Abstract
Global demand for pesticide-free food products is increasing rapidly. Crops of all types are, however, under constant threat from various plant pathogens. To achieve adequate control with minimal pesticide use, close monitoring is imperative. Many plant pathogens spread through the air, so the atmosphere is composed of a wide variety of plant pathogenic and non-plant pathogenic organisms, in particular in agricultural environments. Aerobiology is the science that studies airborne microorganisms and their distribution, especially as agents of infection. Although this discipline has existed for decades, the development of new molecular technologies is contributing to an increase in the use of aerobiological data for several purposes, from day-to-day monitoring to improving our understanding of pathosystems. Although the importance of knowing the size and composition of plant pathogen populations present in the air is recognized, technical constraints hinder the development of agricultural aerobiology. Here we review the application of spore sampling systems in agriculture and discuss the main considerations underlying the implementation of airborne inoculum monitoring. The results of this literature review confirm that the use of aerobiological data to study the escape of inoculum from a source and its role in the development of diseases is well mastered, but point at a lack of knowledge to proceed with the deployment of these systems at the landscape scale. Thus, we conclude that airborne inoculum surveillance networks are still in their early stages and although more and more initiatives are emerging, research must be conducted primarily to integrate evolving technologies and improve the access, analysis, interpretation and sharing of data. These tools are needed to estimate short- and medium-term risks, identify the most appropriate control measures with the lowest environmental risk, develop indicators to document the effects of climate change, and monitor the evolution of new genotypes at multiple scales.
Similar content being viewed by others
Contents
-
1. Introduction
-
11. Perspectives
-
Acknowledgements
-
References
1 Introduction
Air is the means of transport for many plant pathogens, some of which can travel thousands of kilometers while maintaining their viability and ability to cause new epidemics (Stakman and Christensen 1946). Many plant pathogenic fungi are remarkably well adapted to airborne spread. The fact that fungal spores are spread by air currents has been known for almost as long as the existence of the spores themselves. In 1729, Micheli published the results of research on spore production and demonstrated that “clouds” of spores could be released into the air. The atmosphere in agricultural environments is therefore full of particles ranging in size from 0.1 μm for viruses to 100 μm for pollens. Monitoring spores and other particles in the air requires a specific field of scientific expertise, which is called aerobiology.
Aerobiology has developed to a large extent in response to the need to know the quality of air in buildings and of pollen responsible for allergies in humans. Philip Herries Gregory is the pioneer of aerobiology in agriculture. By the late 1930s, he was already studying the movement of fungal spores in the air, including those of Phytophthora infestans which is responsible for potato late blight (Gregory 1945, 1973). Dr. Gregory initiated the aerobiology research program at Rothamsted Research (formerly the Rothamsted Experimental Station) in England, which for decades has contributed and continues to contribute to the advancement of knowledge in aerobiology. Contributions includes the study of processes involved in the airborne spread of plant pathogens (Williams et al. 2001; McCartney et al. 1997), spores disseminated in aerosols (Fitt et al. 1985) and the use of information on airborne inoculum as risk indicators (Rogers et al. 2009; West et al. 2008). Since Dr. Gregory’s early work, the use of spore samplers for the study of airborne inoculum has continued to develop (Lacey 1996) for diseases that affect: cereal (Chamecki et al. 2012; Hellin et al. 2018; Hemmati et al. 2001; Morais et al. 2016; Paulitz et al. 1999), industrial crops (Bashan et al. 1991; Alderman 1993), field crops (Chawda and Rajasab 1994; Fitt et al. 1985; Reich et al. 2016), fruit crops (Charest et al. 2002; Carisse et al. 2009a; Aylor 1995) and vegetables (Carisse et al. 2005; Carisse et al. 2009b).
Although the importance of considering airborne inoculum seems obvious from a disease management perspective, spores of plant pathogenic fungi are very small, many without a distinctive color or shape, and therefore very difficult to identify and count. As a result, the difficulty of counting spores in air samples has long limited the use of information on airborne inoculum in the management of airborne diseases. Over the years, different counting methods have been assessed, but methods using polymerase chain reaction (PCR) have really made it possible to use spore samplers at large scale with standardized counts (Carisse et al. 2009b; Falacy et al. 2007; West et al. 2008). Aerobiological research conducted worldwide during the last two decades represent a turning point as they demonstrated the possibility of using information on airborne inoculum to improve disease management. Published original research articles (Carisse et al. 2012; Van der Heyden et al. 2012; Rogers et al. 2009; West et al. 2008), books (Aylor 2017; Gregory et al. 1959) and reviews (West and Kimber 2015; Mahaffee and Stoll 2016; West et al. 2008; Jackson and Bayliss 2011) covered theoretical and applied, general or specific aspects of aerobiology and airborne spore sampling. In this review, we wish not to repeat former publications, but to discuss the state of knowledge related to the implementation of networks of airborne fungal spore monitoring.
2 Purposes of sampling airborne fungal spores
The primary purpose of aerobiology is to improve our understanding of the epidemiology of airborne diseases at different temporal and spatial scales. Aerobiological data are used in a wide range of studies, several examples of which are provided in Table 1. Regardless of their nature, aerobiological data can be used to model disease development and to evaluate different types of disease forecasting systems (Fall et al. 2015b). In comparative epidemiology, airborne inoculum progress curves can be used to study the role of airborne inoculum on disease development in relation to climatic conditions, production systems, cultivars, control method, etc. (Carisse et al. 2014). In addition, when aerobiological data are combined with genetic analyzes, it become possible to study fungal community and ecology of different pathogen genotypes including those related to fungicide resistance or aggressiveness (Fraaije et al. 2005; Hellin et al. 2020; Nicolaisen et al. 2017).
Sampling air for fungal spores is also motivated by the purposes of biovigilance, surveillance and monitoring. In the case of biovigilance, it can be used to measure the impacts of climate change and production practices or systems on populations of plant pathogens and beneficial agents, or to study the dispersion of emerging pathogens colonizing new agricultural environments, the emergence of new species or the genetic diversity of plant pathogens (Chen et al. 2018). With a purpose of surveillance, air is sampled in a particular region to measure a disease risk for plant pathogens that are present but cause sporadic or variable levels of damage. Also, it is important to monitor periodically introduced plant pathogens originating from areas further away, to detect their arrival in a specific area and react effectively. Potato late blight (P. infestans) and cucumber downy mildew (Pseudoperonospora cubensis) are examples of this type of plant diseases (Cohen and Rotem 1971; Rahman et al. 2020; Fall et al. 2015). Surveillance typically takes place over the long term, ideally at the same sites to facilitate year-to-year comparison. For endemic plant pathogens and for those that migrate via air, transplants, or seeds, a key element of disease management is the ability to determine when secondary inoculum is produced and how much is produced. In the case of monitoring, air sampling is combined with agronomic and meteorological data, to identify risks and optimize phytosanitary measures (Carisse et al. 2012; Van der Heyden et al. 2012). Airborne inoculum monitoring activities typically take place at a high frequency and at the spatial scale of the field, farm, or region, in order to allow day-to-day disease management decision making. Aerobiological data can therefore be used to improve forecasting systems and optimize within seasonal decision-making regarding fungicide applications.
3 Trajectory of fungal spores in the air
As a scientific discipline, aerobiology requires a good understanding of the phenomena involved, namely the production of inoculum (spores), the detachment of spores from their production site, their exit from the plant canopy, their dispersion (transport) in the air over long or short distances, the viability of the spores, and their deposition at new infection sites. The primary inoculum source varies depending on the plant pathogenic fungus and the production system. The sources may be seeds, transplants, volunteer plants, crop or warehouse debris, survival structures (e.g., sclerotia, cleistothecia, oospores), or resting growth structures (mycelium). Depending on the primary inoculum source, the first inoculum production sites are distributed over larger or smaller areas or confined to specific locations with a random or aggregated spatial distribution. For secondary inoculum, the number of production sites generally increases with disease progression.
Plant pathogenic fungi have different spore-release mechanisms, passive or active (Ingold 1971). For many fungal pathogens, spores are released only when they are shaken, splashed, or blown from their mycelia or sporophores; sometimes removed by insects. These spores released mechanisms are considered as passive. For example, most conidia are dislodged from their conidiophores and are dispersed only when exposed to specific physical disturbance such as wind or rain droplets (Fitt et al. 1985). Hence, fungi with passive spore release mechanisms often sporulate on parts of plants exposed to the wind (Ingold 1999). Other fungi have developed active spore release mechanisms that allow spores to escape from the calm air zone regardless of wind exposure (Aylor 1995). Some spores are ejected from fruiting structures (e.g., pseudothecia or perithecia) as a result of increased pressure inside the structure. This pressure can be osmotic or caused by the hydrolysis of substances such as glycogen. For other fungi, the projection of the spores is obtained following the production of a droplet of exudate which suddenly changes position while moving towards the point of attachment of the spore, causing a tension which releases the spore. (Ingold 1971). Some spore release mechanisms are somehow between passive and active for example, spores released in response to the twisting of their sporophores as they desiccate in dry air. Regardless of the mechanism, the exit of spores from the plant cover (laminar zone of calm air) is essential for spores to escape, disperse in the turbulent zone and travel over short or long distances (Gregory 1973; Aylor 2017).
Once spores are released into the air, they disperse, and their concentration decreases as the distance from the release point increases (Aylor 2017; Gregory 1973). Spore dispersion within and outside the plant canopy is difficult to measure since air movement affects the release, dispersion, and deposition of spores at new infection sites (Legg 1983). In the context of airborne inoculum monitoring, spore concentration is generally measured at the exit from the plant canopy, during the dispersion (transport) or deposition phase, the latter being more often used when sampling is conducted for disease risk estimation.
Aerobiology also includes the study of the influence of weather conditions, landscape architecture, plant cover and cultural practices on all stages of trajectory of spores in the air, from spore production to spore deposition at new infection sites. Aerobiology is multidisciplinary in that knowledge of plant patholoy, disease epidemiology, ecology of plant pathogens, mycology, molecular biology, mathematics, statistics, and modelling is required to properly measure and interpret aerobiological data.
4 Types of samplers
Spores can be collected with different types of samplers, depending on the purpose of the study and each type of sampler has its advantages and disadvantages. A number of factors should be considered when choosing the most appropriate sampler: its efficiency, the sampling volume, the length of the sampling period, the amount of time the sampler can stay in the field, the ease of manipulating samples, the size of the particles to be sampled and the frequency of monitoring. The characteristics of each type of samplers were described in depth by West and Kimber (2015), so we only cover the basics in this section (Table 2). In addition, examples of samplers used in different studies are provided in relation to the pursued objectives (Table 1).
