Abstract
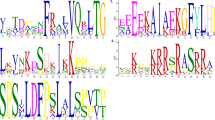
Protein motif is a conserved short region within larger sequence, and it provides an important way to get functional or structural information about a noble protein sequence. This study primarily provides a new approach to identify motifs which are considered to play an important role to determine functional or structural class of any protein. Coconut palm (Cocos nucifera L.) root (wilt) disease (CRWD) became a serious concern for coconut cultivation in the coastal region of South India. It has already been reported that species of the phytoplasma (-belonging to ribosomal group 16SrXI) is primarily associated with CRWD. In this study, our objective is to modify an existing motif identification algorithm, and also identify motifs that describe their conserved region within a CRWD phytoplasma protein sequence. Accordingly, the existing information content measure formulas for a no-gapped and gapped aligned sequence set were re-formulated. Further, an improvement on both these existing measures has been vied by incorporating prior information of BLOSUM90 substitution matrix. Results showed that proposed modifications could successfully identify N-terminal myristoylation motif within an alignment matrix of Arabidopsis thaliana sequences; concurrently our modification has also been validated. Both the approaches, namely frequency- and BLOSUM90-based information content, have been compared using the observed results, in terms of identified motifs. Finally, we could also test this noble BLOSUM90-based approach to correctly predict conserved regions at locations, (163–171) and (117–138), in an aligned secA gene sequence set of phytoplasma, an etiological agent of CRWD.

Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Bork P, Gibson TJ (1996) Applying motif and profile searches. Methods Enzymol 266:162–184
Bork P, Koonin EV (1996) Protein sequence motifs. Curr Opin Struct Biol 6(3):366–376
Charif D, Lobry J (2007) SeqinR 1.0-2: a contributed package to the R project for statistical computing devoted to biological sequences retrieval and analysis. In: Bastolla U, Porto M, Roman H, Vendruscolo M (eds) Structural approaches to sequence evolution: molecules, networks, populations, biological and medical physics, biomedical engineering. Springer Verlag, New York, pp 207–232 ISBN: 978-3-540-35305-8
Dreos R, Ambrosini G, Périer RC, Bucher P (2015) The eukaryotic promoter database: expansion of EPDnew and new promoter analysis tools. Nucleic Acids Res 43(D1):D92–D96
Gupta S, Kapoor V (1982) Fundamentals of mathematical statistics: a modern approach. Sultan Chand and Sons, New Delhi
Harrison NA, Helmick EE, Elliott ML (2008) Lethal yellowing-type diseases of palms associated with phytoplasmas newly identified in Florida, USA. Ann Appl Biol 153(1):85–94
Henikoff S, Henikoff JG (1992) Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci 89(22):10,915–10,919
Hertz GZ, Stormo GD (1995) Identification of consensus patterns in unaligned DNA and protein sequences: a large-deviation statistical basis for penalizing gaps. Proc Third Int Conf Bioinf Genome Res 2:201–216
Hertz GZ, Stormo GD (1999) Identifying DNA and protein patterns with statistically significant alignments of multiple sequences. Bioinformatics 15(7):563–577
Hertz GZ, Hartzell GW, Stormo GD (1990) Identification of consensus patterns in unaligned DNA sequences known to be functionally related. Comput Appl Biosci: CABIOS 6(2):81–92
Huan HX, Tuyet DT, Ha DT, Hung NT (2015) An efficient ant colony algorithm for DNA motif finding. In: Nguyen VH, Le AC, Huynh VN (eds) Knowledge and systems engineering, vol 326. Advances in Intelligent Systems and Computing, Springer, pp 589–601
Johnson DR, Bhatnagar RS, Knoll LJ, Gordon JI (1994) Genetic and biochemical studies of protein N-myristoylation. Annu Rev Biochem 63(1):869–914
Johnson NL, Kotz S, Balakrishnan N (1997) Discrete multivariate distributions, vol 165. Wiley, New York
