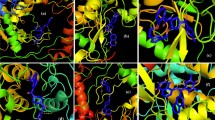
This study identified 3,166 compound candidates inhibiting Mpro. The random forest (RF) molecular access system ML model provided the highest confidence score of 0.95 (bromo-7-nitroindazole) and identified the top 22 candidate compounds. Subjecting the 22 candidate compounds, the native ligand GC-376, and boceprevir to further ADMET property characterization and drug-likeness predictions revealed that one compound had two violations of Lipinski’s rule. Additional MD results showed that only five compounds had more negative binding energies than the native ligand (− 12.25 kcal/mol). Among these compounds, CCX-140 exhibited the lowest score of − 13.64 kcal/mol. Through literature analysis, six compound classes with potential activity for Mpro were discovered. They included benzopyrazole, azole, pyrazolopyrimidine, carboxylic acids and derivatives, benzene and substituted derivatives, and diazine. Four pathologies were also discovered on the basis of the Mpro PPI network.
Conclusion
Results demonstrated the efficiency of LBVS combined with MD. This combined strategy provided positive evidence showing that the top screened drugs, including CCX-140, which had the lowest MD score, can be reasonably advanced to the in vitro phase. This combined method may accelerate the discovery of therapies for novel or orphan diseases from existing drugs.
Graphical abstract






Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
Abbreviations
- ADMET:
-
Absorption, distribution, metabolism, excretion, and toxicity
- ANN:
-
Artificial neural network
- BBB:
-
Blood–brain barrier
- DDX39B:
-
DEAD-box helicase 39B
- COX-2:
-
Cyclooxygenase-2
- hERG:
-
Human ether-a-go-go gene
- LBVS:
-
Ligand-based virtual screening
- MACCS:
-
Molecular access system
- MCS:
-
Maximum common substructure
- MD:
-
Molecular docking
- ML:
-
Machine learning
- MPro :
-
SARS-CoV-2 main protease
- OCT2:
-
Organic cation transporter 2
- PPI:
-
Protein–protein interaction
- RF:
-
Random forest
- ROC:
-
Receiver operating characteristic
- SC-558:
-
1-Phenylsulfonamide-3-trifluoromethyl-5-parabromophenylpyrazole
- SMARTS:
-
SMiles ARbitrary Target Specification
- SMILES:
-
Simplified Molecular-Input Line-Entry System
- SMOTE:
-
Synthetic Minority Over-sampling Technique
- SPTBN2:
-
Nonerythrocytic beta-spectrin 2
- SVM:
-
Support vector machine
References
JHU. COVID-19 Map. In: Johns Hopkins Coronavirus Resource Center. 2022. https://coronavirus.jhu.edu/map.html.
NHS. SARS (severe acute respiratory syndrome). 2022. https://www.nhs.uk/conditions/sars/. Accessed 11 Feb 2022.
Abd El-Aziz TM, Al-Sabi A, Stockand JD. Human recombinant soluble ACE2 (hrsACE2) shows promise for treating severe COVID19. Sig Transduct Target Ther. 2020;5(1):1–2. https://doi.org/10.1038/s41392-020-00374-6.
Clinical Trials. COVID-19 Clinical Trials Drug. 2022.https://clinicaltrials.gov/ct2/results?cond=2019nCoV&Search=Clear&age_v=&gndr=&type=&rslt=. Accessed 23 Jan 2022.
Clinical Trials Arena. Coronavirus treatment: Vaccines/drugs in the pipeline for COVID-19. 2022. https://www.clinicaltrialsarena.com/analysis/coronavirus-mers-cov-drugs/. Accessed 21 Jan 2022.
Gautret P, Lagier JC, Parola P, Hoang VT, Medded L, Mailhe M, et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: preliminary results of an open-label non-randomized clinical trial. 2020.
RAPS. COVID-19 vaccine tracker. 2022. https://www.raps.org/news-and-articles/news-articles/2020/3/covid-19-vaccine-tracker. Accessed 23 Jan 2022.
