Abstract
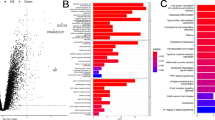
Acute myocardial infarction (AMI) is a coronary artery disease that prevents blood and oxygen flow from reaching the heart. The underlying complex pathophysiology of AMI has made the early detection process very challenging. A potential biomarker with high accuracy in determining AMI can help us in the risk stratification of patients with chest pain and also reduce treatment costs. This study aimed to identify the key genes for early AMI detection from the expression data of peripheral blood samples. We retrieved three GEO datasets from NCBI that represent expression data of healthy individuals and early-stage AMI patients. The differentially expressed genes (DEG) were determined from three datasets by the GEO2R tool on the NCBI webpage. The significant DEGs common in at least 2 datasets were identified by VENNY 2.1 web tool. We then performed a functional enrichment analysis of the selected genes and also the potential hub genes possibly involved in AMI were predicted by a protein–protein interaction network. Finally, a drug–gene interaction network was constructed. We found 5360, 2049, and 579 genes, respectively, from the GSE61144, GSE60993, and GSE29532 datasets to be the significant DEGs in GEO2R analysis. A total of 214 genes were found common in at least two datasets. CD59, FCAR, CLEC5A, CKAP4, and CEACAM8 are the most significant hub genes predicted by protein–protein network analysis that has a close relationship with the early response to inflammation in AMI. Our study suggests CD59, FCAR, CLEC5A, CKAP4, and CEACAM8 as the potential inflammatory biomarkers for the early-stage detection of AMI.






Similar content being viewed by others
Data availability
The datasets supporting the conclusions of this study are included within the article (and its additional files).
References
Ajani UA, Ford ES (2006) Has the risk for coronary heart disease changed among U.S. adults? J Am College Cardiol 48(6), 1177–1182. https://doi.org/10.1016/j.jacc.2006.05.055
Aydin S, Ugur K, Aydin S, Sahin İ, Yardim M (2019) Biomarkers in acute myocardial infarction: current perspectives. Vascular Health Risk Manag 15:1–10. https://doi.org/10.2147/VHRM.S166157
Boley SJ, Feinstein FR, Sammartano R, Brandt LJ, Sprayregen S (1981) New concepts in the management of emboli of the superior mesenteric artery. Int Abstr Surg 153(4):561–569
Brandes U (2001) A faster algorithm for betweenness centrality. J Math Soc 25(2):163–177. https://doi.org/10.1080/0022250X.2001.9990249
Braunwald E (2012) Unstable Angina and Non–ST Elevation Myocardial Infarction. Am J Respir Crit Care Med 185(9):924–932. https://doi.org/10.1164/rccm.201109-1745CI
Chan D, Ng LL (2010) Biomarkers in acute myocardial infarction. BMC Med 8(1):34. https://doi.org/10.1186/1741-7015-8-34
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, Clark NR, Ma’ayan, A. (2013) Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform 14:128. https://doi.org/10.1186/1471-2105-14-128
Chen J, Yu L, Zhang S, Chen X (2016) Network Analysis-Based Approach for Exploring the Potential Diagnostic Biomarkers of Acute Myocardial Infarction. Front Physiol. https://doi.org/10.3389/fphys.2016.00615
Chen Q, Yin Q, Song J, Liu C, Chen H, Li S (2021) Identification of monocyte-associated genes as predictive biomarkers of heart failure after acute myocardial infarction. BMC Med Genomics 14(1):44. https://doi.org/10.1186/s12920-021-00890-6
