Abstract
Glypican-3 (GPC3) is a heparan sulfate proteoglycan that may function as a tumor suppressor in breast cancer (BC). To evaluate the prognostic value of GPC3 in BC, systematic analysis was performed in this study. To evaluate gene alteration during breast carcinogenesis, GPC3 expression was analyzed using the Oncomine, GENT, UALCAN, bcGenExMiner, and UCSC Xena databases. The prognostic role of GPC3 in BC was investigated using KM Plotter and PrognoScan databases. Promoter methylation status and heat map of GPC3 were determined using UALCAN and UCSC Xena. GPC3 expression was significantly downregulated in BC compared to that in normal tissues and was correlated with prognosis. However, estrogen receptor and progesterone receptor status were positively correlated with GPC3 expression, whereas basal-like status, triple-negative breast cancer status, and Scarff, Bloom, and Richardson grade criteria were negatively correlated with GPC3 expression. Further analysis indicated that GPC3 was correlated with Ras-association domain family 6 (RASSF6) expression in BC tissues. GPC3 may thus be considered a significant marker for predicting BC prognosis along with RASSF6. Comparative protein modeling of GPC3 was performed using a FASTA formatted sequence from NCBI, in Swiss model, and GPC3 was visualized using PyMOL tools. The GPC3 active site was identified with CASTp server. The Ramachandran plot in PROCHECK showed significant scores for the protein model in its most favored regions. ProSA server indicated the high accuracy of the protein model. However, large-scale and comprehensive studies are needed to clarify these results.

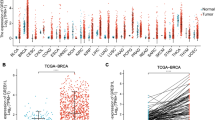
available at https://medical-genomics.kribb.re.kr/GENT/). Boxes represent the median and the 25th and 75th percentiles; dots represent outliers. Red boxes represent tumor tissues; green boxes represent normal tissues. Red and green dashed lines represent the average value of all tumor and normal tissues, respectively









Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42(W1):W252–W258. https://doi.org/10.1093/nar/gku340
Buchanan C, Lago Huvelle MA, Peters GM (2011) Metastasis suppressors: basic and translational advances. Curr Pharm Biotechnol 12(11):1948–1960. https://doi.org/10.2174/138920111798376914
Capurro M, Wanless IR, Sherman M, Deboer G, Shi W, Miyoshi E et al (2003) Glypican-3: a novel serum and histochemical marker for hepatocellular carcinoma. Gastroenterology 125(1):89–97. https://doi.org/10.1016/S0016-5085(03)00689-9
Castillo LF, Tascón R, Lago Huvelle MA, Novack G, Llorens MC, Dos Santos AF et al (2016) Glypican-3 induces a mesenchymal to epithelial transition in human breast cancer cells. Oncotarget 7(37):60133–60154. https://doi.org/10.18632/oncotarget.11107
Collaboration o. D. C. (2015Global) Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 32 cancer groups, 1990 to 2015: a systematic analysis for the global burden of disease study global Burden of Cancer 2015Global Burden of Cancer 2015. JAMA Oncol 3(4):524–548. https://doi.org/10.1001/jamaoncol.2016.5688
Dundas J, Ouyang Z, Tseng J, Binkowski A, Turpaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34(suppl_2):W116–W118. https://doi.org/10.1093/nar/gkl282
Fernández D, Guereño M, Lago Huvelle MA, Cercato M, Peters MG (2018) Signaling network involved in the GPC3-induced inhibition of breast cancer progression: role of canonical Wnt pathway. J Cancer Res Clin Oncol 144(12):2399–2418. https://doi.org/10.1007/s00432-018-2751-0
Filmus J (2001) Glypicans in growth control and cancer. Glycobiology 11(3):19R–23R. https://doi.org/10.1093/glycob/11.3.19R
Filmus J, Selleck SB (2001) Glypicans: proteoglycans with a surprise. J Clin Investig 108(4):497–501. https://doi.org/10.1172/jci13712
Guo W, Dong Z, Guo Y, Shen S, Guo X, Kuang G et al (2016) Decreased expression and frequent promoter hypermethylation of RASSF2 and RASSF6 correlate with malignant progression and poor prognosis of gastric cardia adenocarcinoma. Mol Carcinog 55(11):1655–1666. https://doi.org/10.1002/mc.22416
Han S, Ma X, Zhao Y, Zhao H, Batista A, Zhou S et al (2016) Identification of Glypican-3 as a potential metastasis suppressor gene in gastric cancer. Oncotarget 7(28):44406–44416. https://doi.org/10.18632/oncotarget.9763
Ho M, Kim H (2011) Glypican-3: a new target for cancer immunotherapy. Eur J Cancer 47(3):333–338. https://doi.org/10.1016/j.ejca.2010.10.024
Iglesias BV, Centeno G, Pascuccelli H, Ward F, Peters MG, Filmus J et al (2008) Expression pattern of glypican-3 (GPC3) during human embryonic and fetal development [Research Support, Non-U S Gov't]. Histol Histopathol 23(11):1333–1340
Kuriata A, Gierut AM, Oleniecki T, Ciemny MP, Kolinski A, Kurcinski M et al (2018) CABS-flex 2.0: a web server for fast simulations of flexibility of protein structures. Nucleic Acids Res 46(W1):W338–W343. https://doi.org/10.1093/nar/gky356