Akinlalu AO, Chamundi A, Yakumbur DT, Afolayan FID, Duru IA, Arowosegbe MA, et al. Repurposing FDA-approved drugs against multiple proteins of SARS-CoV-2: an in silico study. Sci Afr. 2021;13:e00845. https://doi.org/10.1016/j.sciaf.2021.e00845.
Hoertel N, Sánchez-Rico M, Cougoule C, Gulbins E, Kornhuber J, Carpinteiro A, et al. Repurposing antidepressants inhibiting the sphingomyelinase acid/ceramide system against COVID-19: current evidence and potential mechanisms. Mol Psychiatry. 2021;26(12):7098–9. https://doi.org/10.1038/s41380-021-01254-3.
Kandeel M, Abdelrahman AHM, Oh-Hashi K, Ibrahim A, Venugopala KN, Morsy MA, et al. Repurposing of FDA-approved antivirals, antibiotics, anthelmintics, antioxidants, and cell protectives against SARS-CoV-2 papain-like protease. J Biomol Struct Dyn. 2021;39(14):5129–36. https://doi.org/10.1080/07391102.2020.1784291.
Singh AK, Singh A, Dubey AK. Repurposed therapeutic strategies towards COVID-19 potential targets based on genomics and protein structure remodeling. IntechOpen. 2021.
Qamar MT, Alqahtani SM, Alamri MA, Chen L-L. Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J Pharm Anal. 2020;10(4):313–9. https://doi.org/10.1016/j.jpha.2020.03.009.
Lu J, Chen SA, Khan MB, Brassard R, Arutyunova E, Lamer T, et al. Crystallization of feline coronavirus Mpro with GC376 reveals mechanism of inhibition. Front Chem. 2022:10.
Mahase E. Covid-19: Pfizer’s paxlovid is 89% effective in patients at risk of serious illness, company reports. BMJ. 2021;n2713. https://doi.org/10.1136/bmj.n2713.
Mengist HM, Mekonnen D, Mohammed A, Shi R, Jin T. Potency, safety, and pharmacokinetic profiles of potential inhibitors targeting SARS-CoV-2 main protease. Front Pharmacol. 2021;11:630500. https://doi.org/10.3389/fphar.2020.630500.
Mody V, Ho J, Wills S, Mawri A, Lawson L, Ebert MCCJC, et al. Identification of 3-chymotrypsin like protease (3CLPro) inhibitors as potential anti-SARS-CoV-2 agents. Commun Biol. 2021;4(1):1–10. https://doi.org/10.1038/s42003-020-01577-x.
Sharun K, Tiwari R, Dhama K. Protease inhibitor GC376 for COVID-19: lessons learned from feline infectious peritonitis. Ann Med Surg (Lond). 2020;61:122–5. https://doi.org/10.1016/j.amsu.2020.12.030.
Andreani J, Le Bideau M, Duflot I, Jardot P, Rolland C, Boxberger M, et al. In vitro testing of combined hydroxychloroquine and azithromycin on SARS-CoV-2 shows synergistic effect. Microb Pathog. 2020;145:104228.
Choudhary R, Sharma AK. Potential use of hydroxychloroquine, ivermectin and azithromycin drugs in fighting COVID-19: trends, scope and relevance. New Microbes New Infect. 2020;35:100684.
Chu CM, Cheng VCC, Hung IFN, Wong MML, Chan KH, Chan KS, et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59(3):252–6. https://doi.org/10.1136/thorax.2003.012658.
Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30(3):269–71. https://doi.org/10.1038/s41422-020-0282-0.
Javelot H, El-Hage W, Meyer G, Becker G, Michel B, Hingray C. COVID-19 and (hydroxy) chloroquine–azithromycin combination: should we take the risk for our patients? Br J Clin Pharmacol. 2020;86(6):1176.
