Abstract
Key message
Genetic diversity appears to be unaffected by disturbance in a stand of the light-demanding Neotropical tree V. michelii. Although spatial genetic structure is modified in post-disturbance cohorts, mixing of seeds from different mother trees in canopy gaps appears to efficiently maintain genetic admixture.
Context
The interplay between genetic and demographic processes has major consequences on population viability. Population size affects demographic trends, while genetic diversity insures viability by reducing risks of inbreeding depression and by maintaining adaptive potential. Yet, the consequences of increases in census size (as opposed to effective size) on genetic diversity of forest populations are poorly known.
Aims
We have studied the structure of genetic diversity in populations of saplings of the light-responsive tree, Virola michelii (Myristicaceae, the nutmeg family), in two plots having undergone different levels of canopy-gap opening disturbance. This allowed us to test the “intermediate disturbance” hypothesis, which generally applies to species diversity, at the intra-specific scale.
Methods
Levels and distribution of genetic diversity were compared between plots and between life stages. Sapling parentage was analysed to infer each adult tree’s contribution to regeneration.
Results
Genetic diversity was higher, and spatial genetic structure was stronger in the post-disturbance than in the control seedling population. Parentage analysis suggested that a limited number of parents contributed to most of the regeneration, but that efficient mixing of their progeny may have enhanced the diversity of saplings occupying canopy gaps.
Conclusion
A mixture of demo-genetic processes may contribute to maintain genetic diversity in spite of, or possibly due to, ecosystem disturbance in V. michelii.
Similar content being viewed by others
1 Introduction
Regeneration dynamics influence the ability of plant species and populations to maintain sustainable levels of genetic diversity. Although a minority of the saplings that grow in a forest will eventually become fertile adults, sapling cohorts are the stock those adults are drawn from; the genetic structure of future adult populations has its foundations in the genetic diversity of present-day juvenile populations. In highly diverse ecosystems such as tropical forests, where a given species’ population density can be extremely low, the study of new generations’ genetic diversity and of parental population effective size is therefore of great importance.
Forest communities are dynamic ecosystems that undergo recurrent perturbation events. The structure of a forest at any point in time is a mixture of ecological successions, including both old-growth cohorts and forest patches recovering from recent disturbances (Connell 1978; Hubbell 1999). Such non-equilibrium processes are considered as one of the causes of the high diversity of some ecosystems, as summarised by the “intermediate disturbance hypothesis” (IDH; Connell 1978). The IDH has been the recurrent subject of controversy (Fox 2013a, b; Sheil and Burslem 2013), particularly from the theoretical point of view. Nevertheless, as far as empirical evidence on highly diverse communities of sessile organisms is concerned, IDH has been tested, and validated, both observationally and experimentally in a variety of case studies (Shea et al. 2004); in particular, a study (Molino and Sabatier 2001) found it to hold in the case of an artificial disturbance experiment carried out in the tropical forests of the Guiana shield. Although this topic is well described at the species level, the IDH has not been tested at the infra-specific level—that is, we do not know the effects of intermediate disturbance on levels and structure of genetic diversity. We therefore set out to test the effect of intermediate disturbance in two species: Virola michelii, which is the subject of the present paper, and Jacaranda copaia (Leclerc et al., companion paper under revision). We did so by studying populations of the two species in the very same plots studied, at the community level, by Molino and Sabatier (2001) and by Baraloto et al. (2012). In the latter two studies, carried out at the Paracou (French Guiana) forest inventory station (Gourlet-Fleury et al. 2004a, b), the forest community was found to respond to intermediate disturbance with increased species and functional diversification. For J. copaia (Leclerc et al., under revision), we found that genetic diversity tended to be reduced in the post-disturbance regeneration, relative to undisturbed areas. For V. michelii, a previous study on the same populations (Scotti et al. 2013) indicated that diversity in post-disturbance cohorts was the same as, or marginally higher than, in undisturbed cohorts, notwithstanding the fact that undisturbed cohorts are the outcome of continued realised dispersal occurring over decades, while post-disturbance cohorts are mostly constituted by seedlings having germinated immediately after disturbance. Here, we investigate the spatial distribution of genetic diversity, as a means to better describe the impact of disturbance on demo-genetic processes. Spatial Genetic Structure (SGS) can be influenced by disturbances and their effects on regeneration processes (Banks et al. 2013). Sezen et al. (2005, 2007), by comparing adult and sub-adult populations in a secondary forest and in an adjacent old-growth forest, found patches of similar genotypes to be larger in the secondary growth. Ueno et al. (2006) showed that sapling populations growing under forest cover have stronger SGS than those in gaps. In a companion paper (Leclerc et al. under revision), we have shown that SGS extends over longer distances after disturbance than in undisturbed cohorts. In the study of the effects of disturbance, it is therefore sensible to assess not only changes in levels of diversity but also modifications of its spatial organisation.