The first category of devices used to collect spores can be described as passive samplers. Their operation is based on the deposition of spores on an adhesive surface (i.e., petri dish, microscope slide), through a slide or a filter (West and Kimber 2015). The simplest example consists of installing microscope slides coated with an adhesive substance (e.g., silicone grease, petroleum jelly) on a vertical, horizontal or inclined support (Fig. 1A). The obvious advantages of this type of device are its low cost and the direct mounting of the slides on a microscope. However, it does not allow the calculation of spore concentrations in the air (i.e., numbers of spores per m3 of air) because the amount of air sampled is unknown. Moreover, the reading of the microscope slides can be difficult and time consuming depending on the thickness of the silicone layer used and the amount of debris present on the sampling surface. However, this type of device can be useful for monitoring in areas that are large or difficult to access, where resources are limited, or if only the presence or absence of a given spore species is needed.
Examples of spore samplers. a) Passive sampler comprising two microscope slides mounted horizontally on a wooden pole; b) rotating-arm impactors; c) Burkard 7-day recording volumetric sampler; d) funnel-like passive spore sampler from Spornado (courtesy of Kristine White and Yaima Arocha Rosete, Sporometrics); and e) Burkard high-volume cyclone (courtesy of Celine Hamon, Vegenov)
Funnel-type samplers also belong to the category of passive samplers. They consist of a vertical cylinder topped by a funnel, at the base of which is installed a filter where are collected the spores captured by the sampler. This type of sampler is suitable for spores travelling long distances and deposited during rainfall events, such as Phakopsora pachyrhizi (Asian soybean rust) (Isard et al. 2011), or for spores dispersed by splashes, such as Fusarium graminearum (Paul et al. 2004). It is possible to quantify the inoculum per volume of water sampled when a weather station is installed nearby or when a rain gauge is installed directly on such a sampler.
The third type of passive sampler consists of a cylinder ending in a fin (or aileron) installed on a ball joint, so that it is facing the wind constantly. Spores are collected in a filter installed at the bottom of the cylinder. This type of device has the advantage of being inexpensive, which allows for large-scale deployment, but it hardly provides the inoculum concentration. In addition, counting spores on the filter can be laborious as they cannot be counted directly without first being dislodged from the filter mesh. This type of device is used in particular for large-scale monitoring, when binary data (presence/absence) is sufficient. This type of sampler has been used to monitor the progression of Asian soybean rust (P. pachyrhizi) throughout North America, from Florida to Canada (von Qualen and Yang 2006). This kind of device is also used to monitor potato late blight (P. infestans) in Ontario (Eugenia Banks, personal communication) (Fig. 1D).
Active (or volumetric) spore samplers are much more widely used, mainly because they allow the calculation of airborne spore concentrations (in spores/m3 of air). As opposed to passive samplers, these samplers are referred as active because the sampled particles are collected actively and use a collection mechanism (a rotor, a pump or an electrostatic force field). They can be classified into four main categories: impaction samplers, cyclone samplers, ionic samplers and virtual impactors (West et al. 2008; West and Kimber 2015). The first can involve impaction on Petri dishes containing a culture medium, on adhesive surfaces (tape, slides, rods) or in a liquid medium (Lacey and West 2006) (Fig. 1B–C). Cyclone samplers collect spores directly into a microcentrifuge tube to facilitate sample manipulation (Fig. 1E). Ionic samplers capture spores on an electron microscope electrode using a powerful electric field. Spores can therefore be counted directly using a scanning electron microscope. The virtual impactor can sample up to 850 liter of air per min (Limpert et al. 1999). It allows to separate the sampled particles according to their size into two air currents. Smaller particles with low inertia are deflected with the main flow while larger particles with greater inertia are directed to the collection device (Schwarzbach 1979).
In general, rotating-arm-type impaction samplers and 7-day volumetric spore samplers are well suited for monitoring fungi and oomycetes spores larger than 10 μm. These are the two most commonly used types of samplers in agriculture (Carisse et al. 2005, 2007, 2008, 2009b, 2012, 2013; Choudhury et al. 2016; Fall et al. 2015a, 2015b; 2015c; Friedrich et al. 2003; Hellin et al. 2018; Klosterman et al. 2014; Kunjeti et al. 2016; Reich et al. 2016; Van der Heyden et al. 2012; 2014). For small particles, a filter sampler (e.g., Button Aerosol Sampler) or a wet cyclone (e.g., Bertin Coriolis sampler) may be more appropriate.
5 Sampling frequency and duration
One of the most important factors for the monitoring of airborne spores is undoubtedly the periodicity at which spores are released into the air. This characteristic dictates the best time of day for sampling, its frequency and duration. The term “sporulation” generally includes the production of spores and their dispersion in the air. For many species, spore production takes place at night, while spore dispersion usually takes place during the day. Periodicity has been studied more particularly since 1953, for Alternaria sp., Cladosporium spp., Ustilago spp., Erysiphe spp., Polythrincium trifolii and P. infestans (Hirst 1953). In Hirst’s pioneer work, it is suggested that spore emission is generally highest around noon (closer to 11h00 for P. infestans and P. trifolii and closer to 13h00 for Alternaria spp., Cladosporium spp., Ustilago spp. and Erysiphe spp.) (Hirst 1953).
Subsequently, with the advancement of technical and practical knowledge, periodicity patterns were more accurately described for different pathogens. For example, a specific circadian dispersion pattern was described for P. infestans (Aylor et al. 2001). Although the results are essentially consistent with Hirst’s work (i.e., emission peaks around 11h00), Aylor et al. (2001) found that spores could be captured between 8h00 and 20h00. For their part, Hildebrand and Sutton (1982) reported that, although the first sporangia of Peronospora destructor could be captured on average 1.5 h after sunrise, the time of emission peaks could vary with relative air humidity, drying of foliage, and wind speed. In this study, spores were collected mainly under winds between 0.3 and 1.0 m/s and emission peaks measured between 10h00 and 14h00 (Hildebrand and Sutton 1982). Similarly, the airborne concentrations of Bremia lactucae sporangia, which are responsible for lettuce downy mildew, are variable, with emission peaks between 8h00 and 14h00 (Fig. 2) (Fall et al. 2015a; Fall et al. 2016).
Example of periodicity for Bremia lactucae. Periodicity can be defined as the description of circadian cycles of spore emission. Emission peaks are influenced by relative air humidity, drying of foliage, and wind speed. Splitting sampling over time extends the sampling time and therefore makes the sampling more representative. The grey and black bars represent two distinct sampling days
Whereas some pathogenic fungi have clear periodicity patterns, others are less characteristic. In carrot cultivation, for example, the emission patterns of Alternaria dauci spores tend to follow a normal curve with a nearly constant spore concentration between 10h00 and 16h00 (Strandberg 1977). Similarly, a study on the inoculum sources of Alternaria spp. suggested that peaks of spores emission ranged from 9h00 to 17h00 (Fernández-Rodríguez et al. 2015). This variation in spores periodicity patterns was also observed for other airborne pathogen like Botrytis cinerea and B. fabae, which were reported to be between 10h00 and 14h00 with occasional peaks between 14h00 and 17h00 in raspberry (Jarvis 1962), between 8h00 and 14h00 in strawberry and bean (Blanco et al. 2006; Fitt et al. 1985).
In order to adjust for this variation in circadian patterns of sporulation, several studies suggest splitting the sampling period. In onions, for example, spore sampling for B. squamosa was initially set for a fixed period of 2 hours between 10h00 and 12h00 (Carisse et al. 2012; Carisse et al. 2005; Carisse et al. 2007, 2008; Carisse et al. 2009b; Van der Heyden et al. 2012), but experiences has shown the importance of splitting the sampling period, which is now distributed between 8h00 and 14h00, 50% of the time (Carisse and Van der Heyden 2017). In a study conducted in Quebec to characterize the spatial distribution patterns of airborne conidia of Podosphaera aphanis in strawberries, the sampling period was fractionated so that effective sampling was carried out 30% of the time between 10h00 and 15h00 (Van der Heyden et al. 2014). Similarly, in a study conducted in New Brunswick, sampling of airborne inoculum from P. infestans was performed for 50% of the time between 6h00 and 15h00 (Fall et al. 2015b). This sampling range was also used for monitoring B. lactucae in lettuce (Fall et al. 2015a). Hence, the sampling period and frequency within the sampling period depend on the period and duration of the spore release as well as how fast the device sampling surface (e.g., rods, tape, etc.) become saturated. To circumvent the need to split the sampling period, others have chosen to use the 7-day sampler, which enables to obtain hourly or daily data (Hellin et al. 2018; Reich et al. 2016; Rogers et al. 2009).
6 Sampling height
Once the sampling period has been determined, the height at which the spore sampler is placed becomes one of the most important criteria to consider. The height at which a sampler should be installed depends largely on the objective of the study and in the best-case scenario, a tower with samplers installed at different heights should be recommended (Aylor 2017). However, it is often difficult to install such towers in the field, with the exception of experimental plots. Hence, a general rule of thumb might be to place the sampler above the canopy for the measurement of local inoculum, while the measurement of incoming inoculum would require the samplers to be placed higher, a few meters above the ground. Several examples are provided in Table 1, and more details are given below for selected examples.
The effect of the sampler height on spore concentration measures made using impaction samplers has been studied for some plant pathogens (Aylor 1995; Aylor and Taylor 1983). The objective of these studies includes estimating the proportion of spores escaping from the canopy from a point source. This is achieved by installing spore samplers at regular distances above a source and counting their contents at regular intervals to characterize vertical dispersion gradients. In a 1981 test, concentrations of P. tabaccina spores, for example, ranged from 384.6 spores/m3 of air at 0.75 m above the ground to 0.9 spores/m3 of air at 4 m above the ground (Aylor and Taylor 1983). On average, concentrations measured at 4 m above the ground represented only 12% of the spore concentrations measured at 0.75 m above the ground (Aylor and Taylor 1983). In the case of Venturia inaequalis, responsible for apple scab, spore concentration also decreased rapidly with height, with concentrations measured at 3 m corresponding to 6% of the concentrations measured at 0.15 m above the ground (Aylor 1995). The case of P. infestans has been extensively studied in this regard. In tests conducted in 1999 and 2000 in the State of New York, the vertical distribution of P. infestans spores showed that spore concentration decreased significantly with height, from nearly 6,000 spores/m3 of air at 0.75 m above ground to less than 10 spores/m3 of air at 3 m above ground in 1999 and from 2000 to 10 spores/m3 of air in 2000 (Aylor et al. 2001). In other words, for crops with low vegetation cover, the concentration of inoculum from a point source in the field quickly fades and the probability of detection is almost null from 3 m above the ground.
For a point source inoculum, the effect of the plume of spores escaping from the canopy dilutes rapidly, and the probability of capturing spores decreases with distance, vertically and horizontally (Severns et al. 2018). The effect of sampling height on the concentration of spores collected from multiple sources is much less documented. However, under field conditions, it can be assumed that there are multiple sources of inoculum and the horizontal flow of spores in the wind direction is equal to the cumulative concentration of spores escaping from the canopy from each source, between the edge of the field or the first source and the location of the sampler (Aylor 2017). Thus, at the field scale, spore concentrations increase in the prevailing wind direction based on the distance from the farthest source and the number of sources between the farthest source and the sampler location (Fig. 3). Hence, it is possible to optimize the position of the spore sampler according to the distance from the edge of the field or the number of inoculum point sources in the direction of the prevailing winds (Fig. 3) (Aylor 1995; Chamecki et al. 2011).