Gadaleta D, Lombardo A, Toma C, Benfenati E. A new semi-automated workflow for chemical data retrieval and quality checking for modeling applications. J Cheminformatics. 2018;10(1):60. https://doi.org/10.1186/s13321-018-0315-6.
DCIS. Daylight Theory: SMARTS - A Language for Describing Molecular Patterns. 2022. https://www.daylight.com/dayhtml/doc/theory/theory.smarts.html. Accessed 18 Apr 2022.
Bajorath J. Machine learning and similarity-based virtual screening techniques. In: Silico drug discovery and design. Unitec house, 2 Albert place. London: Future Science Ltd; 2013. p. 134–46.
Maggiora G, Vogt M, Stumpfe D, Bajorath J. Molecular similarity in medicinal chemistry. J Med Chem. 2014;57(8):3186–204. https://doi.org/10.1021/jm401411z.
Massagué AC, Ojeda MJ, Valls C, Mulero M, Garcia-Vallvé S, Pujadas G. Molecular fingerprint similarity search in virtual screening. Methods. 2015;71:58–63. https://doi.org/10.1016/j.ymeth.2014.08.005.
Mysinger MM, Carchia M, Irwin JJ, Shoichet BK. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J Med Chem. 2012;55(14):6582–94. https://doi.org/10.1021/jm300687e.
Cichonska A, Ravikumar B, Allaway RJ, Park S, Wan F, Isayev O, et al. Crowdsourced mapping extends the target space of kinase inhibitors. bioRxiv; 2020. p. https://doi.org/10.1101/2019.12.31.891812.
PDB. Resolution - Proteopedia, life in 3D. 2022. https://proteopedia.org/wiki/index.php/Resolution. Accessed 20 Apr 2022.
Hu Y, Stumpfe D, Bajorath J. Computational exploration of molecular scaffolds in medicinal chemistry. J Med Chem. 2016;59(9):4062–76. https://doi.org/10.1021/acs.jmedchem.5b01746.
Yang Y. Chapter 3 - temporal data clustering. In: Yang Y, editor. Temporal data mining via unsupervised ensemble learning. Elsevier; 2017. p. 19–34.
Chaudhaery SS, Roy KK, Saxena AK. Consensus superiority of the pharmacophore-based alignment, over maximum common substructure (MCS): 3D-QSAR studies on carbamates as acetylcholinesterase inhibitors. J Chem Inf Model. 2009;49(6):1590–601. https://doi.org/10.1021/ci900049e.
Nighania K. Various ways to evaluate a machine learning models performance. Medium. 2019.
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717. https://doi.org/10.1038/srep42717.
Kumar A, Kumari K, Singh S, Bahadur I, Singh P. Noscapine anticancer drug designed with ionic liquids to enhance solubility: DFT and ADME approach. J Mol Liq. 2021;325:115159. https://doi.org/10.1016/j.molliq.2020.115159.
Pires DEV, Blundell TL, Ascher DB. Pkcsm: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem. 2015;58(9):4066–72. https://doi.org/10.1021/acs.jmedchem.5b00104.
Hermawan A, Wulandari F, Hanif N, Utomo RY, Jenie RI, Ikawati M, et al. Identification of potential targets of the curcumin analog CCA-1.1 for glioblastoma treatment : integrated computational analysis and in vitro study. Sci Rep. 2022;12(1):13928. https://doi.org/10.1038/s41598-022-18348-9.
Fu L, Ye F, Feng Y, Yu F, Wang Q, Wu Y, et al. Both Boceprevir and GC376 efficaciously inhibit SARS-CoV-2 by targeting its main protease. Nat Commun. 2020;11(1) https://doi.org/10.1038/s41467-020-18233-x.
Oerlemans R, Ruiz-Moreno AJ, Cong Y, Kumar ND, Velasco-Velazquez MA, Neochoritis CG, et al. Repurposing the HCV NS3–4A protease drug boceprevir as COVID-19 therapeutics. RSC Med Chem. 2021;12(3):370–9. https://doi.org/10.1039/D0MD00367K.