Chin C-H, Chen S-H, Wu H-H, Ho C-W, Ko M-T, Lin C-Y (2014) cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(4):S11. https://doi.org/10.1186/1752-0509-8-S4-S11
Davies A, Simmons DL, Hale G, Harrison RA, Tighe H, Lachmann PJ, Waldmann H (1989) CD59, an LY-6-like protein expressed in human lymphoid cells, regulates the action of the complement membrane attack complex on homologous cells. J Exp Med 170(3):637–654. https://doi.org/10.1084/jem.170.3.637
Frangogiannis NG (2012) Regulation of the inflammatory response in cardiac repair. Circ Res 110(1):159–173. https://doi.org/10.1161/CIRCRESAHA.111.243162
Frangogiannis NG (2014) The inflammatory response in myocardial injury, repair, and remodelling. Nat Rev Cardiol 11(5):255–265. https://doi.org/10.1038/nrcardio.2014.28
GEO Overview—GEO - NCBI. (n.d.). Retrieved 27 June 2021, from https://www.ncbi.nlm.nih.gov/geo/info/overview.html
GEO2R—GEO—NCBI. (n.d.). Retrieved 27 June 2021, from https://www.ncbi.nlm.nih.gov/geo/geo2r/
Gladka MM, Molenaar B, de Ruiter H, van der Elst S, Tsui H, Versteeg D, Lacraz GPA, Huibers MMH, van Oudenaarden A, van Rooij E (2018) Single-cell sequencing of the healthy and diseased heart reveals cytoskeleton-associated protein 4 as a new modulator of fibroblasts activation. Circulation 138(2):166–180. https://doi.org/10.1161/CIRCULATIONAHA.117.030742
Griffith M, Griffith OL, Coffman AC, Weible JV, McMichael JF, Spies NC, Koval J, Das I, Callaway MB, Eldred JM, Miller CA, Subramanian J, Govindan R, Kumar RD, Bose R, Ding L, Walker JR, Larson DE, Dooling DJ, Wilson RK (2013) DGIdb: mining the druggable genome. Nat Methods 10(12):1209–1210. https://doi.org/10.1038/nmeth.2689
Hong, C., & Hauskrecht, M. (2015). MCODE: Multivariate conditional outlier detection. ArXiv:1505.04097 [Cs, Stat]
Hsieh SL. (2016). C-Type Lectin Receptors in Immunity (Vol. 1–Chapter 3). Springer Japan. https://doi.org/10.1007/978-4-431-56015-9
Iakoubova OA, Tong CH, Chokkalingam AP, Rowland CM, Kirchgessner TG, Louie JZ, Ploughman LM, Sabatine MS, Campos H, Catanese JJ, Leong DU, Young BA, Lew D, Tsuchihashi Z, Luke MM, Packard CJ, Zerba KE, Shaw PM, Shepherd J, Sacks FM (2006) Asp92Asn polymorphism in the myeloid IgA Fc receptor is associated with myocardial infarction in two disparate populations: CARE and WOSCOPS. Arterioscler Thromb Vasc Biol 26(12):2763–2768. https://doi.org/10.1161/01.ATV.0000247248.76409.8b
Iakoubova OA, Tong CH, Rowland CM, Kirchgessner TG, Young BA, Arellano AR, Shiffman D, Sabatine MS, Campos H, Packard CJ, Pfeffer MA, White TJ, Braunwald E, Shepherd J, Devlin JJ, Sacks FM (2008) Association of the Trp719Arg polymorphism in kinesin-like protein 6 with myocardial infarction and coronary heart disease in 2 prospective trials: the CARE and WOSCOPS trials. J Am Coll Cardiol 51(4):435–443. https://doi.org/10.1016/j.jacc.2007.05.057
Kaartinen M, Penttilä A, Kovanen PT (1994) Accumulation of activated mast cells in the shoulder region of human coronary atheroma, the predilection site of atheromatous rupture. Circulation 90(4):1669–1678. https://doi.org/10.1161/01.cir.90.4.1669
Kimberley FC, Sivasankar B, Paul Morgan B (2007) Alternative roles for CD59. Mol Immunol 44(1):73–81. https://doi.org/10.1016/j.molimm.2006.06.019
Kimura H, Fumoto K, Shojima K, Nojima S, Osugi Y, Tomihara H, Eguchi H, Shintani Y, Endo H, Inoue M, Doki Y, Okumura M, Morii E, Kikuchi A (2016) CKAP4 is a Dickkopf1 receptor and is involved in tumor progression. J Clin Investig 126(7):2689–2705. https://doi.org/10.1172/JCI84658