The characteristics of a species’ reproductive phenology and seed dispersal can determine the effect of regeneration events on genetic diversity and contribute to determine how disturbance affects SGS. Patches of closely related individuals can derive from seed dispersal from a limited number of parent trees (Aldrich and Hamrick 1998; Sezen et al. 2005; Davies et al. 2010). Conversely, dispersal agents such as birds and mammals can actively compensate local losses of diversity by actively moving groups of seeds across large distances, so that in some cases local potential parents are under-represented in regeneration after disturbance (Aldrich and Hamrick 1998; Aldrich et al. 1998). Parentage analyses can provide useful information about the influence of fecundity and dispersal processes on genetic diversity patterns in the following generation.
How does disturbance affect dispersal and regeneration, and ultimately genetic diversity? We have addressed this question by building on the data and results presented in Scotti et al. (2013) and by combining SGS and parentage analyses. The results of the present study are discussed in the light of predictions provided by the IDH and of results obtained in the same experimental system at the community and species level.
2 Materials and methods
2.1 Study system
V. michelii has been classified either as a light-demanding (Delcamp et al. 2008) or as a shade-tolerant insect-pollinated, vertebrate-dispersed dioecious canopy rain forest tree species that preferentially grows on terra firme soils. V. michelii has a short seed dormancy of 4–6 weeks, during which embryos develop prior to germination (Forget 1991). The species is common in the Guiana Shield, and its seedlings can rapidly colonise the understory and forest canopy gaps. Forget (1991) has shown that survival of juveniles is greater in gaps than understory and defined the species as dependent on small canopy gaps. However, seedlings can colonise the understory and survive until a canopy gap favouring their growth eventually appears (Forget 1991). V. michelii is common in secondary forests, and it thrives in disturbed forest sites where conditions are favourable for early colonising species.
2.2 Sampling site
The dynamics of V. michelii regeneration was studied at the Paracou experimental site (French Guiana; 5°18′N, 52°55′W). The site has a typical equatorial climate, with mean annual rainfall of about 3000 mm, two rainy seasons and two dry seasons; an elevation between 0 and 100 m a.s.l., a rugged landscape comprised of an alternation of small hills, creeks and seasonally flooded bottomlands; and an alternation of ferralitic, shale and quartzite soils, with rather low fertility (Gourlet-Fleury et al. 2004b). The Paracou forest inventory site is the subject of recurrent censuses since 1984, and the availability of detailed pre- and post-disturbance forest inventory data and of forest disturbance maps makes it a unique tool for the study of regeneration processes. V. michelii adult populations show intermediate levels of spatial genetic structure in the study area (Degen et al. 2001; Hardy et al. 2006). The study was carried out in two contiguous 6.25 ha plots (plots P9 and P11, Fig. 1), separated by a 50-m-wide buffer zone. Plot P11 is a control plot without any logging, while plot P9 underwent experimental logging between 1986 and 1988: a traditional selective felling for timber, representing about 10 trees/ha of diameter at breast height (d.b.h.) >50–60 cm, belonging to commercial species. This treatment induced several canopy gaps that favoured regeneration of light-responding species. The two plots have already been included in the study of the effect of light-gap disturbance on community diversity, described in Molino and Sabatier (2001) and Baraloto et al. (2012), and on genetic diversity in J. copaia (Leclerc et al., submitted to Annals of Forest Science). The two plots are similar in soil conditions, topography and flora; most of their surface is composed of terra firme plateaus (Molino and Sabatier 2001), which is V. michelii’s habitat, occupying flat plateaus. While this setup does not include replicates (the treatment has been applied to only one site, and we have only used one treatment and one control plot), the two plots have been selected because they lie next to each other and are large enough to avoid biases caused by local variation in stem density (Molino and Sabatier 2001).