Laskowski RA, Chistyakov VV, Thornton JM (2005) PDBsum more: new summaries and analyses of the known 3D structures of proteins and nucleic acids. Nucleic Acids Res 33(suppl_1):D266–D268. https://doi.org/10.1093/nar/gki001
Laskowski RA, Rullmann JAC, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8(4):477–486. https://doi.org/10.1007/bf00228148
Liang J, Woodward C, Edelsbrunner H (1998) Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci 7(9):1884–1897. https://doi.org/10.1002/pro.5560070905
Lu X, Ma J, Chu J, Shao Q, Zhang Y, Lu G et al (2017) MiR-129-5p sensitizes the response of Her-2 positive breast cancer to trastuzumab by reducing Rps6. Cell Physiol Biochem 44(6):2346–2356. https://doi.org/10.1159/000486122
Maeda D, Ota S, Takazawa Y, Aburatani H, Nakagawa S, Yano T et al (2009) Glypican-3 expression in clear cell adenocarcinoma of the ovary [Original Article]. Mod Pathol 22:824. https://doi.org/10.1038/modpathol.2009.40
Nakatsura T, Kageshita T, Ito S, Wakamatsu K, Monji M, Ikuta Y et al (2004) Identification of glypican-3 as a novel tumor marker for melanoma. Clin Cancer Res 10(19):6612–6621. https://doi.org/10.1158/1078-0432.ccr-04-0348
Patra P, Ghosh P, Patra BC, Bhattacharya M (2019) Biocomputational analysis and in silico characterization of an angiogenic protein (RNase5) in zebrafish (Danio rerio). Int J Pept Res Ther. https://doi.org/10.1007/s10989-019-09978-1
Peters MG, Farías E, Colombo L, Filmus J, Puricelli L, de Kier B, Joffé E (2003) Inhibition of invasion and metastasis by glypican-3 in a syngeneic breast cancer model. Breast Cancer Res Treat 80(2):221–232. https://doi.org/10.1023/a:1024549729256
Ramachandran GN (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99
Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin 69(1):7–34. https://doi.org/10.3322/caac.21551
Toretsky JA, Zitomersky NL, Eskenazi AE, Voigt RW, Strauch ED, Sun CC et al (2001) Glypican-3 expression in Wilms tumor and hepatoblastoma. J Pediatr Hematol Oncol 23(8):496–499
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108. https://doi.org/10.3322/caac.21262
Yang C, Tabatabaei SN, Ruan X, Hardy P (2017) The dual regulatory role of MiR-181a in breast cancer. Cell Physiol Biochem 44(3):843–856. https://doi.org/10.1159/000485351
Zou M, Li Y, Xia S, Chu Q, Xiao X, Qiu H et al (2017) Knockdown of CAVEOLIN-1 sensitizes human basal-like triple-negative breast cancer cells to radiation. Cell Physiol Biochem 44(2):778–791. https://doi.org/10.1159/000485291
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13721_2020_234_MOESM1_ESM.pptx
Supplementary Fig. 1. GPC3 analysis in breast cancer (Oncomine database). Box plot comparing specific GPC3 expression in normal breast (left) and cancerous breast tissue (right) generated from the Oncomine database. Curtis breast dataset (A-I): analysis is shown for (A) invasive ductal and invasive lobular breast carcinoma (90), (B) invasive ductal breast carcinoma, (C) invasive lobular breast carcinoma, (D) medullary breast carcinoma (32), (E) tubular breast carcinoma (67), (F) ductal breast carcinoma in-situ (10), (G) mucinous breast carcinoma (46), (H) invasive breast carcinoma (21), and (I) breast carcinoma (14); TCGA breast dataset analysis (J-N): analysis is shown for (J) mixed lobular and ductal breast carcinoma (7), (K) invasive ductal and lobular carcinoma (3), (L) invasive breast carcinoma (76), (M) invasive ductal breast carcinoma (389), (N) invasive lobular breast carcinoma (36); Finak breast dataset, analysis is shown for (O) invasive breast carcinoma (53); Karnoub breast dataset, analysis is shown for (P) invasive ductal breast carcinoma (7); and Richardson breast-2 dataset, analysis is shown for (Q) ductal breast carcinoma (40). IDILBC- invasive ductal and invasive lobular breast carcinoma, IDBC- invasive ductal breast carcinoma, ILBC- invasive lobular breast carcinoma, MBC- medullary breast carcinoma, TBC- tubular breast carcinoma, DBCIS- ductal breast carcinoma in-situ, IBC- invasive breast carcinoma, BC- breast carcinoma, MLDBC- mixed lobular and ductal breast carcinoma, IDLC- invasive ductal and lobular carcinoma, DBC- ductal breast carcinoma, TNBC- triple negative breast cancer. (PPTX 174 kb)
13721_2020_234_MOESM2_ESM.pptx
Supplementary Fig. 2. (A) Heat map of GPC3 expression and DNA methylation status; (B) GPC3 expression in different breast cancer DNA methylation clusters. Results generated using the UCSC Xena browser based on the data in TCGA. (PPTX 76 kb)
Rights and permissions
About this article
Cite this article
Barman, U.D., Saha, S.K., Kader, M.A. et al. Clinicopathological and prognostic significance of GPC3 in human breast cancer and its 3D structure prediction. Netw Model Anal Health Inform Bioinforma 9, 24 (2020). https://doi.org/10.1007/s13721-020-00234-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13721-020-00234-x