Horizontal flows of spores in the direction of the wind correspond to the cumulative concentration of spores escaping from the canopy (for each source) between the edge of the field (or first source) and the location of the sampler. Spore concentrations become almost constant at distance d from the point X0 at the edge of the field (or plot) in the direction of the wind from the sampler location (Xi). Adapted from Aylor (2017)
Knowledge of vertical distribution patterns can be exploited to the user’s advantage, particularly to distinguish inoculum originating from within the field versus from outside the field (incoming inoculum). For example, samplers that are part of an airborne inoculum monitoring network dedicated to monitoring the local Botrytis squamosa inoculum, responsible for botrytis leaf blight, are deployed at a height of 1 m early in the season and their height is adjusted according to plant growth (Carisse et al. 2012; Carisse and Van der Heyden 2017; Van der Heyden et al. 2012). For strawberry powdery mildew, samplers installed 1 m above the ground were found to be more representative of a plot than those installed at a height of 0.35 m (Van der Heyden et al. 2014). In research conducted in California on spinach and lettuce, spore samplers used to monitor downy mildew were installed at 0.53 m above the ground, for practical reasons (Choudhury et al. 2016; Klosterman et al. 2014; Kunjeti et al. 2016). In Belgium, the relationship between airborne concentrations of Fusarium graminearum spores and concentrations of deoxynivalenol (DON) was characterized using spore samplers installed 1 m above the ground (Hellin et al. 2018). Concentrations of grape powdery mildew in Oregon and Quebec and B. cinerea in strawberries, are monitored using spore samplers placed at heights of about 1 m (Carisse et al. 2009a, 2014; Thiessen et al. 2016). In a different context, spore samplers were also used between 2009 and 2012 for the implementation of a monitoring network for the inoculum of P. infestans in potatoes, aimed at characterizing regional dispersion patterns and identifying exogenous inoculum sources (Fall et al. 2015b). In this context, the samplers were installed at 2.9 m high to collect spores mostly coming from outside the fields.
7 Spatial distribution patterns and dispersal gradients
During the establishment of a spore sampling network, it is important to know as much as possible about the spatial distribution of the pathogen airborne spore concentrations to be monitored. More precisely, the study of spatial distribution patterns allows one to deduce the nature of the dispersion processes of the pathogens or diseases of interest and hence, improve the sampling. Does the pathogen spread over short distances from one plant to its neighbor? Or does it travel over long distances from one field to another, or even from one region to another? In other words, learning the spatial scope of the mechanisms involved in inoculum dispersion processes for the monitored species is the aim. In an agricultural field or a plantation, diseased plants among the plants present can have three types of spatial distribution: regular, completely random and aggregated (Fig. 4).
The type of spatial distribution has an impact on the effectiveness of sampling strategies. In the case of traditional field scouting (symptom scouting), the probability of finding a diseased plant is higher when the distribution of the disease in the field is completely random, compared to an aggregated distribution (Madden and Hughes 1999; Mahaffee and Stoll 2016). The spatial distribution of symptoms in the field was characterized for different pathosystems, revealing distribution patterns that varied between pathosystems. For example, the spatial distribution of Sclerotinia sclerotiorum symptoms in bean cultivation is largely dominated by a completely random pattern, both in pods and foliage (Jones et al. 2011). Patterns for hop powdery mildew (Podosphaera macularis) also follow a completely random distribution in general (Turechek and Mahaffee 2004). A study conducted in Quebec suggests a completely random spatial distribution pattern for strawberry powdery mildew for 72% of the sampling dates (Van der Heyden et al. 2014). In the case of field scouting, the sampling routes and number of samples can be adjusted to the spatial distribution, making this epidemiological characteristic a non-limiting factor.
Unlike field scouting for symptoms, air sampling is more likely to detect the onset of an epidemic if the distribution of inoculum sources is aggregated and the sampler is positioned to intercept the plume of spores emitted (Aylor and Irwin 1999; Mahaffee and Stoll 2016). A good knowledge of the distribution patterns is therefore important to develop a reliable inoculum monitoring network based on spore samplers. However, despite the importance of the spatial distribution patterns, specific to each pathogen, very few studies described these patterns of spatial distribution of airborne spores at the field and regional scales.
At the field or plot scale, one of the first studies describing spatial dispersion patterns for spores targeted apple scab, caused by Venturia inaequalis. The results of this study, conducted on 40 108-m2 plots in Quebec, suggested a patchy aggregated distribution pattern for most sampling dates and a spatial autocorrelation range between 25 and 53 m (Charest et al. 2002). Similarly, the spatial distribution of P. aphanis spores in strawberries is characterized by an aggregated pattern with no distinct focus (Van der Heyden et al. 2014). However, spatial distribution patterns vary with time and disease intensity. For example, for B. squamosa in onions, spore concentrations measured in small plots reveal random dispersion patterns at the beginning of the season or when the inoculum concentration is low and more aggregated patterns when inoculum concentrations are higher (Carisse et al. 2007).
For some pathogens, spatial distribution and dispersion patterns have been characterized at a larger scale (regional or national). Such knowledge is particularly relevant to predict the risks of development of an epidemic at the territory scale rather than the field scale. Potato late blight, caused by P. infestans, is probably the most documented in this regard. In New Brunswick, spatial distribution patterns of P. infestans spores have been characterized, and the results suggest a heterogeneous distribution with the increase in spore concentrations during the season (Fall et al. 2015b). In addition, the authors note that this information can be used to guide decision making with a limited number of spore samplers. Results from the same study also suggest that epidemics of downy mildew are not very affected by the size of the initial spore load, and support monitoring of the inoculum throughout the production period. Spatial distribution patterns of P. effusa spores in spinach have been characterized as highly aggregated, with the distribution of spores fitting a power-law distribution, in the Salinas Valley, California (Choudhury et al. 2016). In this study, however, spore samplers were installed 0.53 m above the ground, which possibly represents the plot better than the region.
The aggregation patterns observed at the field scale in the studies above are generally the result of auto-infection processes (Mundt 2009). The concepts of auto- and alloinfection are important in epidemiology because they are largely associated with the dispersion of pathogens. Both concepts were first introduced by Robinson (1976), who considered that autoinfection occurs when an infected plant or part of a plant (donor) contaminates the same plant or part of a plant (recipient), and conversely, that alloinfection occurs when the donor and recipient are different. Autoinfection therefore is a process that occurs over short distances compared to the alloinfection process (Mundt 2009; Zawolek and Zadoks 1992). In some studies, autoinfection is considered to be an infection resulting from propagules belonging to the same genotype, produced from the same foci of contiguous plants, whereas alloinfection is considered as an infection resulting from propagules produced from other genotypic units in the population (Mundt and Browning 1985; Zawolek and Zadoks 1992). Autoinfection and allonfection generally occur simultaneously, and their occurrence can be expressed as a proportion of the sum of infections. Thus, the ratio of allo- and autoinfection varies throughout an epidemic, the proportion of autoinfection being higher at the beginning. The disease tends to accumulate locally until a dispersion threshold is reached, beyond which transmission of the disease to another location becomes likely (Aylor 2017).
When a focus of infection has reached the dispersal threshold, it becomes a source of inoculum that emits over varying distances. Dispersal gradients from single point source were described for several plant pathogens. Some pathogens are disseminated over short distances from a single source. For example, Fusarium head blight seeds infection caused by Gibberella zeae was shown to decrease to 10% of the maximum within 5 to 22m from the focal center of an inoculated plot (Fernando et al. 1997; Paulitz et al. 1999), while spores of Mycosphaerella fijiensis, responsible for Black leaf streak disease, showed a gradient of up to 12.5 m (Rieux et al. 2014). Other plant diseases like P. infestans can be dispersed over several 100 m to up to 700 km (Severns et al. 2018). Dispersion rates are also influenced by the manner in which spores are released. In simple terms, spores can be divided into two categories: type 1 and type 2 (Fig. 5). Type 1 spores, such as Botrytis spp. conidia, are passively released from their attachment structure (e.g., conidiophores) by turbulent and unstable winds (Aylor 2017). The wind speed required to detach and disperse type 1 spores is species-specific (Aylor 2017). Unlike type 1 spores, type 2 spores are released regardless of wind speed. These spores can be actively released, as with Peronospora destructor, which uses a hygroscopic torsion mechanism to release its sporangia (Leach 1982; Leach et al. 1982).
Types of spores according to their release mechanism: a) passive release (type 1) and b) active release (type 2). Adapted from Aylor (2017)
The architecture of the crop canopy (height, density) and the position of infection sites on plants also influence the rates of auto- and alloinfection. The higher the crop and the denser the canopy (e.g., potatoes, beans), the lower the probability that spores escape from the canopy, unless sporulation takes place on the top of the canopy. When infections are located at the base of plants instead of the top, the number of spores escaping from the canopy will also be lower. All these factors make it difficult to quantify the phases of autoinfection. It is only when a dispersion threshold corresponding to alloinfections and specific to the pathosystem is reached, that it is possible to intercept spores using spore samplers.
The understanding of spatial distribution patterns can be facilitated by a good comprehension of the structure of the plant pathogen populations. In turn, this knowledge could also help to better plan the deployment of aerobiology experiments. For example, the presence of different mating types in the same location may suggest the production of overwintering inoculum, and thus the probabilities of the inoculum being local are greater. In their study of M. graminicola, Zhan et al. (2002) suggests an equivalent proportion of the two sexual types at the field or country scale. In this context, the sources of inoculum may be local, and the deployment of experiments adapted accordingly. In other cases, the presence of compatible sexual types may vary from region to region. This is the case of P. infestans for whom the mating types A1 and A2 can be present in the same field which implies that the overwintering inoculum could play a role in the initiation of epidemics (Widmark et al. 2007). In other regions only A1 or A2 types are present, which implies an important role for external inoculum sources (Blandón-Díaz et al. 2011; Peters et al. 2014). Thus, this knowledge can also influence the planning of research initiatives (Fall et al. 2015).
8 Spore sampler networks
Results obtained by monitoring using spore samplers should be interpreted with great caution, as the collected spores may come from a large but distant source or from a smaller source nearby (West and Kimber 2015). For this reason, the concepts of auto- and alloinfection must also be defined at the landscape scale (region or larger territory). Thus, the plant, plot or field becomes the basic unit; inoculum accumulation locally takes place at the scale of the smallest unit in the system; and autoinfection is also defined within this unit (Mundt 2009). When a critical level is reached, spores are transported outside the boundaries of this basic unit (plant, plot, or field), and it becomes possible to proceed to epidemiological surveillance. At the landscape level, it becomes possible to measure alloinfection on a small scale and auto- and allo-infection on a larger scale. It is at this scale that spore sampling networks can be deployed.