Li M, Chen F, Zhang Y, Xiong Y, Li Q, Huang H (2020) Identification of post-myocardial infarction blood expression signatures using multiple feature selection strategies. Front Physiol 11:483. https://doi.org/10.3389/fphys.2020.00483
Liu Z, Ma C, Gu J, Yu M (2019) Potential biomarkers of acute myocardial infarction based on weighted gene co-expression network analysis. Biomed Eng Online 18(1):9. https://doi.org/10.1186/s12938-019-0625-6
Ma Y, Yabluchanskiy A, Lindsey ML (2013) Neutrophil roles in left ventricular remodeling following myocardial infarction. Fibrogenesis Tissue Repair 6(1):11. https://doi.org/10.1186/1755-1536-6-11
Montalescot G, Cayla G, Collet J-P, Elhadad S, Beygui F, Le Breton H, Choussat R, Leclercq F, Silvain J, Duclos F, Aout M, Dubois-Randé J-L, Barthélémy O, Ducrocq G, Bellemain-Appaix A, Payot L, Steg P-G, Henry P, Spaulding C, Investigators ABOARD (2009) Immediate vs delayed intervention for acute coronary syndromes: a randomized clinical trial. JAMA 302(9):947–954. https://doi.org/10.1001/jama.2009.1267
Nichols M, Townsend N, Scarborough P, Rayner M (2014) Cardiovascular disease in Europe 2014: epidemiological update. Eur Heart J 35(42):2950–2959. https://doi.org/10.1093/eurheartj/ehu299
O, ’Gara Patrick T., Kushner, F. G., Ascheim, D. D., Casey, D. E., Chung, M. K., de, L. J. A., Ettinger, S. M., Fang, J. C., Fesmire, F. M., Franklin, B. A., Granger, C. B., Krumholz, H. M., Linderbaum, J. A., Morrow, D. A., Newby, L. K., Ornato, J. P., Ou, N., Radford, M. J., Tamis, -Holland Jacqueline E., … Zhao, D. X. (2013). 2013 ACCF/AHA Guideline for the Management of ST-Elevation Myocardial Infarction. Journal of the American College of Cardiology, 61(4), e78–e140. https://doi.org/10.1016/j.jacc.2012.11.019
Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, Kanehisa M (1999) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 27(1):29–34. https://doi.org/10.1093/nar/27.1.29
OLIVEROS, J. C. (2007). VENNY. An interactive tool for comparing lists with Venn diagrams. Http://Bioinfogp.Cnb.Csic.Es/Tools/Venny/Index.Html, https://ci.nii.ac.jp/naid/20001505977/.
Ong S-B, Hernández-Reséndiz S, Crespo-Avilan GE, Mukhametshina RT, Kwek X-Y, Cabrera-Fuentes HA, Hausenloy DJ (2018) Inflammation following acute myocardial infarction: Multiple players, dynamic roles, and novel therapeutic opportunities. Pharmacol Ther 186:73–87. https://doi.org/10.1016/j.pharmthera.2018.01.001
Park, H.-J., Noh, J. H., Eun, J. W., Koh, Y.-S., Seo, S. M., Park, W. S., Lee, J. Y., Chang, K., Seung, K. B., Kim, P.-J., & Nam, S. W. (2015). Assessment and diagnostic relevance of novel serum biomarkers for early decision of ST-elevation myocardial infarction. Oncotarget, 6(15), 12970–12983. https://doi.org/10.18632/oncotarget.4001
Reed GW, Rossi JE, Cannon CP (2017) Acute myocardial infarction. The Lancet 389(10065):197–210. https://doi.org/10.1016/S0140-6736(16)30677-8
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Silbiger VN, Luchessi AD, Hirata RDC, Lima-Neto LG, Cavichioli D, Carracedo A, Brión M, Dopazo J, García-García F, dos Santos ES, Ramos RF, Sampaio MF, Armaganijan D, Sousa AGMR, Hirata MH. (2013) Novel genes detected by transcriptional profiling from whole-blood cells in patients with early onset of acute coronary syndrome. Clinica Chimica Acta 421:184–190. https://doi.org/10.1016/j.cca.2013.03.011