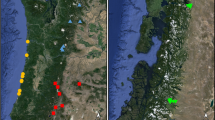
Maps of the study site and sampling. a Position of the two study plots (P9, P11) in the larger context of the Paracou forest inventory site; b positions of the sampled individuals: large dots, adult trees; small dots, saplings; grey areas represent canopy gaps as detected at the time of the experimental perturbation
2.3 Sampling and molecular methods
The samples and the molecular data are the same as in Scotti et al. (2013) (Electronic Supplementary Material 1). In summary, all saplings in the study area (524 in total, of which 312 in the treatment plot (P9) and 212 in the control plot (P11); Fig. 1), plus 11 adult trees in the two plots and 66 in the remaining Paracou site plots (Fig. 1), were characterised at 687 AFLP markers. For all the analyses, the subset of 163 AFLP markers that minimise linkage disequilibrium (Scotti et al. 2013) were used; samples with a low genotyping quality were excluded from further analyses.
2.4 Spatial genetic structure
Spatial autocorrelation of kinship was estimated based on the “Kinship” statistic (Eqs. (1) and (15) in Hardy 2003). Calculations were performed by SPAGeDi (Hardy and Vekemans 2002) version 1.3 with the following maximum distances for distance classes: 20, 40, 60, 80, 100, 200 and 300 m (intervals were defined more densely at short distances to analyse relatedness in sapling clumps). Three independent analyses were executed for the treatment and the control plots with inbreeding coefficient F set to −0.1, 0 and 0.1 to take unknown mating system parameters into account, based on independent inbreeding estimates obtained on codominant markers (Montaigne et al., under revision). The whole sapling population was used as reference in the calculation of relative estimates of kinship coefficients. Parameters b and Sp (Vekemans and Hardy 2004) were computed. The method developed by Smouse et al. (2008), as implemented in GenAlEx 6.501 (Peakall and Smouse 2012), was used to test differences between autocorrelograms obtained from different populations.
2.5 Genetic clustering
Population genetic structure was assessed by automated Bayesian assignment using BAPS (Corander et al. 2006, 2008a, b) at the individual level. The algorithm was allowed to pick the most likely K (number of clusters), with 1 ≤ K ≤ 20 and 20 runs for each K. Bayesian clustering was performed with and without prior spatial information, while keeping all the remaining settings equal.
2.6 Parentage analyses
Parentage analyses were performed to identify the relative contribution of individual trees to regeneration, and to assess the effect of dispersal on the genetic diversity in the control and in the treatment plot. All sexed and not sexed trees with diameter at breast height >20 cm at the date of controlled logging were used as potential parents (direct observations performed in 2005 on the whole Paracou inventory site showed that the absolute minimum flowering diameter was 15 cm for male trees and 20 cm for females; Toebosch, unpublished). Parentage analyses were performed using FaMoz (Gerber et al. 2003), with typing error rate set to 0.001 for both limit of detection (LOD) calculation and simulations, the same three levels of inbreeding coefficient F (−0.1, 0, 0.1) used for the spatial autocorrelation of kinship estimates, and using all samples, including those with missing data. Simulations were first performed to assess the power and accuracy of the test, and then parentage analyses were run on the empirical sapling dataset to identify the most likely parent pair for each sapling. A parent pair was assigned to a seedling when the ΔLOD relative to the second most likely parent pair was higher than 3.0. Parent pairs with incompatible sexes (two males or two females) were not retained; unsexed trees were always retained as parent when significant. Parent pair analyses were preferred over single-parent analyses because parent identification was unclear in the latter case (i.e. ΔLODs were generally small), although it provided a larger number of potential parents.