There are as many types of spore sampling networks as there are networks, in other words, there are no universal rules that apply to all networks. Of course, the structure of the network should correspond to the basic sampling objectives. Airborne inoculum monitoring is a scouting technique, so the higher the number of samplers and the frequency of sampling, the more reliable and representative the data will be. While in general it is the availability of resources that dictates how data will be collected, understanding the elements discussed in this review should enable managers, regardless of their network size, to make the best possible decisions given their own context. Ideally, one sampler would be positioned in each field of the network, but the obvious logistical constraints do not allow this kind of density. Hence, airborne inoculum monitoring networks can be built on an existing disease scouting network and structured to combine the two types of data collection, but they evolve gradually to adapt to the needs of the users (Van der Heyden et al. 2012).
Though implemented on different continents, few examples of spore sampler networks are documented in the literature. The network of passive samplers installed across the eastern United States to monitor the south-north progression of spores of Asian soybean rust (Phakopsora pachyrhizi) is among the most extensive. Through the ipmPIPE information platform (https://www.ipmpipe.org), this network was designed to model the progression of the disease across the US and in Canada and to demonstrate that the use of spore samplers was less costly than and as effective as sentinel plot monitoring, which had formerly been used for the network (Isard et al. 2011). Results of this project suggest that the information collected at the continent scale can be used to limit the number of inoculum sources, but above all, to delay the progression of the disease along the south-north axis (Isard et al. 2011).
In Belgium, spore sampler networks have been assessed for monitoring the airborne inoculum of wheat yellow rust (Puccinia striiformis f. sp. tritici) (Duvivier et al. 2016; Hellin et al. 2018; Dedeurwaerder et al. 2011). The approach was followed in Wallonia in 2008-2009, using active volumetric samplers (7 day recording volumetric sampler). Results suggest that spore sampler networks could be used to estimate the concentration of P. striiformis spores and predict yellow rust epidemics (Dedeurwaerder et al. 2011). Similarly, a monitoring network including the use of spore samplers is being deployed in England to monitor various diseases, in wheat, canola and potatoes among other crops. In this example, the use of spore samplers is coupled with sentinel plots (untreated), a network of weather stations allowing the use of predictive models, and a network of collaborators providing disease observations (https://www.fera.co.uk/our-science/active-r-and-d/in-field-diagnostics). This network is operated jointly by a private research organization (Fera). The data are accessible through a web platform, and producers can subscribe to a text message alert service.
A slightly different approach is being assessed in Australia for monitoring diseases in wheat (Pyrenophora tritici-repentis) and canola (e.g., Leptosphaeria maculans). The working group is composed of researchers from the South Australian Research and Development Institute, in collaboration with the Australian Department of Agriculture and grower groups. The approach is based on the use of two types of spore samplers: fixed Cyclone and mobile Jet Spore-type mounted on the roof of a light truck (Narayan et al. 2010). The approach is still under development (2017-2022), but a new initiative has been launched to add grapes, sugar cane, cotton, and nursery trees to the list of monitored plants.
In Brittany, the Vigispore Initiative, by the private research organization Végénov, was launched in January 2017. This initiative aims to assess the use of fixed spore samplers such as the Cyclone for monitoring airborne concentrations of P. destructor, B. squamosa, and B. allii spores in shallot cultivation (Dr Celine Hamon, personal communication). The use of spore samplers in combination with forecast model predictions has been identified as a potential avenue to reduce the fungicide treatment frequency index in the region. This initiative, deployed on two sites in 2017 and 2018 and six sites in 2019, is inspired by the spore sampler network deployed in the Jardins-de-Napierville Regional County (Quebec) for onions.
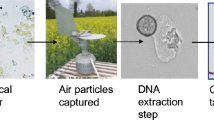
The Quebec spore collection network mentioned above has been running annually since 2008 (Van der Heyden et al. 2012). Unlike other networks that aim to monitor at larger spatial scales (state, country or continent), it is designed for regional monitoring at the 30-km-by-30-km scale. This spore sampler network is an example of sustainable airborne inoculum monitoring, and as such, involves researchers affiliated with private organizations, researchers from federal agencies, provincial collaborators, technical support clubs, onion producers and even, to some degree, processors. Spore sampling is carried out using rotating-arm type of samplers installed in about 20 fields, depending on the distribution of producing fields. Spore samplers are in operation three times per week and sample the air between 8h00 and 14h00 50% of the time. Spore concentrations are counted using qPCR assays and results are transmitted in near-real time to growers and their farm advisors, accompanied by the disease risks predicted by a forecasting model, so that the best decision can be made regarding appropriate phytosanitary treatments or actions (Fig. 6). The work carried out through this monitoring initiative has resulted in a significant reduction in the environmental risk index, health risk index and treatment frequency index (Carisse and Van der Heyden 2017). More precisely, this approach reduced the three indices by 32%, 14%, and 28% respectively on average in 2015–2017, compared to the reference period (2007–2009).
9 Spore counting
The effectiveness of airborne spore monitoring systems for disease management decision making depends in particular on the speed of spore counting and the accuracy in identifying the targeted spores. The simplest method of counting in terms of equipment is certainly microscopic counting. However, in general, samples contain so many different spores and other particles that it is essential to determine what needs to be counted before starting. Although this method has been widely used, it can be long and tedious, even for experienced personnel. Several factors must be considered, including the size of the particles to be identified, the magnification and staining (aniline blue, safranin, etc.). Nevertheless, this counting method has been widely used, particularly for B. squamosa in onions (Carisse et al. 2012; Carisse et al. 2005; Carisse et al. 2008; Van der Heyden et al. 2012a), P. aphanis in strawberries (Carisse et al. 2013; Van der Heyden et al. 2014b), P. infestans in potatoes (Fall et al. 2015b), B. lactucae in lettuce (Fall et al. 2015a; Fall et al. 2016), V. inequalis in apples (Charest et al. 2002), and many others (Table 1). Visual identification and counting of spores by microscopy remain uneasy and of limited accuracy. Spores found in airborne samples generally have a different appearance than in an aqueous solution or when coming from a pure culture. They are normally dehydrated and can be deformed due to impaction and rotation of the sampling surface. In many cases, morphological characteristics of spores are very similar between species, making species identification very difficult for some spores. This is notably the case for powdery mildews (e.g., Erysiphe necator, Podosphaerea aphanis), which are relatively difficult to identify when other species of the same genus originating from nearby weeds may be present on the sample. Also, it is more and more important to distinguish genotypes (e.g., formae speciales, clonal lineage, races, fungicide resistant genotypes), and this is not possible on the basis of spore appearance. In addition, the presence of soil particles, dust, pollen and other debris can complicate spore identification (Fig. 7). Thus, it is often necessary to measure the dimensions of each particle and compare with reference specimens, or regularly adjust the focus to find all particles and quantify them adequately (Lacey and West 2006). Such constraints can substantially increase the time required to identify spores and complicate precise species identification. However, microscopic counts can be used to determine presence/absence or estimate categories of spore density such as low, moderate, and high for a given spore species.
In order to overcome the shortcomings of microscopy in terms of specificity, standardize the counting methods and reduce the response time, different approaches have been developed over the years. Among the alternative methods, the use of monoclonal antibodies combined with a lateral flow device has been assessed for some pathogens (Kennedy and Wakeham 2008). However, these methods have very low sensitivity, and although they appear to be specific, their detection limit is often inadequate. For Peronospora destructor, for example, the detection limit of such a system was found to be about 500 sporangia (Kennedy and Wakeham 2008). Hence, molecular approaches using PCR and qPCR methods were assessed, and have proven to be more appropriate tools for spore counting. The first plant pathology trials were carried out in the early 2000s, with Penicillium roqueforti as a model organism. This trial, conducted using conventional PCR combined with agarose gel electrophoresis, allowed the specific detection of 10 spores, which represents a substantial improvement over immunoassays (Williams et al. 2001). Although it was not possible to obtain a spore concentration in the sample, this trial paved the way for several developments. Quantitative PCR methods were designed for different systems, including Sclerotinia sclerotiorum (Rogers et al. 2009) and Erysiphe necator (Falacy et al. 2007). After publication of the marker for the identification and quantification of B. squamosa under real field conditions (Carisse et al. 2009b), the qPCR approach has been intensively used. Subsequently, molecular markers were developed for fungi including, among others, Botrytis cinerea (Carisse et al. 2014; Suarez et al. 2005), Peronospora effusa (Klosterman et al. 2014), P. infestans (Fall et al. 2015c), B. lactucae (Kunjeti et al. 2016), P. cubensis and P. humuli (Summers et al. 2015; Rahman et al. 2020), and Fusarium graminearum (Hellin et al. 2018).
With air sampling, especially in agricultural fields, a variety of inhibitors may be present in the samples to be analyzed by qPCR, which can increase the frequency of false negatives. Thus, care must therefore be taken to include controls to detect these false negatives (McDevitt et al. 2007). Several approaches like the dilution of sample DNA or the use of additive and PCR enhancer such as BSA or Polyvinylpyrrolidone can be used to circumvent the presence of PCR inhibitors. A more common practice is to include a second exogenous target added to the system in a known amount that is constant from sample to sample (Haudenshield and Hartman 2011). In addition to detecting false negatives caused by the presence of inhibitors, these exogenous internal controls also enable this inhibition to be quantified and used to correct the values obtained (Bilodeau et al. 2011).
10 Thresholds and interpretation of aerobiological data
For an efficient disease management, it is essential to be able to predict the risk of an epidemic. Thresholds are therefore an important tool in integrated or reasoned management as they make it possible to select the best time to respond as well as the type of response. Regardless of their type, thresholds are indicators that reduce the frequency of incorrect decisions (i.e., responding when unnecessary and not responding when necessary). The economic threshold is the disease severity level at which the cost of treatment becomes lower than the estimated cost of losses, whereas an action threshold, also called treatment threshold, is the disease level at which treatments is less effective due to high disease pressure. A damage threshold or tolerance threshold is reached when the estimated losses at harvest are higher than the acceptable disease level (for example: 5% grey mould severity in grapes).
Thresholds are established in accordance with the relationship between the values of the indicator (disease incidence or severity, spore concentration) and the damage (loss of yield, lower quality) (Fig. 8). When this relationship is known, as well as the acceptable damage level, it is simple to establish the threshold. However, in practice, the threshold varies with the crop development stage, the expected harvest date, the type of market (fresh, storage, processing), and the grower’s risk tolerance level. Fungal diseases with aerial dispersion are generally referred to as “polycyclic,” i.e., the population of the plant pathogenic fungus increases with each secondary reproductive cycle. The number of reproductive cycles is difficult to predict because it depends on the fungus, the crop’s sensitivity, weather conditions, and the length of the growing season.
