Abstract
Background
The World Health Organization announced the end of the Coronavirus Disease of 2019 (COVID-19) global health emergency on May 5, 2023. However, the reports from different countries indicate an elevation in the number of COVID-19-related hospitalizations and deaths through the last months. The subvariant XBB.1.5 (Kraken) was the cause of 49.1% of COVID-19 cases by the end of January 2023. Although, the subvariant EG.5 (Eris) has surpassed the XBB.1.5 recently. EG.5 is a close subvariant descending from XBB.1.9.2 subvariant of Omicron. EG.5.1 is a sublineage carrying two crucial spike mutations F456L and Q52H. Up to now, it is not well-established whether its infectivity, severity, and immune evasion have shown any change or not. Also, BA.2.86 another subvariant of Omicron descending from BA.2 bears over 30 mutations which could affect its infectivity and transmissibility.
Methods
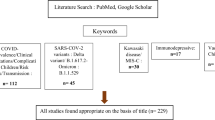
Scopus, PubMed, Google Scholar, and Google were searched with six keywords up to 20 November 2023 and highly reliable research and reports were selected to refer to in this article.
Purpose
This brief review aims to overview the most reliable data about EG.5 and BA.2.86 based on scientific evidence.
Conclusion
Based on the currently available data these two new subvariants have similar features with currently circulating variants of Omicron and are less immune evasive than ancestral SARS-CoV-2.


Similar content being viewed by others
Data availablity
All data generated or analyzed during this study are included in this published article.
Abbreviations
- ACE2:
-
Angiotensin-converting enzyme 2
- COVID-19:
-
Coronavirus disease of 2019
- RBD:
-
Receptor binding domain
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- VUM:
-
Variant under monitoring
- VOI:
-
Variant of interest
- WHO:
-
World Health Organization
- ICU:
-
Intensive Care Unit
- NTD:
-
N-terminal domain
- US:
-
United States
- UK:
-
United Kingdom
References
Mohamadian M, Chiti H, Shoghli A, Biglari S, Parsamanesh N, Esmaeilzadeh A. COVID-19: virology, biology and novel laboratory diagnosis. J Gene Med. 2021;23:e3303. https://doi.org/10.1002/jgm.3303.
WHO COVID-19 Dashboard. Geneva: World Health Organization; 2020. [Internet]. https://covid19.who.int. Accessed 13 Dec 2023.
Chatterjee S, Bhattacharya M, Nag S, Dhama K, Chakraborty C. A detailed overview of SARS-CoV-2 omicron: its sub-variants, mutations and pathophysiology, clinical characteristics, immunological landscape, immune escape, and therapies. Viruses. 2023;15:167. https://doi.org/10.3390/v15010167.
Bazargan M, Elahi R, Esmaeilzadeh A. OMICRON: virology, immunopathogenesis, and laboratory diagnosis. J Gene Med. 2022;24:e3435. https://doi.org/10.1002/jgm.3435.
Looi M-K. COVID-19: scientists sound alarm over new B.A 2.86 “Pirola” variant. Br Med J. 2023;382:1964. https://doi.org/10.1136/bmj.p1964.
Looi M-K. Covid-19: Hospital admissions rise in England amid fears of new variant and waning immunity. Br Med J. 2023;382:1833. https://doi.org/10.1136/bmj.p1833.
Abbasi J. What to Know About EG. 5, the latest SARS-CoV-2 “Variant of Interest.” JAMA. 2023. https://doi.org/10.1001/jama.2023.16498.
Esmaeilzadeh A, Maleki AJ, Moradi A, Siahmansouri A, Yavari MJ, Karami P, et al. Major severe acute respiratory coronavirus-2 (SARS-CoV-2) vaccine-associated adverse effects; benefits outweigh the risks. Expert Rev Vaccines. 2022;21:1377–94. https://doi.org/10.1080/14760584.2022.2116008.