Oughtred R, Rust J, Chang C, Breitkreutz B-J, Stark C, Willems A, et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 2021;30(1):187–200. https://doi.org/10.1002/pro.3978.
Chin C-H, Chen S-H, Wu H-H, Ho C-W, Ko M-T, Lin C-Y. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8(4):S11. https://doi.org/10.1186/1752-0509-8-S4-S11.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504. https://doi.org/10.1101/gr.1239303.
Brownlee J. How to know if your machine learning model has good performance. Machine Learning Mastery. 2018.
El Khouli RH, Macura KJ, Barker PB, Phil D, Habba MR, Jacobs MA, et al. The relationship of temporal resolution to diagnostic performance for dynamic contrast enhanced (DCE) MRI of the breast. J Magn Reson Imaging. 2009;30(5):999–1004. https://doi.org/10.1002/jmri.21947.
McHugh ML. Interrater reliability: the kappa statistic. Biochem Med (Zagreb). 2012;22(3):276–82.
Azzahra SNA, Hanif N, Hermawan A. MDM2 is a potential target gene of glycyrrhizic acid for circumventing breast cancer resistance to tamoxifen: integrative bioinformatics analysis. Asian Pac J Cancer Prev. 2022;23(7):2341–50. https://doi.org/10.31557/APJCP.2022.23.7.2341.
Gill AJ. Succinate dehydrogenase (SDH)-deficient neoplasia. Histopathology. 2018;72(1):106–16. https://doi.org/10.1111/his.13277.
Szymura SJ, Bernal GM, Wu L, Zhang Z, Crawley CD, Voce DJ, et al. DDX39B interacts with the pattern recognition receptor pathway to inhibit NF-κB and sensitize to alkylating chemotherapy. BMC Biol. 2020;18(1):32. https://doi.org/10.1186/s12915-020-0764-z.
Chan JF-W, Kok K-H, Zhu Z, Chu H, To KK-W, Yuan S, et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020;9(1):221–36. https://doi.org/10.1080/22221751.2020.1719902.
Belotserkovskaya R, Oh S, Bondarenko VA, Orphanides G, Studitsky VM, Reinberg D. FACT facilitates transcription-dependent nucleosome alteration. Science. 2003;301(5636):1090–3. https://doi.org/10.1126/science.1085703.
Cui J, Wang L, Ren X, Zhang Y, Zhang H. LRPPRC: a multifunctional protein involved in energy metabolism and human disease. Front Physiol. 2019;10:595. https://doi.org/10.3389/fphys.2019.00595.
Zhang H-R, Lai S-Y, Huang L-J, Zhang Z-F, Liu J, Zheng S-R, et al. Myosin 1b promotes cell proliferation, migration, and invasion in cervical cancer. Gynecol Oncol. 2018:149. https://doi.org/10.1016/j.ygyno.2018.01.024.
Ngo HB, Lovely GA, Phillips R, Chan DC. Distinct structural features of TFAM drive mitochondrial DNA packaging versus transcriptional activation. Nat Commun. 2014;5(1):3077. https://doi.org/10.1038/ncomms4077.
Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman A. The integrated stress response. EMBO Rep. 2016;17(10):1374–95. https://doi.org/10.15252/embr.201642195.
Amorim IS, Lach G, Gkogkas CG. The role of the eukaryotic translation initiation factor 4E (eIF4E) in neuropsychiatric disorders. Front Genet. 2018;9:561. https://doi.org/10.3389/fgene.2018.00561.
Romaniello R, Citterio A, Panzeri E, Arrigoni F, De Rinaldis M, Trabacca A, et al. Novel SPTBN2 gene mutation and first intragenic deletion in early onset spinocerebellar ataxia type 5. Ann Clin Transl Neurol. 2021;8(4):956–63. https://doi.org/10.1002/acn3.51345.