(a) Plots of the severity of powdery mildew on strawberry leaves (down-pointing white triangles), the airborne concentration of Podosphaera aphanis spores (black circles), and crop losses (grey bars) in relation to the day of year. (b) Proportion of scabbed leaves per shoot (Venturia inaequalis) at the beginning of the growing season and percentage of scabbed fruit at harvest. The black circles were used for model development and white triangles for validation purposes (Carisse et al. 2009; Carisse et al. 2012; Carisse et al. 2013)
Action thresholds based on airborne spore concentration are difficult to define, as the concentration varies with the number and proximity of spore sources. For this reason, information on spore concentration is often interpreted in terms of the time of capture of the first spores (Thiessen et al. 2016; Van der Heyden et al. 2012) and the increase in the aerial spore concentration (Carisse et al. 2012). For onion leaf blight (Botrytis squamosa), a first action threshold was developed by comparing airborne concentration of conidia with number of lesions per leaf under field conditions. The threshold was determined based on concentrations corresponding to disease-based thresholds already used to time fungicide applications with values of 10-15 and 25-35 conidia per m3 corresponding to 1 and 2.5 lesions per leaf, respectively (Carisse et al. 2005) (Fig. 9a). Based on this study, airborne inoculum-based thresholds (first conidia and total of 15 conidia per m3 caught) were compared to a disease-based threshold (one lesion/leaf) to time the initiation of fungicide spray program for onion leaf blight management (Van der Heyden et al. 2012) (Fig. 9b). In this study, best disease management was obtained when fungicide spray programs were initiated at first airborne B. squamosa conidia detected, followed by a threshold of a total 15 conidia per m3, and by initiation when the first lesion was detected. To further improve onion leaf blight management, a disease risk indicator was developed by combining weather-based risk of infection, amount of airborne inoculum and forecasted favorability of weather conditions for infection (Carisse et al. 2012).
a) Relationship between an economic treatment threshold and the aerial concentration of Botrytis squamosa spores. The dotted black line is the linear regression, and the solid grey lines represent the prediction interval (Carisse et al. 2005). b) Relationship between the progression of onion leaf blight and the indicator used to initiate the treatment program; down-pointing black triangles correspond to the threshold of 1 lesion per leaf, white circles to the threshold of 15 spores per m3 of air, and black circles to the threshold of 1 spore per m3 of air (Van der Heyden et al. 2012a)
Regardless of the approach followed, thresholds are highly dependent on the sampling method and disease management strategy. When the conditions of use of the spore samplers are well-defined and validated according to the development of diseases, the threshold must then be determined. In other words, the question is how to express the number of captured spores into risk and the decision of applying or not a fungicide treatment (synthetic or biological). Each fungus has its own survival strategy. Some produce a large number of spores when one individual spore has little chance of infecting a plant. In this case, the threshold will be high, e.g., with powdery mildews (strawberries, grapes). For example, Van der Heyden et al. (2014) find a linear relationship between airborne concentration of Podosphaera aphanis conidia with strawberry powdery mildew severity and reported that 5 and 10% disease severity corresponded to 50 and 100 conidia/m3. Other fungi produce fewer, very aggressive spores, as is the case with downy mildews (potatoes, vines). In these cases, thresholds must be lower. Thresholds should be determined based on controlled studies and field observations. Fall et al. (2015) studied the relationship between airborne sporangia concentration of four clonal lineages of Phytophthora infestans (US-8, US-11, US-23, and US-24) and late blight of potato development under controlled conditions. Even though the shape of the disease-inoculum curve were similar for all clonal lineages, infection efficiency of sporangia was different. The sporangia concentration required to cause one lesion per leaf was 10 sporangia per m3 for US-23 and 15 to 25 sporangia per m3 for the other clonal lineages of P. infestans.
Data from spore samplers should be interpreted by considering many other risk factors such as weather, crop susceptibility, disease level, and aggressiveness of the pathogen (Carisse et al. 2012; Fall et al. 2015b). A large number of spores in conditions that are unfavorable to disease development may represent a lower risk than a small number of spores under favorable conditions. Thresholds should also be interpreted in accordance with the growth stage of the crop. Therefore, the presence of spores does not necessarily mean that an action (treatment) is necessary.
11 Perspectives
In recent decades, global trade in plant materials and products has contributed to the spread of many pathogens. In addition, climate change tends to increase the length of the growing season, especially in northern territories (Evans et al. 2008). Consequently, these changes could influence seasonal epidemics of resident (endogenous) pathogens, promote the production and survival of overwintering inoculum, increase the frequency and scale of seasonal epidemics for exogenous pathogens, and change the status of some plant pathogens from periodically introduced to endemic (Van der Heyden et al. 2020). Moreover, pesticides use is under heavy scrutiny, and growers feel the pressure of the public, buyers and consumers, to reduce their dependence on synthetic pesticides. Tools are needed for the industry to estimate short- and middle-term risks and to identify the most appropriate control measures and times to apply phytosanitary measures, including those with low environmental risk. For governments, indicators to monitor the impacts of climate change and introduced pathogens are required. Monitoring the territory, particularly with networks of spore samplers, is perfectly coherent in this context.
The development of quantitative molecular approaches has allowed the deployment of spore sampler networks at different scales (Carisse et al. 2012; Carisse and Van der Heyden 2017; Fall et al. 2015c; Van der Heyden et al. 2012, Chen et al. 2018). The combination of these approaches allows an accurate monitoring of a predetermined number of pathogens, usually between one and four. However, molecular methods are evolving rapidly, and new promising approaches are continuously being developed. As an example, scientists are currently attempting to combine isothermal PCR technologies (LAMP-PCR, RPA-PCR) and automated spore sampling instruments to send real-time risk alerts to growers and crop specialists (West et al. 2018). Presently, these devices are expensive (~$30,000 USD per unit), their availability is limited, and the number of pathogens that they can monitor simultaneously is also limited. Several research groups are working on the development of molecular tools compatible with these approaches to combined spore sampling and real-time data transmission (Lees et al. 2019; Si Ammour et al. 2017). Automated image recognition technologies are also progressing rapidly, and with them, a true real-time monitoring approach is getting closer (see https://www.scanittech.com or https://bioscout.com.au).
It is impossible to ignore the emerging technology of single-molecule sequencing, also known as third-generation sequencing, and its potential to improve biomonitoring at multiple scales. This technology could quickly become a benchmark in terms of diagnosis and has several advantages over previous approaches. First, it allows the sequencing of long DNA fragments that are easier to assemble and align with reference genomes and sequencing entire small and medium-sized genomes such as those of bacteria and certain fungi. Presently, there are two third-generation sequencing platforms: Pacific Biosystems (PacBio) and Oxford Nanopore Technologies (ONT). The ONT platform offers a faster response time (between 3 and 5 h), requires only a small amount of initial DNA, and has lower costs of acquisition (Hu et al. 2019). In 2014, ONT launched the first portable third-generation sequencer, the MinION. Its small size and ability to connect to a local computer make it suitable for use in all laboratories. Several bioinformatics tools have been adapted for the MinION sequencer and allow real-time identification of plant pathogenic species present in a sample. This new technology has already been shown to be useful for phytosanitary biomonitoring (Chalupowicz et al. 2018; Bronzato Badial et al. 2018). In these trials, a number of plant species (e.g., tomato, pepper, melon, Gypsophila, strawberry, lemon) presenting unknown symptoms or knowingly inoculated with 16 bacterial, fungal, and viral pathogens, were analyzed using Nanopore’s tools. For all inoculated or symptomatic plants, the pathogen species were identified within 2 h (Chalupowicz et al. 2018).
Since the discovery that some pathogens are dispersed in the air, aerobiology has evolved from the establishment of principles (Gregory 1945), to molecular identification and quantification of airborne spores (Carisse et al. 2009), and statistical and mathematical modeling (Aylor 2017). Nowadays, the combined use of hightrouput sequencing (HTS) and bioinformatics in aerobiology, allows the simultaneous identification of several targets. An example of this new approach is presented in Chen et al. (2018) were the use of metagenomics using HTS is combined to a spore-sampling network (AeroNet) to detect specific taxonomic groups. In an example from forestry in Canada, Tremblay et al. 2018 identified regulated and unregulated pathogenic forest fungi in spore trap samplers. They identified potential species to be controlled as they could cause diseases not yet recorded in the region (Bérubé et al. 2018a, 2018b) and may require management procedures. In Europe, HTS methods have been used to study fungal diversity in different environmental substrates. In this study, it was shown that the assessment of airborne fungal community diversity can be achieved by using simple and robust passive spore traps (cellulose filters, coated microscope slides) (Aguayo et al. 2018). Redondo et al. (2020) evaluated the airborne fungal spore community and the fungal spore community deposited with active and passive spore traps in three different vegetation types in Sweden using metagenomics.
The use of these new technologies also accelerates the discovery of new genotypes with specific epidemiological characteristics, in terms of aggressiveness, host specificity or resistance to fungicides. Combined with the use of spore samplers organized in networks, these new tools will make it possible to carry out genotype-specific biosurveillance. Examples of this type of monitoring are not yet very abundant, but it is likely that this type of use will become more and more common. Among the available examples is the development of a monitoring tool for two host specialized clades of P. cubensis (Rahman et al. 2020), identified as a result of a genome wide association study (Wallace et al. 2020). Similarly, tools for simultaneous monitoring of Plasmopara viticola clade riparia and clade aestivalis was recently developed (Carisse et al. 2021). Another example of such an approach is provided for Z. tritici, for which molecular markers have been developed to monitor specific DNA substitutions associated with fungicide resistance to demethylation inhibitors and succinate dehydrogenase (Hellin et al. 2020). Hence, it is expected that aerobiology will continue to evolve through the integration of new technologies such as metagenomics, but also through other knowledge such as deep learning, neural networks or more automated methods of sample analysis.
In conclusion, airborne inoculum monitoring networks combined with advanced DNA analysis, and image and data analysis technologies open the door to a new generation of crop disease management tools and to better understanding of pathosystems. Regardless of the level of sophistication, once in place, airborne inoculum monitoring networks allow early detection of resident or immigrant pathogens; thus, control measures can be deployed before the disease gets out of control. In addition, DNA analysis can detect changes in the population of plant pathogens such as virulence or genetic substitutions related to resistance to fungicides. The observations from the networks allow the identification of the source (reservoir) of inoculum, to describe the dispersion of the pathogen at a territory scale, to identify the zones most at risk (hotspots), and to study the impact of climate change and cultural practices on the development of diseases. In other words, to improve our knowledge of pathosystems. As research data and data from inoculum tracking networks become available, it will be possible to specify the best location and improve spatial resolution.
In the context of the “big data” era, airborne inoculum monitoring networks are still in the juvenile phase and research must be done essentially to integrate available technologies, improve access, analysis, interpretation and data sharing. It is likely that with our constant need for more and more data, that local networks will be integrated into regional networks and then into national networks. It is therefore essential to properly structure the root networks; it is in this spirit that we have prepared this literature review.
Data availability
Not applicable.