3 Results
Spatial autocorrelation analyses were run for three values of the inbreeding coefficient F. Given that no difference was observed between inbreeding levels, only the results for F = 0 are shown here. Large differences could be observed between the autocorrelograms of the treatment and the control plot (Fig. 2): the treatment plot (Fig. 2a) showed positive autocorrelation up to around 80 m and negative farther than 100 m, and a generally monotonically declining curve, with a highly significant slope (log-log b = −0.02, P value <0.0001; Sp = 0.028). The control plot (Fig. 2b) also showed a globally negative slope (log-log b = −0.01, P value <0.0001; Sp = 0.013), but with positive kinship values across the range, a convex curve, above-average significant values within 50 m, under-average significant values between 100 and 150 m, and non-significant values beyond that distance. Differences between the two autocorrelograms were significant (Omega = 40.7, P = 0.01)
Spatial autocorrelation for kinship in sapling cohorts in a the treatment plot (P9) and b the control plot (P11). Distance—mean distance for pairs within each distance class. Dashed lines represent the upper (97.5 %) and lower (2.5 %) bounds of the neutral distribution as obtained by 1000 permutations
To visualise the spatial structure underlying these patterns, an automated Bayesian analysis was run to infer clusters of genetically related saplings. Since the results obtained with and without prior spatial information were qualitatively similar, only the analysis with prior spatial information is described here. The most likely partition (P = 0.94) involved 14 clusters (Electronic Supplementary Material 2). Individuals belonging to the same cluster were visually aggregated. The pattern appeared to be similar for the treatment and the control plot, but the number of groups occurring in the two plots differed: 12 clusters out of 14 occurred in the treatment population (plot P9), while only 8 appeared in the control population (plot P11). Two clusters were restricted to plot P11 (one of which made of a single individual), and six to plot P9 (one of which made of a single individual).
Parentage analyses were performed to identify patterns of seed dispersal. Simulation tests showed that, conditional on sampled parental genotypes, parenthood was correctly identified (i.e. the most likely parents and parent pairs were the correct ones) for 88 to 100 % of the simulated progeny arrays (out of 5000 simulated offspring genotypes and over three values of inbreeding coefficient F (−0.1, 0, 0.1)). A parent pair was identified among the 77 genotyped mature trees for 217 saplings out of 383 (57 %). At least one parent was found within the study plots for 57 saplings out of 383 (15 %), and both parents were found within the plots for one of them. Of these 57 saplings, 47 had their second parent in the neighbourhood of the study plots (<300 m apart); 110 additional saplings had at least one parent in the same neighbourhood. Twenty-seven putatively mature trees, out of the 77 trees sampled, appeared in identified parent pairs. Six trees of both sexes contributed individually more than 5 % of the offspring and 84 % cumulatively (Table 1). The best contributor (which could not be sexed; 27 % of the total parenthood) was in the neighbourhood of the study area, while the only contributor tree inside the study area was a female having contributed 14 % of the saplings (Table 1). Seedlings derived from all six parents were found scattered throughout the study area (Electronic Supplementary Material 3), with no particular trend for clumped distributions. Nevertheless, Bayesian clusters, which are non-randomly distributed (see above), had partially non-overlapping sets of potential parents. Differences in the composition of the putative parent pool were inspected for clusters for which at least 20 parents could be identified (Electronic Supplementary Material 4, Table 1 and Fig. 3). The proportions of parental contributions to the largest cluster (cluster 13) were not statistically different from those of the remaining five clusters included in this analysis, while the proportions of all other clusters statistically differed from each other at the 5 % statistical threshold (Table 1).