Becker JH, Lin JJ, Doernberg M, Stone K, Navis A, Festa JR, et al. Assessment of cognitive function in patients after COVID-19 infection. JAMA Netw Open. 2021;4(10):e2130645. https://doi.org/10.1001/jamanetworkopen.2021.30645.
Bora VR, Patel BM. The deadly duo of COVID-19 and cancer! Front Mol Biosci. 2021;8:643004. https://doi.org/10.3389/fmolb.2021.643004.
Hojyo S, Uchida M, Tanaka K, Hasebe R, Tanaka Y, Murakami M, et al. How COVID-19 induces cytokine storm with high mortality. Inflamm Regener. 2020;40(1):37. https://doi.org/10.1186/s41232-020-00146-3.
Liu Y-H, Chen Y, Wang Q-H, Wang L-R, Jiang L, Yang Y, et al. One-year trajectory of cognitive changes in older survivors of COVID-19 in Wuhan, China: a longitudinal cohort study. JAMA Neurology. 2022; https://doi.org/10.1001/jamaneurol.2022.0461.
Merad M, Martin JC. Pathological inflammation in patients with COVID-19: a key role for monocytes and macrophages. Nat Rev Immunol. 2020;20(6):355–62. https://doi.org/10.1038/s41577-020-0331-4.
Povlow A, Auerbach AJ. Acute cerebellar ataxia in COVID-19 infection: a case report. J Emerg Med. 2021;60(1):73–6. https://doi.org/10.1016/j.jemermed.2020.10.010.
Saini G, Aneja R. Cancer as a prospective sequela of long COVID-19. Bioessays. 2021;43(6):e2000331. https://doi.org/10.1002/bies.202000331.
Sullivan T, Miao Z, Dairaghi DJ, Krasinski A, Wang Y, Zhao BN, et al. CCR2 antagonist CCX140-B provides renal and glycemic benefits in diabetic transgenic human CCR2 knockin mice. Am J Physiol Renal Physiol. 2013;305(9):F1288–F97. https://doi.org/10.1152/ajprenal.00316.2013.
Perez-Gomez MV, Sanchez-Niño MD, Sanz AB, Martín-Cleary C, Ruiz-Ortega M, Egido J, et al. Horizon 2020 in diabetic kidney disease: the clinical trial pipeline for add-on therapies on top of renin angiotensin system blockade. J Clin Med. 2015;4(6):1325–47. https://doi.org/10.3390/jcm4061325.
Sullivan TJ, Miao Z, Zhao BN, Ertl LS, Wang Y, Krasinski A, et al. Experimental evidence for the use of CCR2 antagonists in the treatment of type 2 diabetes. Metabolism. 2013;62(11):1623–32. https://doi.org/10.1016/j.metabol.2013.06.008.
Stipp MC, Acco A. Involvement of cytochrome P450 enzymes in inflammation and cancer: a review. Cancer Chemother Pharmacol. 2021;87(3):295–309. https://doi.org/10.1007/s00280-020-04181-2.
Zhao M, Ma J, Li M, Zhang Y, Jiang B, Zhao X, et al. Cytochrome P450 enzymes and drug metabolism in humans. Int J Mol Sci. 2021;22(23):12808. https://doi.org/10.3390/ijms222312808.
Tao J, Aristotelidis R, Zanowick-Marr A, Chambers LC, McDonald J, Mylonakis EE, et al. Evaluation of the treatment efficacy and safety of remdesivir for COVID-19: a meta-analysis. SN Compr Clin Med. 2021;3(12):2443–54. https://doi.org/10.1007/s42399-021-01014-y.
Aleissa MM, Silverman EA, Paredes Acosta LM, Nutt CT, Richterman A, Marty FM. New perspectives on antimicrobial agents: remdesivir treatment for COVID-19. Antimicrob Agents Chemother. 2020;65(1). https://doi.org/10.1128/AAC.01814-20.