Stelzl U, Worm U, Lalowski M, Haenig C, Brembeck FH, Goehler H, Stroedicke M, Zenkner M, Schoenherr A, Koeppen S, Timm J, Mintzlaff S, Abraham C, Bock N, Kietzmann S, Goedde A, Toksöz E, Droege A, Krobitsch S, Wanker EE (2005) A human protein-protein interaction network: a resource for annotating the proteome. Cell 122(6):957–968. https://doi.org/10.1016/j.cell.2005.08.029
Sung P-S, Chang W-C, Hsieh S-L (2020) CLEC5A: a promiscuous pattern recognition receptor to microbes and beyond. Adv Exp Med Biol 1204:57–73. https://doi.org/10.1007/978-981-15-1580-4_3
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, von Mering C (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Thygesen K, Alpert JS, Jaffe AS, Simoons ML, Chaitman BR, White HD, Joint ESC/ACCF/AHA/WHF Task Force for Universal Definition of Myocardial Infarction, Authors/Task Force Members Chairpersons, Thygesen, K., Alpert, J. S., White, H. D., Biomarker Subcommittee, Jaffe, A. S., Katus, H. A., Apple, F. S., Lindahl, B., Morrow, D. A., ECG Subcommittee, Chaitman, B. R., … Wagner, D. R. (2012). Third universal definition of myocardial infarction. Journal of the American College of Cardiology, 60(16), 1581–1598. https://doi.org/10.1016/j.jacc.2012.08.001
Tilea I, Varga A, Serban RC (2021) Past, present, and future of blood biomarkers for the diagnosis of acute myocardial infarction—promises and challenges. Diagnostics 11(5):881. https://doi.org/10.3390/diagnostics11050881
Xie Y, Wang Y, Zhao L, Wang F, Fang J (2021) Identification of potential biomarkers and immune cell infiltration in acute myocardial infarction (AMI) using bioinformatics strategy. Bioengineered 12(1):2890–2905. https://doi.org/10.1080/21655979.2021.1937906
Xiong Y-Y, Gong Z-T, Tang R-J, Yang Y-J (2021) The pivotal roles of exosomes derived from endogenous immune cells and exogenous stem cells in myocardial repair after acute myocardial infarction. Theranostics 11(3):1046–1058. https://doi.org/10.7150/thno.53326
Zhao H, Jiang A, Yu M, Bao H (2020) Identification of biomarkers correlated with diagnosis and prognosis of endometrial cancer using bioinformatics analysis. J Cell Biochem 121(12):4908–4921. https://doi.org/10.1002/jcb.29819
Zhao L, Xu S, Fjaertoft G, Pauksen K, Håkansson L, Venge P (2004) An enzyme-linked immunosorbent assay for human carcinoembryonic antigen-related cell adhesion molecule 8, a biological marker of granulocyte activities in vivo. J Immunol Methods 293(1):207–214. https://doi.org/10.1016/j.jim.2004.08.009
Author information
Authors and Affiliations
Contributions
MTS, SS, PB, MTI, MNH carried out the data analysis, bioinformatics analysis, interpreted the results and wrote the article. MAS, MOR contributed to the bioinformatics analysis and revised the article. MTS, MOR, contributed to the interpretation of the results and revised the article. MTS, SS, PB, MOR, NI, and MMIR examined the literature and prepare the figures. MTS, SS, PB, NI, MMIR, and MOR contributed to the interpretation of the results and revised the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics statement
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sarker, M.T., Saha, S., Biswas, P. et al. Identification of blood-based inflammatory biomarkers for the early-stage detection of acute myocardial infarction. Netw Model Anal Health Inform Bioinforma 11, 28 (2022). https://doi.org/10.1007/s13721-022-00371-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13721-022-00371-5