Graphical representation of the contribution of each potential parent tree within each cluster, relative to the total number of parents identified for each cluster. Grey scale corresponds to contributions from 0 % (white boxes) to 50 % (black boxes). Clusters are sorted from top to bottom by decreasing number of potential parents identified. Tree names are given under the form of Paracou inventory codes
4 Discussion
In a previous paper (Scotti et al. 2013), we have shown that diversity in the post-disturbance (treatment) plot was higher than in the control plot. The results presented here concur to indicate that regeneration processes occurring after disturbance, and in particular after the opening of canopy gaps, may influence the genetic structure of populations in V. michelii. Indeed, differences in diversity levels between post-disturbance and control juvenile populations are matched by marked differences in spatial genetic structure and levels of genetic relatedness. Variation in relative relatedness is much larger in the post-disturbance (treatment) plot, with relative kinship as high as 0.07 at short distances, falling to negative values at distances beyond 150 m. On the contrary, the control plot is characterised by a relatively flatter trend, with low but strictly positive kinship coefficients at all distances and a lower Sp value (0.13 for the control plot vs 0.028 for the treatment plot). These differences suggest that the post-disturbance (treatment) plot is occupied by denser clumps of related saplings than the control plot; this is in contrast with other cases reported in the literature. The spatial distribution of Bayesian clusters provided a graphical illustration of these differences between sub-populations. Both plots are occupied by geographically clustered genetic groups. Fourteen genetic clusters have been identified (although isolation-by-distance structures may introduce biases in the results of Bayesian assignment methods (Guillot and Santos 2009), spatial structure shows relatively weak kinship coefficients, and should not have a large impact on the outcome of Bayesian assignment). Out of 14 clusters, 6 are common to both plots, 2 are restricted to the control plot and 6 more are present only in the post-disturbance (treatment) plot. These six groups represent 31 % of all juvenile stems (55 % of stems in plot P9) and may explain both the strength of the observed structure and the increased diversity in plot P9 relative to plot P11. In this interpretation, common clusters would represent the “background” genetic pool, undergoing turnover that happens in control conditions, while additional, genetically different sapling clumps appear as a consequence of the increased availability of favourable space after disturbance.
The large imbalance of parental contributions, indicated by parentage analysis, suggests that spatial structure is generated by the non-random realised dispersal of clumps of related seeds in areas favouring regeneration, as already observed in another Virola species (Russo and Augspurger 2004). These seeds may be derived from few trees having experienced larger seed output at the time of canopy opening (Jones and Hubbell 2006). The results presented here describe a picture of overall mixing of genotypes at the landscape level, with a slight majority (56 %) of seeds having no parent in the study area or in neighbouring plots (a relevant portion of the saplings, whose parents are not accounted for, must nevertheless be the progeny of trees located in the forest surrounding the focal plots, and not included in our sampling). Parents located at distances larger than 1 km from the study plots account for 50 % of the identified parental contribution: 7 % from females, 11 % from males, and 32 % from trees with undetermined sex.
Distant realised dispersal and strong spatial genetic structure seem to be contradictory, unless groups of related seeds are dispersed together over long distances: dispersers (most likely large birds and monkeys) may prefer canopy gaps as resting areas, fetching fruits from distant trees to the disturbed areas, thus increasing diversity in post-disturbance populations (Karubian et al. 2010). This form of kin-structured dispersal has been reported previously (Torimaru et al. 2007; García and Grivet 2011) and may actually be responsible for the clumped distribution of genetic clusters. Pollinators may also contribute to clumped genotype distribution, but it is unlikely that single pollinators carry large quantities of pollen from one tree to several distant trees, because virola flowers are small (2–3 mm), and therefore pollinators should be correspondingly small. Realised dispersal distances observed here are slightly larger than the seed dispersal distances previously observed in the genus (Howe 1981; Howe and Richter 1982; Russo and Augspurger 2004; Holbrook and Loiselle 2007, 2009). In V. michelii, direct observations reported seed dispersal up to 60 m in the undisturbed plot P11 (Forget et al. 2000). If we can consider seed dispersal distance as the background reference value in the absence of disturbance, the difference with the values we have observed supports the idea of increased mixing in the post-disturbance conditions (keep in mind, though, that we deal with indirect estimates of realised dispersal distance, not direct measures of seed dispersal; but see Oddou-Muratorio et al. (2010) for a discussion about comparisons between direct and indirect methods to estimate seed dispersal). By providing more diverse habitats than the understory of closed canopy, disturbances—and in particular light gaps—may further increase genetic diversity by favouring a more diverse array of genotypes (Ueno et al. 2006); alternatively, the large amount of light available in gaps may release the growing cohorts from selection at least at early life stages, also increasing diversity.