Wang G, Xiao B, Deng J, Gong L, Li Y, Li J, et al. The role of cytochrome P450 enzymes in COVID-19 pathogenesis and therapy. Front Pharmacol. 2022;13:791922. https://doi.org/10.3389/fphar.2022.791922.
Stergiopoulos C, Tsopelas F, Valko K. Prediction of hERG inhibition of drug discovery compounds using biomimetic HPLC measurements. ADMET DMPK. 2021;9(3):191–207. https://doi.org/10.5599/admet.995.
Munawar S, Windley MJ, Tse EG, Todd MH, Hill AP, Vandenberg JI, et al. Experimentally validated pharmacoinformatics approach to predict herg inhibition potential of new chemical entities. Front Pharmacol. 2018:9. https://doi.org/10.3389/fphar.2018.01035.
Kumar S, Singh B, Kumari P, Kumar PV, Agnihotri G, Khan S, et al. Identification of multipotent drugs for COVID-19 therapeutics with the evaluation of their SARS-CoV2 inhibitory activity. Comput Struct Biotechnol J. 2021;19:1998–2017. https://doi.org/10.1016/j.csbj.2021.04.014.
Theodoridou A, Gika H, Diza E, Garyfallos A, Settas L. In vivo study of pro-inflammatory cytokine changes in serum and synovial fluid during treatment with celecoxib and etoricoxib and correlation with VAS pain change and synovial membrane penetration index in patients with inflammatory arthritis. MJR. 2017;28(1):33–40. https://doi.org/10.31138/mjr.28.1.33.
Prasher P, Sharma M, Gunupuru R. Targeting cyclooxygenase enzyme for the adjuvant COVID-19 therapy. Drug Dev Res. 2021; https://doi.org/10.1002/ddr.21794.
Ke Y-Y, Peng T-T, Yeh T-K, Huang W-Z, Chang S-E, Wu S-H, et al. Artificial intelligence approach fighting COVID-19 with repurposing drugs. Biom J. 2020;43(4):355–62. https://doi.org/10.1016/j.bj.2020.05.001.
Gimeno A, Mestres-Truyol J, Ojeda-Montes MJ, Macip G, Saldivar-Espinoza B, Cereto-Massagué A, et al. Prediction of novel inhibitors of the main protease (M-pro) of SARS-CoV-2 through consensus docking and drug reposition. Int J Mol Sci. 2020;21(11):E3793. https://doi.org/10.3390/ijms21113793.
Magro P, Zanella I, Pescarolo M, Castelli F, Quiros-Roldan E. Lopinavir/ritonavir: repurposing an old drug for HIV infection in COVID-19 treatment. Biom J. 2021;44(1):43–53. https://doi.org/10.1016/j.bj.2020.11.005.
Cvetkovic RS, Goa KL. Lopinavir/ritonavir. Drugs. 2003;63(8):769–802. https://doi.org/10.2165/00003495-200363080-00004.
Croxtall JD, Perry CM. Lopinavir/ritonavir. Drugs. 2010;70(14):1885–915. https://doi.org/10.2165/11204950-000000000-00000.
Marzolini C, Stader F, Stoeckle M, Franzeck F, Egli A, Bassetti S, et al. Effect of systemic inflammatory response to SARS-CoV-2 on lopinavir and hydroxychloroquine plasma concentrations. Antimicrob Agents Chemother. 2020;64(9). https://doi.org/10.1128/AAC.01177-20.
Ma C, Sacco MD, Hurst B, Townsend JA, Hu Y, Szeto T, et al. Boceprevir, GC-376, and calpain inhibitors II, XII inhibit SARS-CoV-2 viral replication by targeting the viral main protease. Cell Res. 2020;30(8):678–92. https://doi.org/10.1038/s41422-020-0356-z.
Cáceres CJ, Cardenas-Garcia S, Carnaccini S, Seibert B, Rajao DS, Wang J, et al. Efficacy of GC-376 against SARS-CoV-2 virus infection in the K18 hACE2 transgenic mouse model. Sci Rep. 2021;11(1):9609. https://doi.org/10.1038/s41598-021-89013-w.