Centers for Disease Control and Prevention (CDC). COVID Data Tracker. [Internet]. 2023. https://covid.cdc.gov/covid-data-tracker/#variant-proportions. Accessed 3 Sept 2023.
Elahi R, Hozhabri S, Moradi A, Siahmansouri A, Jahani Maleki A, Esmaeilzadeh A. Targeting the cGAS-STING pathway as an inflammatory crossroad in coronavirus disease 2019 (COVID-19). Immunopharmacol Immunotoxicol. 2023. https://doi.org/10.1080/08923973.2023.2215405.
Viana R, Moyo S, Amoako DG, Tegally H, Scheepers C, Althaus CL, et al. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature. 2022;603:679–86. https://doi.org/10.1038/s41586-022-04411-y.
Kumar S, Karuppanan K, Subramaniam G. Omicron (BA. 1) and sub-variants (BA. 11, BA. 2, and BA. 3) of SARS-CoV-2 spike infectivity and pathogenicity: a comparative sequence and structural-based computational assessment. J Med Virol. 2022;94:4780–91. https://doi.org/10.1002/jmv.27927.
Mohapatra RK, Kandi V, Verma S, Dhama K. Challenges of the omicron (B. 1.1. 529) variant and its lineages: a global perspective. ChemBioChem. 2022;23: e202200059. https://doi.org/10.1002/cbic.202200059.
Parums DV. The XBB. 1.5 (‘Kraken’) Subvariant of Omicron SARS-CoV-2 and its Rapid Global Spread. Med Sci Monit Int Med J Exp Clin Res. 2023;29:e939580-1.
Pagel C. Covid is on the rise again—so what next? BMJ. 2023. https://doi.org/10.1136/bmj.p1885.
World Health Organization (WHO). EG.5 initial risk evaluation. 9 August 2023. [Internet]. https://www.who.int/docs/default-source/coronaviruse/09082023eg.5_ire_final.pdf
Dyer O. Covid-19: infections climb globally as EG. 5 variant gains ground. BMJ. 2023. https://doi.org/10.1136/bmj.p1900.
Scarpa F, Pascarella S, Ciccozzi A, Giovanetti M, Azzena I, Locci C, et al. Genetic and structural analyses reveal the low potential of the SARS-CoV-2 EG. 5 variant. J Med Virol. 2023;95:e29075. https://doi.org/10.1002/jmv.29075.
Girma A. The many mutations of the COVID-19 variant: current perspectives on EG. 5/Eris. Environmental health insights. 2023;17. https://doi.org/10.1177/11786302231217805
Pascarella S, Ciccozzi M, Bianchi M, Benvenuto D, Cauda R, Cassone A. The electrostatic potential of the Omicron variant spike is higher than in Delta and Delta-plus variants: a hint to higher transmissibility? J Med Virol. 2021. https://doi.org/10.1002/jmv.27528.
Mouliou DS. The deceptive COVID-19: lessons from common molecular diagnostics and a novel plan for the prevention of the next pandemic. Diseases. 2023;11:20. https://doi.org/10.3390/diseases11010020.
Mouliou DS, Gourgoulianis KI. False-positive and false-negative COVID-19 cases: respiratory prevention and management strategies, vaccination, and further perspectives. Expert Rev Respir Med. 2021;15:993–1002. https://doi.org/10.1080/17476348.2021.1917389.
Yisimayi A, Song W, Wang J, Jian F, Yu Y, Chen X, et al. Repeated Omicron infection alleviates SARS-CoV-2 immune imprinting. bioRxiv. 2023. https://doi.org/10.1101/2023.05.01.538516.
Kaku Y, Kosugi Y, Uriu K, Ito J, Kuramochi J, Sadamasu K, et al. Antiviral efficacy of the SARS-CoV-2 XBB breakthrough infection sera against Omicron subvariants including EG. 5. bioRxiv. 2023. https://doi.org/10.1101/2023.08.08.552415.
yalemedicine. [Internet]. 2023. https://www.yalemedicine.org/news/covid-eg5-eris-latest-coronavirus-strain. Accessed 3 Sept 2023.