Overall, the pattern of genetic diversity observed in virola matches the pattern of increased species diversity after disturbance shown, at the community and functional level, in the very same plots respectively by Molino and Sabatier (2001) and Baraloto et al. (2012). Thus, the same mechanisms driving community diversity may be acting as well at the within-species level for V. michelii. Nevertheless, it must be recalled that genetic diversity in the adult population is higher than in juveniles from both plots (Scotti et al. 2013), suggesting that imbalances in the genetic composition of the juvenile population is somehow compensated during the phases of demographic reduction that lead to the adult cohort, possibly due to density-dependent thinning mechanisms (Choo et al. 2012); moreover, the current adult population is likely to be composed of trees belonging to multiple cohorts, likely increasing their diversity. The patterns shown by virola regeneration indicate that, notwithstanding the very unequal contribution of parental trees to large juvenile populations, genetic diversity may be maintained by regeneration bursts, and it can be expected that present-day saplings will give rise to adult stands as diversified as the current ones. This seems to contrast with patterns observed in pioneer or secondary-forest trees and palms but is consistent with genetic admixture in pasturelands in primary forest species, and contrasts with the results obtained, in the very same experimental site and setup, on J. copaia (Leclerc et al. submitted). Globally, species with the same ecological and dispersal characteristics as virola may take advantage from intermediate-level disturbance. Clearly, this cannot be generalised to all species and all kinds of perturbation, and the lack of replication imposed upon our analysis by the original setup of the controlled disturbance constrains the possibility to more rigorously test hypotheses; nevertheless, this case study shows that at least some tropical rainforest species can positively respond to, and even benefit from, events with moderate ecological impact. Despite the lack of a highly disturbed treatment, our results are a good indication that intermediate disturbance hypothesis (Grime 1973; Connell 1978; Molino and Sabatier 2001) may hold not only for species diversity but also for the maintenance of (neutral) genetic diversity.
References
Aldrich PR, Hamrick JL (1998) Reproductive dominance of pasture trees in a fragmented tropical forest mosaic. Science 281:103–105
Aldrich PR, Hamrick JL, Chavarriaga P, Kochert G (1998) Microsatellite analysis of demographic genetic structure in fragmented populations of the tropical tree Symphonia globulifera. Mol Ecol 7:933–944
Banks SC, Cary GJ, Smith AL et al (2013) How does ecological disturbance influence genetic diversity? Trends Ecol Evol 28:670–679
Baraloto C, Hérault B, Paine CET et al (2012) Contrasting taxonomic and functional responses of a tropical tree community to selective logging. J Appl Ecol 49:861–870
Choo J, Juenger TE, Simpson BB (2012) Consequences of frugivore-mediated seed dispersal for the spatial and genetic structures of a neotropical palm. Mol Ecol 21:1019–1031
Connell JH (1978) Diversity in tropical rain forests and coral reefs. Science 199:1302–1310
Corander J, Marttinen P, Mäntyniemi S (2006) Bayesian identification of stock mixtures from molecular marker data. Fish Bull 104:550–558
Corander J, Marttinen P, Siren J, Tang J (2008a) Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinforma 9:539
Corander J, Sirén J, Arjas E (2008b) Bayesian spatial modeling of genetic population structure. Comput Stat 23:111–129
Davies SJ, Cavers S, Finegan B, Navarro C, Lowe AJ (2010) Genetic consequences of multigenerational and landscape colonisation bottlenecks for a Neotropical forest pioneer tree, Vochysia ferruginea. Trop Plant Biol 3:14–27