References
Aguayo J, Fourrier-Jeandel C, Husson C, Ioos R (2018) Assessment of passive traps combined with high-throughput sequencing to study airborne fungal communities. Appl Environ Microb 84(11):e02637–e02617. https://doi.org/10.1128/AEM.02637-17
Aizenberg V, Reponen T, Grinshpun SA, Willeke K (2000) Performance of Air-O-Cell, Burkard, and Button samplers for total enumeration of airborne spores. Am Ind Hyg Assoc J 61:855–864. https://doi.org/10.1080/15298660008984598
Alderman SC (1993) Aerobiology of Claviceps purpura in Kentucky bluegrass. Plant Dis 77:1045–1049. https://doi.org/10.1094/PD-77-1045
Aylor DE (1995) Vertical variation of airborne concentration of Venturia inaequalis ascospores in an apple orchard. Phytopathology 85:175–181. https://doi.org/10.1094/Phyto-85-175
Aylor DE (2017) Airborne dispersion of pollen and spores. The American Phytopathological Society Saint-Paul Minesota. https://doi.org/10.1094/9780890545430.
Aylor DE, Irwin ME (1999) Aerial dispersal of pests and pathogens: implications for integrated pest management. Agr Forest Meteorol 97:233–234. https://doi.org/10.1016/S0168-1923(99)00068-4
Aylor DE, Kiyomoto RK (1993) Relationship between airborne concentration of Venturia inaequalis ascospores and development of apple scab. Agr Forest Meteorol 63:133–147. https://doi.org/10.1016/0168-1923(93)90057-O
Aylor D, Taylor G (1983) Escape of Peronospora tabacina spores from a field of diseased tobacco plants. Phytopathology 73(4):525–529
Aylor DE, Fry WE, Mayton H, Andrade-Piedra JL (2001) Quantifying the rate of release and escape of Phytophthora infestans sporangia from a potato canopy. Phytopathology 91:1189–1196. https://doi.org/10.1094/PHYTO.2001.91.12.1189
Bashan Y, Levanony H, Or R (1991) Wind dispersal of Alternaria alernata a cause of leaf blight of cotton. J Phytopathol 133:225–238. https://doi.org/10.1111/j.1439-0434.1991.tb00157.x
Bérubé JA, Potvin A, Stewart D, Gagné PN, Ponchart JP, Phelan J, Varga A, James D, Tremblay ED, Duceppe MO, Kimoto T, Bilodeau GJ (2018a) Species, distribution and spore density of Heterobasidion in Canada. Forest Pest Management Forum abstract proceeding. 2018:42–43
Bérubé JA, Gagne PN, Ponchart JP, Tremblay ED, Bilodeau GJ (2018b) Detection of Diplodia corticola spores in Ontario and Québec based on High throughput sequencing (HTS) methods. Canadian Journal of Plant Pathol 40(3):378–386. https://doi.org/10.1080/07060661.2018.1498394
Bilodeau GJ, Koike ST, Uribe P, Martin FN (2011) Development of an Assay for Rapid Detection and Quantification of Verticillium dahliae in Soil. Phytopathology 102(3):331–343. https://doi.org/10.1094/PHYTO-05-11-0130
Blanco C, de Santos BL, Romero F (2006) Relationship between concentrations of Botrytis Cinerea conidia in air, environmental conditions, and the incidence of grey mold in strawberry flowers and fruits. Eur J Plant Pathol 114:415–425. https://doi.org/10.1007/s10658-006-0007-3
Blandón-Díaz JU, Widmark AK, Hannukkala A, Andersson B, Högberg N, Yuen JE (2011) Phenotypic Variation Within a Clonal Lineage of Phytophthora infestans Infecting both Tomato and Potato in Nicaragua. Phytopathology 102(3):323–330. https://doi.org/10.1094/PHYTO-02-11-0033
Bock CH, Cotty PJ (2006) Methods to Sample air borne propagules of Aspergillus flavus. Eur J Plant Pathol 114:357–362. https://doi.org/10.1007/s10658-005-4514-4
Brischetto C, Bove F, Languasco L, Rossi V (2020) Can Spore Sampler Data Be Used to Predict Plasmopara viticola Infection in Vineyards? Frontiers in Plant Science 11(1187). https://doi.org/10.3389/fpls.2020.01187
Bronzato Badial A, Sherman D, Stone A, Gopakumar A, Wilson V, Schneider W, King J (2018) Nanopore sequencing as a surveillance tool for plant pathogens in plant and insect tissues. Plant Dis 102:1648–1652. https://doi.org/10.1094/PDIS-04-17-0488-RE
Cao X, Duan X, Zhou Y, Luo Y (2012) Dynamics in concentrations of Blumeria graminis f. sp tritici conidia and its relationship to local weather conditions and disease index in wheat. European Journal of Plant Pathology 132(4):525–535. https://doi.org/10.1007/s10658-011-9898-8
Cao X, Yao D, Xu X, Zhou Y, Ding K, Duan X, Fan J, Luo Y (2014) Development of Weather- and Airborne Inoculum-Based Models to Describe Disease Severity of Wheat Powdery Mildew. Plant Disease 99(3):395–400. https://doi.org/10.1094/PDIS-02-14-0201-RE
Carisse O, van der Heyden H (2017) Networked real time disease risk evaluation: A cost-effective approach to disease management. Phytopathology 107(12):145–147
Carisse O, McCartney HA, Gagnon JA, Brodeur L (2005) Quantification of airborne inoculum as an aid in the management of leaf blight of onion caused by Botrytis squamosa. Plant Dis 89:726–733. https://doi.org/10.1094/PD-89-0726
Carisse O, Savary S, Willocquet L (2007) Spatiotemporal relationships between disease development and airborne inoculum in unmanaged and managed botrytis leaf blight epidemics. Phytopathology 98:38–44. https://doi.org/10.1094/PHYTO-98-1-0038
Carisse O, Savary S, Willocquet L (2008) Spatiotemporal relationships between disease development and airborne inoculum in unmanaged and managed botrytis leaf blight epidemics. Phytopathology 98:38–44. https://doi.org/10.1094/PHYTO-98-1-0038
Carisse O, Bacon R, Lefebvre A (2009a) Grape powdery mildew (Erysiphe necator) risk assessment based on airborne conidium concentration. Crop Prot 28(12):1036–1044. https://doi.org/10.1016/j.cropro.2009.06.002
Carisse O, Tremblay DM, Lévesque CA, Gindro K, Ward P, Houde A (2009b) Development of a TaqMan real-time PCR assay for quantification of airborne conidia of Botrytis squamosa and management of Botrytis leaf blight of onion. Phytopathology 99:1273–1280. https://doi.org/10.1094/PHYTO-99-11-1273
Carisse O, Levasseur A, Van der Heyden H (2012) A new risk indicator for botrytis leaf blight of onion caused by Botrytis squamosa based on infection efficiency of airborne inoculum. Plant Pathol 61:1154–1164. https://doi.org/10.1111/j.1365-3059.2012.02594.x
Carisse O, Morissette-Thomas V, van der Heyden H (2013) Lagged association between powdery mildew leaf severity, airborne inoculum, weather, and crop losses in strawberry. Phytopathology 103:811–821. https://doi.org/10.1094/PHYTO-11-12-0300-R
Carisse O, Tremblay DM, Lefebvre A (2014) Comparison of Botrytis cinerea airborne inoculum progress curves from raspberry, strawberry and grape plantings. Plant Pathol 63:983–993. https://doi.org/10.1111/ppa.12192
Carisse O, Tremblay DM, Hébert PO, Van der Heyden H, Delmotte D (2021) Evidence for differences in the temporal progress of Plasmopara viticola clades riparia and aestivalis airborne inoculum monitored in vineyards in eastern Canada using a specific multiplex qPCR assay. Plant Dis.:PDIS-06-20-1164. https://doi.org/10.1094/PDIS-06-20-1164-RE
Carvalho E, Sindt C, Verdier A, Galan C, O’Donoghue L, Parks S, Thibaudon M (2008) Performance of the Coriolis air sampler, a high-volume aerosol-collection system for quantification of airborne spores and pollen grains. Aerobiologia 24:191–201. https://doi.org/10.1007/s10453-008-9098-y
Chalupowicz L, Dombrovsky A, Gaba V, Luria N, Reuven M, Beerman A, Lachman O, Dror O, Nissan G, Manulis-Sasson S (2018) Diagnosis of plant diseases using the Nanopore sequencing platform. Plant Pathol 68:229–238. https://doi.org/10.1111/ppa.12957
Chamecki M, Dufault NS, Isard SA (2011) Atmospheric dispersion of wheat rust spores: a new theoretical framework to interpret field data and estimate downwind dispersion. J Appl Meteorol Clim 51:672–685. https://doi.org/10.1175/JAMC-D-11-0172.1
Charest J, Dewdney M, Paulitz T, Philion V, Carisse O (2002) Spatial distribution of Venturia inaequalis airborne ascospores in orchards. Phytopathology 92(7):769–779. https://doi.org/10.1094/phyto.2002.92.7.769
Chawda HT, Rajasab AH (1994) Aerobiology of Alternaria porri and its relation to purple blotch disease in onion. Indian J Mycol Plant Pathol 24:41–45. https://doi.org/10.1023/A:1011244104920
Chen W, Hambleton S, Seifert KA, Carisse O, Diarra MS, Peters RD, Lowe C, Chapados JT, Lévesque CA (2018) Assessing performance of spore samplers in monitoring aeromycobiota and fungal plant pathogen diversity in Canada. Appl Environ Microbiol 84:e02601–e02617. https://doi.org/10.1128/AEM.02601-17
Choudhury RA, Koike ST, Fox AD, Anchieta A, Subbarao KV, Klosterman SJ, McRoberts N (2016) Spatiotemporal patterns in the airborne dispersion of spinach downy mildew. Phytopathology 107:50–58. https://doi.org/10.1094/PHYTO-04-16-0162-R
Cohen Y, Rotem J (1971) Dispersal and viability of sporangia of Pseudoperonospora cubensis. T Brit Mycol Soc 57(1):67–74. https://doi.org/10.1016/S0007-1536(71)80081-5
Dedeurwaerder G, Duvivier M, Mvuyenkure S, Renard ME, Van Hese V, Marchal G, Moreau JM, Legrève A (2011) Spore traps network: a new tool for predicting epidemics of wheat yellow rust. Communications in agricultural and applied biological sciences 76:667–670
Dhar N, Mamo BE, Subbarao K, Koike S, Fox A, Anchieta A, Klosterman SJ (2019) Measurements of aerial spore load by qPCR facilitates lettuce downy mildew risk advisement. Plant Disease. 104:82–93. https://doi.org/10.1094/PDIS-03-19-0441-RE