Kaiser F, Eisold A, Labudde D (2015) A novel algorithm for enhanced structural motif matching in proteins. J Comput Biol 22:698–713
Ladunga IS (2003) Finding homologs in amino acid sequences using network BLAST searches. Curr Protoc Bioinf 25:3–4
Li F, Liu J, Valls L, Hiser C, Ferguson-Miller S (2015) Identification of a key cholesterol binding enhancement motif in translocator protein 18 kDa. Biochemistry 54(7):1441–1443
Liu S, Trenkler G (2008) Hadamard, Khatri-Rao, Kronecker and other matrix products. Int J Inf Syst Sci 4(1):160–177
Liu Z, Li F, Zhang B, Li S, Wu J, Shi Y (2015) Structural basis of plant homeodomain finger 6 (PHF6) recognition by retinoblastoma binding protein 4 (RBBP4) component of the nucleosome remodeling and deacetylase (NuRD) complex. J Biol Chem 290:6630–6638
Manimekalai R, Soumya V, Nair S, Thomas GV, Baranwal V (2014) Molecular characterization identifies 16srxi-b group phytoplasma (candidatus phytoplasma oryzae-related strain) associated with root wilt disease of coconut in india. Sci Hortic 165:288–294
Myrie W, Harrison N, Douglas L, Helmick E, Gore-Francis J, Oropeza WC, McLaughlin (2014) First report of lethal yellowing disease associated with subgroup 16SrIV-A phytoplasmas in Antigua, West Indies. New Dis Rep 29(1):12
Nipah JO, Jones P, Dickinson MJ (2007) Detection of lethal yellowing phytoplasma in embryos from coconut palms infected with cape St Paul wilt disease in Ghana. Plant Pathol 56:777–784
Ou J, Zhu LJ (2013) motifStack: plot stacked logos for single or multiple DNA, RNA and amino acid sequence. R Package Version 1(8):1
Perera L, Meegahakumbura MK, Wijesekara HRT, Fernando WBS, Dickinson MJ (2012) A phytoplasma is associated with the Weligama coconut leaf wilt disease in Sri Lanka. J Plant Pathol 94(1):205–209
Pruitt KD, Tatusova T, Maglott DR (2007) NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 35(suppl 1):D61–D65
Schneider TD, Stormo GD, Gold L, Ehrenfeucht A (1986) Information content of binding sites on nucleotide sequences. J Mol Biol 188(3):415–431
Shil S, Das KK, Dutta A (2015) Searching of conserved motifs within a partial seca gene sequence of phytoplasma associated with root (wilt) disease of coconut (Cocos nucifera) in India: using a frequency based approach. Int J Bio-resour Stress Manag 6(2):208–213
Sun T, Shi X, Friso G, Van Wijk K, Bentolila S, Hanson MR (2015a) A zinc-finger motif-containing protein is essential for chloroplast rna editing. PLoS Genetics 11(3):e1005,028–e1005,028
Sun W, Chen H, Wang J, Sun HW, Yang SK, Sang YL, Lu XB, Xu XH (2015b) Expression analysis of genes encoding mitogen-activated protein kinases in maize provides a key link between abiotic stress signaling and plant reproduction. Funct Integr Genom 15(1):107–120
Team RC (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing. http://www.R-project.org/
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL\_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Wu T, Brutlag D (1994) Identification of protein motifs using conserved amino acid properties and partitioning techniques. In: Proceedings. International conference on intelligent systems for molecular biology; ISMB, vol 3, pp 402–410
Xia X (2011) Position weight matrix, Gibbs sampler, and the associated significance tests in motif characterization and prediction. Scientifica 2012:917,540–917,540
Yamauchi S, Fusada N, Hayashi H, Utsumi T, Uozumi N, Endo Y, Tozawa Y (2010) The consensus motif for N-myristoylation of plant proteins in a wheat germ cell-free translation system. FEBS J 277(17):3596–3607
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shil, S., Das, K.K. & Saxena, V.K. Identification of Protein Motifs in Phytoplasma Associated with Root (Wilt) Disease of Coconut (Cocos nucifera) Using an Improved Statistical Measure. Agric Res 8, 56–64 (2019). https://doi.org/10.1007/s40003-018-0351-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40003-018-0351-5