Degen B, Caron H, Bandou E et al (2001) Fine-scale spatial genetic structure of eight tropical tree species as analysed by RAPDs. Heredity 87:497–507
Delcamp M, Gourlet-Fleury S, Flores O, Gamier E (2008) Can functional classification of tropical trees predict population dynamics after disturbance? J Veg Sci 19:209–220
Forget PM (1991) Comparative recruitment patterns of two non-pioneer canopy tree species in French Guiana. Oecologia 85:434–439
Forget P-M, Milleron T, Feer F, Henry O, Dubost G (2000) Effects of dispersal pattern and mammalian herbivores on seedling recruitment for Virola michelii (Myristicaceae) in French Guiana. Biotropica 32:452–462
Fox JW (2013a) The intermediate disturbance hypothesis should be abandoned. Trends Ecol Evol 28:86–92
Fox JW (2013b) The intermediate disturbance hypothesis is broadly defined, substantive issues are key: a reply to Sheil and Burslem. Trends Ecol Evol 28:572–573
García C, Grivet D (2011) Molecular insights into seed dispersal mutualisms driving plant population recruitment. Acta Oecol 37:632–640
Gerber S, Chabrier P, Kremer A (2003) FaMoZ: a software for parentage analysis using dominant, codominant and uniparentally inherited markers. Mol Ecol Notes 3:479–481
Gourlet-Fleury S, Favrichon V, Schmitt L, Petronelli P (2004a) Consequences of silvicultural treatments on stand dynamics at Paracou. In: Gourlet-Fleury S, Guehl JM, Laroussinie O (eds) Ecology & management of a neotropical rainforest. Lessons drawn from Paracou, a long-term experimental research site in French Guiana. Elsevier, Paris, pp 254–280
Gourlet-Fleury S, Guehl J-M, Laroussinie O (2004b) Ecology & management of a neotropical rainforest. Lessons drawn from Paracou, a long-term experimental research site in French Guiana. Elsevier, Paris
Grime JP (1973) Competitive exclusion in herbaceous vegetation. Nature 242:344–347
Guillot G, Santos F (2009) A computer program to simulate multilocus genotype data with spatially autocorrelated allele frequencies. Mol Ecol Resour 9:1112–1120
Hardy OJ (2003) Estimation of pairwise relatedness between individuals and characterization of isolation-by-distance processes using dominant genetic markers. Mol Ecol 12:1577–1588
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hardy OJ, Maggia L, Bandou E et al (2006) Fine-scale genetic structure and gene dispersal inferences in 10 Neotropical tree species. Mol Ecol 15:559–571
Holbrook KM, Loiselle BA (2007) Using toucan-generated dispersal models to estimate seed dispersal in Amazonia Ecuador. In: Dennis AJ, Schupp EW, Green EW, Westcott DA (eds) Seed dispersal: theory and its application in a changing world. CAB International, Wallingford
Holbrook KM, Loiselle BA (2009) Dispersal in a Neotropical tree, Virola flexuosa (Myristicaceae): does hunting of large vertebrates limit seed removal? Ecology 90:1449–1455
Howe F (1981) Dispersal of a Neotropical nutmeg (Virola sebifera) by birds. Auk 98:88–98
Howe HF, Richter WM (1982) Effects of seed size on seedling size in Virola surinamensis; a within and between tree analysis. Oecologia 53:347–351
Hubbell SP (1999) Light-gap disturbances, recruitment limitation, and tree diversity in a neotropical forest. Science 283:554–557
Jones FA, Hubbell SP (2006) Demographic spatial genetic structure of the Neotropical tree, Jacaranda copaia. Mol Ecol 15:3205–3217
Karubian J, Sork VL, Roorda T, DurÃEs R, Smith TB (2010) Destination-based seed dispersal homogenizes genetic structure of a tropical palm. Mol Ecol 19:1745–1753
Molino J-F, Sabatier D (2001) Tree diversity in tropical rain forests: a validation of the intermediate disturbance hypothesis. Science 294:1702–1704
Oddou-Muratorio S, Bontemps A, Klein EK et al (2010) Comparison of direct and indirect genetic methods for estimating seed and pollen dispersal in Fagus sylvatica and Fagus crenata. For Ecol Manag 259:2151–2159