Rizza SA, Talwani R, Nehra V, Temesgen Z. Boceprevir. Drugs of Today. 2011;47(10):743. https://doi.org/10.1358/dot.2011.47.10.1656503.
Kim Y, Lovell S, Tiew K-C, Mandadapu SR, Alliston KR, Battaile KP, et al. Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses. J Virol. 2012;86(21):11754–62. https://doi.org/10.1128/JVI.01348-12.
Didziapetris R, Japertas P, Avdeef A, Petrauskas A. Classification analysis of P-glycoprotein substrate specificity. J Drug Target. 2003;11(7):391–406. https://doi.org/10.1080/10611860310001648248.
Mandò C, Savasi VM, Anelli GM, Corti S, Serati A, Lisso F, et al. Mitochondrial and oxidative unbalance in placentas from mothers with SARS-CoV-2 infection. Antioxidants. 2021;10(10):1517. https://doi.org/10.3390/antiox10101517.
Refolo G, Ciccosanti F, Di Rienzo M, Basulto Perdomo A, Romani M, Alonzi T, et al. Negative regulation of mitochondrial antiviral signaling protein-mediated antiviral signaling by the mitochondrial protein LRPPRC during hepatitis C virus infection. Hepatology. 2019;69(1):34–50. https://doi.org/10.1002/hep.30149.
Dong A, Zhao J, Griffin C, Wu R. The genomic physics of COVID-19 pathogenesis and spread. Cells. 2022;11(1):80. https://doi.org/10.3390/cells11010080.
McCarthy MK, Weinberg JB. The immunoproteasome and viral infection: a complex regulator of inflammation. Front Microbiol. 2015;6(1):630500. https://doi.org/10.3389/fphar.2020.630500.
West AP, Khoury-Hanold W, Staron M, Tal MC, Pineda CM, Lang SM, et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature. 2015;520(7548):553–7. https://doi.org/10.1038/nature14156.
Sooryanarain H, Rogers AJ, Cao D, Haac MER, Karpe YA, Meng X-J. ISG15 modulates type I interferon signaling and the antiviral response during hepatitis E virus replication. J Virol. 2017;91(19):e00621–17. https://doi.org/10.1128/JVI.00621-17.
Zhou D, Park J-G, Wu Z, Huang H, Fiches GN, Biswas A, et al. FACT subunit SUPT16H associates with BRD4 and contributes to silencing of antiviral interferon signaling. Mol Biol. 2021.
Wenzel J, Lampe J, Müller-Fielitz H, Schuster R, Zille M, Müller K, et al. The SARS-CoV-2 main protease Mpro causes microvascular brain pathology by cleaving NEMO in brain endothelial cells. Nat Neurosci. 2021;24(11):1522–33. https://doi.org/10.1038/s41593-021-00926-1.
Acknowledgements
The authors thank Badan Penerbit dan Publikasi, Universitas Gadjah Mada for their assistance in writing.
Funding
The authors did not receive any particular grant from the public, commercial, or non-profit funding agency.
Author information
Authors and Affiliations
Contributions
GPWCY contributed to the design, acquisition, the writing and revision of the article, drafted the article, and the finalized the version to be published. NH contributed to the data acquisition, writing and revision of the article. AH contributed to supervision, review and evaluation of the data and the final approval of the version to be published. This manuscript is a part of the bachelor thesis of GPWCY.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This work does not involve any studies with human participants or animals.
Competing interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yuda, G.P.W.C., Hanif, N. & Hermawan, A. Computational Screening Using a Combination of Ligand-Based Machine Learning and Molecular Docking Methods for the Repurposing of Antivirals Targeting the SARS-CoV-2 Main Protease. DARU J Pharm Sci 32, 47–65 (2024). https://doi.org/10.1007/s40199-023-00484-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40199-023-00484-w