Abstract
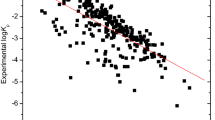
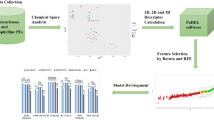
The prediction of maximum steady state flux values of a chemical compound from its structural features plays an important role in design of transdermal drug delivery systems. In this study, we developed the quantitative structure property relationship (QSPR) models to estimate the maximum steady state flux of 245 drugs-like compounds through the polydimethylsiloxane membranes. A correlation-based feature selection was used for descriptor selection. The selected descriptors, surface tension, polarity, and count of hydrogen accept sites, which are interpretable and can be used to explain the permeability of chemicals. These descriptors are used for developing the QSPR prediction models by multiple linear regression, artificial neural network, support vector machine (SVM) and Instance-Based Learning algorithms using K nearest neighbor machine learning approaches. The models were assessed by internal and external validation. All four approaches yield the QSPR models with good statistics. The models developed by SVM have better prediction performance. These models can be useful for predicting the permeability new untested compounds.





Similar content being viewed by others
References
ACD/ChemSketch11, Advanced Chemistry Development, Inc, 8 King Street East, Suite 107, Toronto, M5C 1B5 Canada. http://www.acdlabs.com
Aha D, Kibler DW, Albert MK (1991) Instance-based learning algorithms. Mach Learn 6:37–66
Burges CA (1998) Tutorial on support vector machines for pattern recognition. Data Min Knowl Discov 2:1–43
Chen Y, Yang WL, Matheson LE (1993) Prediction of flux through polydimethylsiloxane membranes using atomic charge calculations. Int J Pharm 94:81–88
Chen Y, Vayuhasuwan P, Matheson LE (1996) Prediction of flux through polydimethylsiloxane membranes using atomic charge calculations: application to an extended data set. Int J Pharm 137:149–158
Cronin MTD, Dearden JC, Gupta R, Moss GP (1998) An investigation of the mechanism of flux across polydimethylsiloxane membranes by quantitative structure–permeability relationships. J Pharm Pharmacol 50:143–152
Doucet JP, Barbault F, Xia H, Panaye A, Fan B (2007) Nonlinear SVM approaches to QSPR/QSAR studies and drug design. Curr Comput Aided Drug Des 3:263–389
Draper NR, Smith H (1981) Applied regression analysis. Wiley, New York
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701
Gramatica P, Giani E, Papa E (2007) Statistical external validation and consensus modeling: a QSPR case study for Koc prediction. J Mol Graph Model 25:755–766
Guha R, Dutta D, Jurs PC, Chen T (2006) Local lazy regression: making use of the neighborhood to improve qsar predictions. J Chem Inf Model 46:1836–1847
Hall MA, Holmes G (2003) Benchmarking attribute selection techniques for discrete class data mining. IEEE Trans Knowl Data Eng 15:1437–1447
Hasegawa K, Funatsu K (2010) Non-linear modeling and chemical interpretation with aid of support vector machine and regression. Curr Comput Aided Drug Des 6:24–36
Haykin S (2006) Neural networks. A comprehensive foundation, 2nd edn. Perarson Prentice Hall, New Delhi
Heikamp K, Bajorath J (2014) Support vector machines for drug discovery. Expert Opin Drug Discov 9:93–104
Konovalov DA, Llewellyn LE, Heyden YV, Coomans D (2008) Robust cross-validation of linear regression QSAR models. J Chem Inf Model 48:2081–2094
Lavecchia A (2015) Machine learning approaches in drug discovery: methods and applications. Drug Discov Today 20:318–331
Louis B, Agrawal VK, Khadikar PV (2010) Prediction of intrinsic solubility of generic drugs using MLR, ANN and SVM analyses. Eur J Med Chem 45:4018–4025
Maa W, Luana F, Zhaoa C, Zhanga X, Liua M, Hua Z, Fanb B (2006) QSAR prediction of the penetration of drugs across a polydimethylsiloxane membrane. QSAR Comb Sci 25:895–904
Mitchell JBO (2014) Machine learning methods in chemoinformatics. WIREs Comput Mol Sci 4:468–481
Mitra I, Saha A, Roy K (2010) Exploring quantitative structure–activity relationship studies of antioxidant phenolic compounds obtained from traditional Chinese medicinal plants. Mol Simulat 36:1067–1079
Moss GP, Wilkinson SC, Sun Y (2012) Mathematical modelling of percutaneous absorption. Curr Opin Colloid Interface Sci 17:166–172
Ng S, Rouse J, Sanderson F, Eccleston G (2012) The relevance of polymeric synthetic membranes in topical formulation assessment and drug diffusion study. Arch Pharm Res 34:579–593
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H (2012) Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model 52:396–408
Ruby PK, Pathak SM, Deepika A (2014) Critical attributes of transdermal drug delivery system (TDDS)—a generic product development review. Drug Dev Ind Pharm 14 40(11):1421–1428
Shevade SK, Keerthi SS, Bhattacharyya C, Murthy KRK (1999) Improvements to SMO algorithm for SVM regression. Technical report CD-99-16, Control Division Dept of Mechanical and Production Engineering, National University of Singapore, Singapore
Smola AJ, Scholkopf B (2004) A tutorial on support vector regression. Stat Comput 14:199–222
Tropsha A, Gramatica P, Gombar VK (2003) The importance of being earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci 22:69–77
Varnek A, Baskin I (2012) Machine learning methods for property prediction in chemoinformatics: Quo Vadis? J Chem Inf Model 52:1413–1437
Wang WJ, Xu ZB, Lu WZ, Zhang XY (2003) Determination of the spread parameter in the Gaussian kernel for classification and regression. Neuro Comput 55:643–663
Witten IH, Frank E (2005) Data mining: practical machine learning tools and techniques. Morgan Kaufmann, San Francisco
Acknowledgments
This article does not contain any studies with human and animal subjects performed by any of the authors. These authors (B. Shaik, R. Gupta, B. Louis and V. K. Agrawal) declare that they have no conflict of interest. We acknowledge and thank Dr. Weiping Ma for the data set, which is used in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shaik, B., Gupta, R., Louis, B. et al. Prediction of permeability of drug-like compounds across polydimethylsiloxane membranes by machine learning methods. Journal of Pharmaceutical Investigation 45, 461–473 (2015). https://doi.org/10.1007/s40005-015-0194-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40005-015-0194-z