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics (Oxford, England) 28:2537–2539
Russo SE, Augspurger CK (2004) Aggregated seed dispersal by spider monkeys limits recruitment to clumped patterns in Virola calophylla. Ecol Lett 7:1058–1067
Scotti I, Montaigne W, Cseke K, Traissac S (2013) RaBoT: a rarefaction-by-bootstrap method to compare genome-wide levels of genetic diversity. Ann For Sci 70:631–635
Sezen UU, Chazdon RL, Holsinger KE (2005) Genetic consequences of tropical second-growth forest regeneration. Science 307:891
Sezen UU, Chazdon RL, Holsinger KE (2007) Multigenerational genetic analysis of tropical secondary regeneration in a canopy palm. Ecology 88:3065–3075
Shea K, Roxburgh SH, Rauschert ESJ (2004) Moving from pattern to process: coexistence mechanisms under intermediate disturbance regimes. Ecol Lett 7:491–508
Sheil D, Burslem DFRP (2013) Defining and defending Connell’s intermediate disturbance hypothesis: a response to Fox. Trends Ecol Evol 28:571–572
Smouse PE, Peakall R, Gonzales E (2008) A heterogeneity test for fine-scale genetic structure. Mol Ecol 17:3389–3400
Torimaru T, Tani N, Tsumura Y, Nishimura N, Tomaru N (2007) Effects of kin-structured seed dispersal on the genetic structure of the clonal dioecious shrub Ilex leucoclada. Evol Int J Org Evol 61:1289–1300
Ueno S, Tomaru N, Yoshimaru H, Manabe T, Yamamoto S (2006) Effects of canopy gaps on the genetic structure of Camellia japonica saplings in a Japanese old-growth evergreen forest. Heredity 96:304–310
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–935
Acknowledgments
The authors wish to thank Mathieu Toebosch for sexing adult trees, Saint-Omer Cazal for plant material collection and DNA extractions, and Audrey Jolivot and Jean-Yves Goret for producing plot maps. We wish to thank Eric Marcon, Caroline Scotti-Saintagne, Rosane G. Collevatti, Andrea Piotti and Pierre-Michel Forget for critically reading the manuscript.
Funding
The research was funded by the “ECOFOR—Biodiversité et gestion forestière” program. IS is currently supported by an “Investissement d’Avenir” grant managed by Agence Nationale de la Recherche (CEBA, ref. ANR-10-LABX-0025).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Ricardo Alia
Contribution of the co-authors
IS conceived the study and ran data analyses; WM and CS collected and analysed data; ST analysed data; all co-authors wrote the manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Electronic Supplementary Material 1
Sampling and molecular methods. (DOCX 14 kb)
Electronic Supplementary Material 2
Voronoi tessellation of individuals, colour-coded according to genetic clusters identified by automated Bayesian assignment. (GIF 72 kb)
Electronic Supplementary Material 3
Distribution of seedlings derived from parent trees listed in Table 1. Dots: seedlings; circles: parental trees (parental tree names are displayed next to their position; only the two trees closer than 1 Km apart from the study areas are shown). x and y axes show coordinates in the UTM system. (a) 10010089; (b) 5030500; (c) 3030733; (d) 9010869; (e) 16110208; (f) 16060457. (GIF 39 kb)
Electronic Supplementary Material 4
Table of the contribution of each potential parent tree within each cluster, relative to the total number of parents identified for each cluster. Tree names are given under the form of Paracou inventory codes. (DOCX 12 kb)
Rights and permissions
About this article
Cite this article
Scotti, I., Montaigne, W., Cseke, K. et al. Life after disturbance (II): the intermediate disturbance hypothesis explains genetic variation in forest gaps dominated by Virola michelii Heckel (Myristicaceae). Annals of Forest Science 72, 1035–1042 (2015). https://doi.org/10.1007/s13595-015-0508-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13595-015-0508-3