Abstract
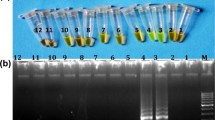
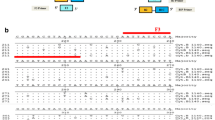
The aim of this study was to develop PCR assay for simultaneous detection of tissues from cattle and buffalo in a single PCR reaction. Species-specific primers for cattle and buffalo species were designed through homology comparisons and alignment of partial sequences of Cyt.b gene from common food animals. The conditions for species-specific PCR amplification were optimized in terms of quantity and concentration of various components for PCR mix and annealing temperature for each set of designed species-specific primer pairs. The assay revealed the amplification of genomic DNA from cattle to amplicon size of 655 bp, whereas DNA template from buffalo to amplicon length of 249 bp. Simultaneous amplification of DNA templates from cattle and buffalo was observed in duplex PCR assay. Thus, the developed assay could identify the tissues of cattle and buffalo origin, detecting even both the species together and being advantageous over species-specific PCR.



Similar content being viewed by others
References
Zhang C (2013) Semi-nested multiplex PCR enhanced method sensitivity of species detection in further-processed meats. Food Control 31:326–330
Amaral JS, Santos CG, Melo VS, Costa J, Oliveira MBPP, Mafra I (2015) Identification of duck, partridge, pheasant, quail, chicken and turkey meats by species-specific PCR assays to assess the authenticity of traditional game meat sausages. Food Control 47:190–195
Cheng X, He W, Huang F, Huang M, Zhou G (2014) Multiplex real-time PCR for the identification and quantification of DNA from duck, pig and chicken in Chinese blood curds. Food Res Int 60:30–37
Kallea E, Kubista M, Rensing C (2014) Multi-template polymerase chain reaction. Biomol Detect Quantif 2:11–29
Park YH, Uzzaman MR, Park JW, Kim SW, Lee JH, Kim KS (2013) Detection of meat origin (species) using polymerase chain reaction. Korean J Food Sci Anim Resour 33(6):696–700
Ali ME, Razzak MA, Hamid SBA, Rahman MM, Amin MA, Rashid NRA (2015) Multiplex PCR assay for the detection of five meat species forbidden in Islamic foods. Food Chem 177:214–224
Guan F, Jin YT, Zhao J, Xu AC, Luo YY (2018) A PCR method that can be further developed into PCR-RFLP assay for eight animal species identification. J Anal Methods Chem. https://doi.org/10.1155/2018/5890140
Abbas O, Zadravec M, Baeten Mikus VT, Lesic T, Vulic A, Pleadin J (2018) Analytical methods used for the authentication of food of animal origin. Food Chem 246:6–17
Hanapi UK, Desa MN, Ismail A, Mustafa S (2015) A higher sensitivity and efficiency of common primer multiplex PCR assay in identification of meat origin using NADH dehydrogenase subunit 4 gene. J Food Sci Technol 52(7):4166–4175
Kumari R, Rank DN, Kumar S, Joshi CG, Lal SV (2015) Real time PCR- an approach to detect meat adulteration. Buffalo Bull 34(1):124–129
You J, Huang L, Zhuang J, Mou Z (2014) Species specific multiplex real-time PCR assay for identification of deer and common domestic animals. Food Sci Biotechnol 23:133–139
Panwar N, Gahlot GC, Gahlot K, Ashraf M, Singh A (2015) Rapid identification of goat (Capra hircus) and sheep (Ovis aries) species in raw meat using duplex PCR assay. Indian J Anim Res 49:4–10
Vaithiyanathan S, Kulkarni VV (2016) Species identification of cattle and buffalo fat through PCR assay. J Food Sci Technol 53(4):2077–2082
Gould MP, Bosworth CM, McMahon S, Grandhi S, Grimberg BT et al (2016) PCR-free enrichment of mitochondrial DNA from human blood and cell lines for high quality next-generation DNA sequencing. PLoS ONE 11(5):e0156884
Ruihua Z, Ping J, Chuanbo S, Deyong S, Feng Z, Chaochao H (2018) The analysis of genetic variation in the mitochondrial genome and its application for the identification of Papilio species. Mitochondrial DNA 3(2):687–690
Langdon QK, Peris D, Kyle B, Hittinger CT (2018) sppIDer: a species identification tool to investigate hybrid genomes with high-throughput sequencing. Mol Biol Evol 35(11):2835–2849
Adiningsiha MW, Soejoedonoc RD, Purnawarmanc T, Latifc H, Poetric ON, Putrid DD (2018) Authentication of sumateran wild boar (sus scrofa vittatus) meat contamination by polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP) technique of cytochrome b gene. Trop Anim Sci J 41(3):157
Makwana MD, Rathod PT, Roy SK, Patel BK (2016) Detection of different meat species by multiplex PCR assay. Int J Agric Sci 8(52):2595–2597
Ewart KM, Frankham GJ, McEwing R, The DT, Hogg CJ et al (2018) A rapid multiplex PCR assay for presumptive species identification of rhinoceros horns and its implementation in Vietnam. PLoS ONE 13(8):e0202433
Zarei M, Maktabi S, Nasiri M (2016) Fraud identification of cow’s milk in buffalo’s milk and it’s products using the polymerase chain reaction. Jundishapur J Health Sci 8(4):e60326
Acknowledgements
The authors gratefully acknowledge Indian Council of Agricultural Research, Indian Veterinary Research Institute (IVRI), Izatnagar, India (Grant No. LPT-2014-2018-II), for providing necessary facilities and financial support to accomplish this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest to publish this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Significance Statement The high market value of meat often leads to targets for species substitution of meat. Detection of species-specific DNA has been basis for the most of the advanced analytical methods utilized to date for meat authentication. The co-amplification of separate regions of a single gene or specific fragments, each typical of a different animal species in a single PCR reaction, using different pairs of primers in the same reaction mix is the unique advantage with duplex PCR.
Rights and permissions
About this article
Cite this article
Kumbhar, V.H., Kumar, R.R., Mendiratta, S.K. et al. Duplex PCR Assay for Identification of Tissue of Cattle and Buffalo Origin Targeting Mitochondrial Cytochrome b Gene. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 90, 287–291 (2020). https://doi.org/10.1007/s40011-019-01102-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-019-01102-z