Duvivier M, Dedeurwaerder G, Bataille C, De Proft M, Legrève A (2016) Real-time PCR quantification and spatio-temporal distribution of airborne inoculum of Puccinia triticina in Belgium. European Journal of Plant Pathology 145(2):405–420. https://doi.org/10.1007/s10658-015-0854-x
Evans N, Baierl A, Semenov MA, Gladders P, Fitt BDL (2008) Range and severity of a Plant Dis increased by global warming. J R Soc Interface 5(22):525–531. https://doi.org/10.1098/rsif.2007.1136
Falacy JS, Grove GG, Mahaffee WF, Galloway H, Glawe DA, Larsen RC, Vandemark GJ (2007) Detection of Erysiphe necator in air samples using the polymerase chain reaction and species-specific primers. Phytopathology 97:1290–1297. https://doi.org/10.1094/PHYTO-97-10-1290
Fall ML, Tremblay DM, Gobeil-Richard M, Couillard J, Rocheleau H, Van der Heyden H, Lévesque CA, Beaulieu C, Carisse O (2015) Infection efficiency of four Phytophthora infestans clonal lineages and DNA-based quantification of sporangia. PLOS ONE 10:e0136312. https://doi.org/10.1371/journal.pone.0136312
Fall ML, Van der Heyden H, Beaulieu C, Carisse O (2015a) Bremia lactucae infection efficiency in lettuce is modulated by temperature and leaf wetness duration under Quebec field conditions. Plant Dis 99:1010–1019. https://doi.org/10.1094/PDIS-05-14-0548-RE
Fall ML, Van der Heyden H, Brodeur L, Leclerc Y, Moreau G, Carisse O (2015b) Spatiotemporal variation in airborne sporangia of Phytophthora infestans: characterization and initiative toward improving potato late blight risk estimation. Plant Pathol 64:178–190. https://doi.org/10.1111/ppa.12235
Fall ML, Van der Heyden H, Carisse O (2016) A Quantitative dynamic simulation of Bremia lactucae airborne conidia concentration above a lettuce canopy. PLOS ONE 11:e0144573. https://doi.org/10.1371/journal.pone.0144573
Fernández-Rodríguez S, Sadyś M, Smith M, Tormo-Molina R, Skjøth CA, Maya-Manzano JM, Silva-Palacios I, Gonzalo-Garijo Á (2015) Potential sources of airborne Alternaria spp. spores in South-west Spain. Sci Total Environ 533:165–176. https://doi.org/10.1016/j.scitotenv.2015.06.031
Fernando WGD, Paulitz TC, Seaman WL, Dutilleul P, Miller JD (1997) Head blight gradients caused by Gibberella zeae from area sources of inoculum in wheat field plots. Phytopathology 87:414–421. https://doi.org/10.1094/PHYTO.1997.87.4.414
Fitt BDL, Creighton NF, Bainbridge A (1985) Role of wind and rain in dispersal of Botrytis fabae conidia. T Brit Mycol Soc 85(2):307–312. https://doi.org/10.1016/S0007-1536(85)80193-5
Fraaije BA, Cools HJ, Fountaine J, Lovell DJ, Motteram J, West JS, Lucas JA (2005) Role of ascospores in further spread of QoI-resistant cytochrome b alleles (G143A) in field populations of Mycosphaerella graminicola. Phytopathology 95:933–941. https://doi.org/10.1094/PHYTO-95-0933
Friedrich S, Leinhos GME, Löpmeier FJ (2003) Development of ZWIPERO, a model forecasting sporulation and infection periods of onion downy mildew based on meteorological data. Eur J Plant Pathol 109:35–45. https://doi.org/10.1023/A:1022024612563
Gregory PH (1945) The dispersion of airborne spores. Trans Br Mycol Soc 28:26–72. https://doi.org/10.1016/S0007-1536(45)80041-4
Gregory PH (1973) The Microbiology of the Atmosphere, 2nd edn. Leonard Hill, London
Gregory PH, Guthrie EJ, Bunce ME (1959) Experiments on splash dispersal of fungus spores. Microbiology 20 (2):328–354. https://doi.org/10.1099/00221287-20-2-328
González-Fernández E, Piña-Rey A, Fernández-González M, Aira MJ, Rodríguez-Rajo FJ (2020) Identification and evaluation of the main risk periods of Botrytis cinerea infection on grapevine based on phenology, weather conditions and airborne conidia. J Agric Sci 158(1-2):88–98. https://doi.org/10.1017/S0021859620000362
Haudenshield JS, Hartman GL (2011) Exogenous controls increase negative call veracity in multiplexed, quantitative PCR assays for Phakopsora pachyrhizi. Plant Dis 95:343–352. https://doi.org/10.1094/PDIS-01-10-0023
Hellin P, Duvivier M, Dedeurwaerder G, Bataille C, De Proft M, Legrève A (2018) Evaluation of the temporal distribution of Fusarium graminearum airborne inoculum above the wheat canopy and its relationship with Fusarium head blight and DON concentration. Eur J Plant Pathol 151:1049–1064. https://doi.org/10.1007/s10658-018-1442-7
Hellin P, Duvivier M, Clinckemaillie A, Bataille C, Legrève A, Heick TM, Jørgensen LN, Andersson B, Samils B, Rodemann B, Berg G, Kildea S (2020) Multiplex qPCR assay for simultaneous quantification of CYP51-S524T and SdhC-H152R substitutions in European populations of Zymoseptoria tritici. Plant Pathology 69(9):1666–1677. https://doi.org/10.1111/ppa.13252
Hemmati F, Pell JK, Mccartney HA, Deadman ML (2001) Airborne concentrations of conidia of Erynia neoaphidis above cereal fields. Mycol Res 105:485–489. https://doi.org/10.1017/S0953756201003537
Hildebrand PD, Sutton JC (1982) Weather variables in relation to an epidemic of onion downy mildew. Phytopathology 72:219–224. https://doi.org/10.1094/Phyto-72-219
Hirst JM (1953) Changes in atmospheric spore content: Diurnal periodicity and the effects of weather. T Brit Mycol Soc 36:375–IN378. https://doi.org/10.1016/S0007-1536(53)80034-3
Hu Y, Green GS, Milgate AW, Stone EA, Rathjen JP, Schwessinger B (2019) Pathogen detection and microbiome analysis of infected wheat using a portable DNA sequencer. Phytobiomes J 3:92–101. https://doi.org/10.1094/PBIOMES-01-19-0004-R
Ingold CT (1971) Fungal spores: their liberation and dispersal. Oxford University Press, London
Ingold CT (1999) Active liberation of reproductive units in terrestrial fungi. Mycologist 13(3):113–116. https://doi.org/10.1016/S0269-915X(99)80040-8
Isard SA, Barnes CW, Hambleton S, Ariatti A, Russo JM, Tenuta A, Gay DA, Szabo LJ (2011) Predicting soybean rust incursions into the North American continental interior using crop monitoring, spore trapping, and aerobiological modeling. Plant Dis 95:1346–1357. https://doi.org/10.1094/PDIS-01-11-0034
Jackson SL, Bayliss KL (2011) Spore traps need improvement to fulfil plant biosecurity requirements. Plant Pathol 60(5):801–810. https://doi.org/10.1111/j.1365-3059.2011.02445.x
Jarvis WR (1962) The dispersal of spores of Botrytis cinerea fr. in a raspberry plantation. Trans Brit Mycol Society 45:549–559. https://doi.org/10.1016/S0007-1536(62)80015-1
Jones SJ, Gent DH, Pethybridge SJ, Hay FS (2011) Spatial characteristics of white mould epidemics and the development of sequential sampling plans in Australian bean fields. Plant Pathol 60:1169–1182. https://doi.org/10.1111/j.1365-3059.2011.02466.x
Kennedy R, Wakeham AJ (2008) Development of detection systems for the sporangia of Peronospora destructor. Eur J Plant Pathol 122:147–155. https://doi.org/10.1007/s10658-008-9346-6
Klosterman SJ, Anchieta A, McRoberts N, Koike ST, Subbarao KV, Voglmayr H, Choi YJ, Thines M, Martin FN (2014) Coupling spore traps and quantitative PCR assays for detection of the downy mildew pathogens of spinach (Peronospora effusa) and beet (P. schachtii). Phytopathology 104:1349–1359. https://doi.org/10.1094/PHYTO-02-14-0054-R
Kunjeti SG, Anchieta A, Martin FN, Choi YJ, Thines M, Michelmore RW, Koike ST, Tsuchida C, Mahaffee W, Subbarao KV, Klosterman SJ (2016) Detection and quantification of Bremia lactucae by spore trapping and quantitative PCR. Phytopathology 106:1426–1437. https://doi.org/10.1094/PHYTO-03-16-0143-R
Lacey J (1996) Spore dispersal-its role in ecology and disease: the British contribution to fungal aerobiology. Mycol Res 100:641–660. https://doi.org/10.1016/S0953-7562(96)80194-8
Lacey ME, West J (2006) The air spora: a manual for catching and identifying airborne biological particles. Springer, Boston, MA. https://doi.org/10.1007/978-0-387-30253-9
Leach CM (1982) Active sporangium discharge by Peronospora destructor. Phytopathology 72:881–885. https://doi.org/10.1094/Phyto-72-881
Leach CM, Hildebrand PD, Sutton JC (1982) Sporangium discharge by Peronospora destructor: influence of humidity, red-infrared radiation, and vibration. Phytopathology 72:1052–1056. https://doi.org/10.1094/Phyto-72-1052
Lees A, Roberts DM, Lynott J, Sullivan L, Brierley J (2019) Real-Time PCR and LAMP assays for the detection of spores of Alternaria solani and sporangia of Phytophthora infestans to inform disease risk forecasting. Plant Dis 103(12):3172–3180. https://doi.org/10.1094/PDIS-04-19-0765-RE
Legg BJ (1983) Movement of plant pathogens in the crop canopy. Philos Trans R Soc Lond B Biol Sci 302:559–574. https://doi.org/10.1098/rstb.1983.0075
Limpert E, Godet F, Müller K (1999) Dispersion of cereal mildews across Europe. Agr Forest Meteorol 97:293–308. https://doi.org/10.1016/S0168-1923(99)00073-8
Madden LV, Hughes G (1999) Sampling for plant disease incidence. Phytopathology 89:1088–1103. https://doi.org/10.1094/PHYTO.1999.89.11.1088
Mahaffee WF, Stoll R (2016) The ebb and flow of airborne pathogens: Monitoring and use in disease management decisions. Phytopathology 106(5):420–431. https://doi.org/10.1094/PHYTO-02-16-0060-RVW
McCartney HA, Fitt BDL, Schmechel D (1997) Sampling bioaerosols in plant pathology. J Aerosol Sci 28(3):349–364. https://doi.org/10.1016/S0021-8502(96)00438-7
McDevitt JJ, Lees PSJ, Merz WG, Schwab KJ (2007) Inhibition of quantitative PCR analysis of fungal conidia associated with indoor air particulate matter. Aerobiologia 23(1):35–45. https://doi.org/10.1007/s10453-006-9047-6
Micheli PA (1729) Nova Plantarum Genera. 234 pp.