World Health Organization (WHO). Weekly-epidemiological-update-on-covid-19. [Internet]. 2023. https://www.who.int/publications/m/item/weekly-epidemiological-update-on-covid-19. Accessed 1 Sept 2023.
Lee WS, Wheatley AK, Kent SJ, DeKosky BJ. Antibody-dependent enhancement and SARS-CoV-2 vaccines and therapies. Nat Microbiol. 2020;5:1185–91. https://doi.org/10.1038/s41564-020-00789-5.
Food and Drug administration (FDA). Vaccines, blood, and biologics. Updated COVID-19 Vaccines for Use in the United States Beginning in Fall. 2023. [Internet]. https://www.fda.gov/vaccines-blood-biologics/updated-covid-19-vaccines-use-united-states-beginning-fall-2023. Accessed 16 June 2023.
Meo SA, Meo AS, Klonoff DC. Omicron new variant BA.2.86 (Pirola): epidemiological, biological, and clinical characteristics—a global data-based analysis. Eur Rev Med Pharmacol Sci. 2023;27:9470–6. https://doi.org/10.26355/EURREV_202310_33975.
Hu Y, Zou J, Kurhade C, Deng X, Chang HC, Kim DK, et al. Less neutralization evasion of SARS-CoV-2 BA. 2.86 than XBB sublineages and CH. 1.1. bioRxiv. 2023. https://doi.org/10.1101/2023.09.10.557047.
Khan K, Lustig G, Reedoy K, Jule Z, Romer C, Karim F, et al. Evolution and neutralization escape of the SARS-CoV-2 BA. 2.86 subvariant. medRxiv. 2023. https://doi.org/10.1101/2023.09.08.23295250.
Uriu K, Ito J, Kosugi Y, Tanaka YL, Mugita Y, Guo Z, et al. Transmissibility, infectivity, and immune evasion of the SARS-CoV-2 BA.2.86 variant. Lancet Infect Dis. 2023;23:e460–1. https://doi.org/10.1016/S1473-3099(23)00575-3.
Wang Q, Guo Y, Liu L, Schwanz LT, Li Z, Nair MS, et al. Antigenicity and receptor affinity of SARS-CoV-2 BA.2.86 spike. Nature. 2023. https://doi.org/10.1038/s41586-023-06750-w.
Yang S, Yu Y, Jian F, Song W, Yisimayi A, Chen X, et al. Antigenicity and infectivity characterisation of SARS-CoV-2 BA. 2.86. Lancet Infect Dis. 2023;23:e457–9. https://doi.org/10.1016/S1473-3099(23)00573-X.
Centers for Disease Control and Prevention (CDC). Respiratory Viruses. [Internet]. 2023. https://www.cdc.gov/respiratory-viruses/whats-new/covid-19-variant.html
Mouliou DS, Gourgoulianis KI. COVID-19 ‘asymptomatic’ patients: an old wives’ tale. Expert Rev Respir Med. 2022;16:399–407. https://doi.org/10.1080/17476348.2022.2030224.
Funding
The authors have no relevant financial or non-financial interests to disclose.
Author information
Authors and Affiliations
Contributions
FE, AJM, and AS contributed to data gathering, writing the manuscript, and designing figures. AE contributed to the hypothesis, correspondence, and revising of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest or personal relationships that could have appeared to influence any aspects of this work.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Esmaeilzadeh, A., Ebrahimi, F., Jahani Maleki, A. et al. EG.5 (Eris) and BA.2.86 (Pirola) two new subvariants of SARS-CoV-2: a new face of old COVID-19. Infection 52, 337–343 (2024). https://doi.org/10.1007/s15010-023-02146-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s15010-023-02146-0