Morais D, Sache I, Suffert F, Laval V (2016) Is the onset of septoria tritici blotch epidemics related to the local pool of ascospores? Plant Pathology 65(2):250–260. https://doi.org/10.1111/ppa.12408
Mundt CC (2009) Importance of autoinfection to the epidemiology of polycyclic foliar disease. Phytopathology 99:1116–1120. https://doi.org/10.1094/PHYTO-99-10-1116
Mundt C, Browning J (1985) Development of crown rust epidemics in genetically diverse oat populations: effect of genotype unit area. Phytopathology 75:607–610. https://doi.org/10.1094/Phyto-75-607
Narayan P, Gonzalez F, Zeller L, Walker R (2010) Final Report - Flying Spore Traps Project CRC30032 (with Ground Vehicle Test Results) ACT. CRC for National Plant Biosecurity, Australia
Neufeld KN, Isard SA, Ojiambo PS (2013) Relationship between disease severity and escape of Pseudoperonospora cubensis sporangia from a cucumber canopy during downy mildew epidemics. Plant Pathology 62(6):1366–1377. https://doi.org/10.1111/ppa.12040
Nicolaisen M, West JS, Sapkota R, Canning GGM, Schoen C, Justesen AF (2017) Fungal communities including plant pathogens in near surface air are similar across northwestern Europe. Frontiers in Microbiology 8(1729). https://doi.org/10.3389/fmicb.2017.01729
Nieguitsila A, Arné P, Durand B, Deville M, Benoît-Valiergue H, Chermette R, Cottenot-Latouche S, Guillot J (2011) Relative efficiencies of two air sampling methods and three culture conditions for the assessment of airborne culturable fungi in a poultry farmhouse in France. Envion Res 111:248–253. https://doi.org/10.1016/j.envres.2010.12.005
Ovaskainen O, Abrego N, Somervuo P, Palorinne I, Hardwick B, Pitkänen J-M, Andrew NR, Niklaus PA, Schmidt NM, Seibold S, Vogt J, Zakharov EV, Hebert PDN, Roslin T, Ivanova NV (2020) Monitoring Fungal Communities With the Global Spore Sampling Project. Frontiers in Ecology and Evolution 7(511). https://doi.org/10.3389/fevo.2019.00511
Paul PA, El-Allaf SM, Lipps PE, Madden LV (2004) Rain splash dispersion of Gibberella zeae within wheat canopies in Ohio. Phytopathology 94:1342–1349. https://doi.org/10.1094/PHYTO.2004.94.12.1342
Paulitz TC, Dutilleul P, Yamasaki SH, Fernando WGD, Seaman WL (1999) A generalized two-dimensional Gaussian model of disease foci of head blight of wheat caused by Gibberella zeae. Phytopathology 89:74–83. https://doi.org/10.1094/PHYTO.1999.89.1.74
Peters RD, Al-Mughrabi KI, Kalischuk ML, Dobinson KF, Conn KL, Alkher H, Islam MR, Daayf F, Lynn J, Bizimungu B, De Koeyer D, Lévesque CA, Kawchuk LM (2014) Characterization of Phytophthora infestans population diversity in Canada reveals increased migration and genotype recombination. Canadian Journal of Plant Pathol 36(1):73–82. https://doi.org/10.1080/07060661.2014.892900
Rahman A, Standish J, D'Arcangelo KN, Quesada-Ocampo LM (2020) Clade-specific biosurveillance of Pseudoperonospora cubensis using spore traps for precision disease management of cucurbit downy mildew. Phytopathology. 111:312–320. https://doi.org/10.1094/PHYTO-06-20-0231-R
Redondo MA, Berlin A, Boberg J, Oliva J (2020) Vegetation type determines spore deposition within a forest–agricultural mosaic landscape. FEMS Microbiol Ecol 96(Issue 6):fiaa082. https://doi.org/10.1093/femsec/fiaa082
Reich J, Chatterton S, Johnson D (2016) Temporal dynamics of Botrytis cinerea and Sclerotinia sclerotiorum in seed Alfalfa fields of southern Alberta, Canada. Plant Dis 101:331–343. https://doi.org/10.1094/PDIS-04-16-0492-RE
Rieux A, Soubeyrand S, Bonnot F, Klein EK, Ngando JE, Mehl A, Ravigne V, Carlier J, de Lapeyre de Bellaire L (2014) Long-distance wind-dispersal of spores in a fungal plant pathogen: Estimation of anisotropic dispersal kernels from an extensive field experiment. PLoS ONE 9(8):e103225. https://doi.org/10.1371/journal.pone.0103225
Robinson RA (1976) Plant pathosystems. Springer-Verlag Berlin Heidelberg. https://doi.org/10.1007/978-3-642-66359-8.
Rogers SL, Atkins SD, West JS (2009) Detection and quantification of airborne inoculum of Sclerotinia sclerotiorum using quantitative PCR. Plant Pathol. 58:324–331. https://doi.org/10.1111/j.1365-3059.2008.01945.x
Schwarzbach E (1979) A High Throughput Jet Trap for Collecting Mildew Spores on Living Leaves. J Phytopathol 94(2):165–171. https://doi.org/10.1111/j.1439-0434.1979.tb01546.x
Severns PM, Sackett KE, Farber DH, Mundt CC (2018) Consequences of long-distance dispersal for epidemic spread: patterns, scaling, and mitigation. Plant Dis 103(2):177–191. https://doi.org/10.1094/PDIS-03-18-0505-FE
Si Ammour M, Bilodeau GJ, Tremblay DM, van der Heyden H, Yaseen T, Varvaro L, Carisse O (2017) Development of real-time isothermal amplification assays for on-site detection of Phytophthora infestans in potato leaves. Plant Dis 101:1269–1277. https://doi.org/10.1094/PDIS-12-16-1780-RE
Stakman EC, Christensen CM (1946) Aerobiology in relation to plant disease. Bot Rev 12(4):205–253. http://www.jstor.org/stable/4353339
Strandberg JO (1977) Spore production and dispersal of Alternaria dauci. Phytopathology 67:1262–1266. https://doi.org/10.1094/Phyto-67-1262
Suarez MB, Walsh K, Boonham N, O’Neill T, Pearson S, Barker I (2005) Development of real-time PCR (TaqMan®) assays for the detection and quantification of Botrytis cinerea in planta. Plant Physiol Bioch 43:890–899. https://doi.org/10.1016/j.plaphy.2005.07.003
Summers CF, Adair NL, Gent DH, McGrath MT, Smart CD (2015) Pseudoperonospora cubensis and P. humuli detection using species-specific probes and high definition melt curve analysis. Canadian Journal of Plant Pathol 37:315–330. https://doi.org/10.1080/07060661.2015.1053989
Thiessen LD, Keune JA, Neill TM, Turechek WW, Grove GG, Mahaffee WF (2016) Development of a grower-conducted inoculum detection assay for management of grape powdery mildew. Plant Pathol 65:238–249. https://doi.org/10.1111/ppa.12421
Tremblay ED, Duceppe M-O, Berube JA, Kimoto T, Lemieux C, Bilodeau GJ (2018) Screening for exotic forest pathogens to increase survey capacity using metagenomics. Phytopathology 108(12):1509–1521. https://doi.org/10.1094/PHYTO-02-18-0028-R
Turechek WW, Mahaffee WF (2004) Spatial pattern analysis of hop powdery mildew in the Pacific Northwest: Implications for sampling. Phytopathology 94:1116–1128. https://doi.org/10.1094/PHYTO.2004.94.10.1116
Van der Heyden H, Carisse O, Brodeur L (2012) Comparison of monitoring based indicators for initiating fungicide spray programs to control Botrytis leaf blight of onion. Crop Prot 33(0):21-28. https://doi.org/10.1016/j.cropro.2011.11.008.
Van der Heyden H, Dutilleul P, Charron J-B, Bilodeau GJ, Carisse O (2020) Factors Influencing the Occurrence of Onion Downy Mildew (Peronospora destructor) Epidemics: Trends from 31 Years of Observational Data. Agronomy 10(5):738. https://doi.org/10.3390/agronomy10050738
Van der Heyden H, Dutilleul P, Brodeur L, Carisse O (2014) Spatial distribution of single-nucleotide polymorphisms related to fungicide resistance and implications for sampling. Phytopathology 104:604–613. https://doi.org/10.1094/PHYTO-03-13-0085-R
von Qualen R, Yang X-B (2006) Spore traps help researchers watch for soybean rust. http://lib.dr.iastate.edu/cropnews/1308
Wallace EC, D’Arcangelo KN, Quesada-Ocampo LM (2020) Population Analyses Reveal Two Host-Adapted Clades of Pseudoperonospora cubensis, the Causal Agent of Cucurbit Downy Mildew, on Commercial and Wild Cucurbits. Phytopathology 110(9):1578–1587. https://doi.org/10.1094/PHYTO-01-20-0009-R
West JS, Kimber RBE (2015) Innovations in air sampling to detect plant pathogens. Ann. Appl. Biol 166:4–17. https://doi.org/10.1111/aab.12191
West JS, Atkins SD, Emberlin J, Fitt BDL (2008) PCR to predict risk of airborne disease. Trends Microbiol 16(8):380–387. https://doi.org/10.1016/j.tim.2008.05.004
West JS, Canning K, King K, Fraaije B, Wili S (2018) Final Report - Arable crop disease alert system, PR594. AHDB Cereals & Oilseeds.
Widmark AK, Andersson B, Cassel-Lundhagen A, Sandström M, Yuen JE (2007) Phytophthora infestans in a single field in southwest Sweden early in spring: symptoms, spatial distribution and genotypic variation. Plant Pathology 56(4):573–579. https://doi.org/10.1111/j.1365-3059.2007.01618.x
Williams RH, Ward E, McCartney HA (2001) Methods for integrated air sampling and DNA analysis for detection of airborne fungal pores. Appl Environ Microb 67(6):2453–2459. https://doi.org/10.1128/AEM.67.6.2453-2459.2001
Zawolek M, Zadoks J (1992) Studies in focus development: An optimum for the dual dispersal of plant pathogens. Phytopathology 82:1288–1297. https://doi.org/10.1094/Phyto-82-1288
Zhan J, Kema GHJ, Waalwijk C, McDonald BA (2002) Distribution of mating type alleles in the wheat pathogen Mycosphaerella graminicola over spatial scales from lesions to continents. Fungal Genetics and Biology 36(2):128–136. https://doi.org/10.1016/S1087-1845(02)00013-0
Acknowledgements
We would like to thank the members of the MAPAQ advisory committee for their comments and suggestions. We are also thankful to Agriculture and Agri-Food Canada for the implication of Dr. Odile Carisse to this work. Finally, we wish to acknowledge the anonymous reviewers for their useful and constructive comments and suggestions.
Funding
This work was carried out under component 3.2 of the Prime-Vert program (2013–2018) and received financial assistance from the Ministère de l’Agriculture, des Pêcheries et de l’Alimentation (MAPAQ) through the Green Fund (grant number: PV-3.2-PHYT-22).
Author information
Authors and Affiliations
Contributions
The idea behind this literature review came from H.V.D.H. and O.C.; the literature search and the preparation of the draft manuscript was carried out by H.V.D.H. and O.C. Critical review and exhaustive editing were done by P.D. The final version of the manuscript was revised by J-B.C. and G.B. prior to its submission.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Van der Heyden, H., Dutilleul, P., Charron, JB. et al. Monitoring airborne inoculum for improved plant disease management. A review. Agron. Sustain. Dev. 41, 40 (2021). https://doi.org/10.1007/s13593-021-00694-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s13593-021-00694-z