Abstract
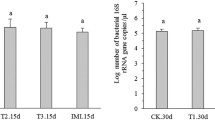
Honeybees are subjected to direct contact with transgenic maize pollen due to their feeding activities on pollen. The potential side effects of transgenic cry1Ah-maize pollen on the midgut bacteria of the larvae and worker bees of Apis mellifera ligustica were investigated through denaturing gradient gel electrophoresis under both laboratory and field conditions. Newly emerged bees were fed transgenic cry1Ah-maize pollen, normal maize pollen, linear cry1Ah gene DNA, supercoiled plasmid DNA, and sugar syrup under the laboratory conditions. The results showed that there were no significant differences in the midgut bacterial community composition among the five treatments. No significant effects were observed in the midgut communities between larvae and adult honeybees fed transgenic cry1Ah-maize pollen and non-transgenic maize pollen in the field trials.







Similar content being viewed by others
References
Al-Deeb, M.A., Wilde, G.E., Blair, J.M., Todd, T.C. (2003) Effect of Bt corn for corn rootworm control on nontarget soil microarthropods and nematodes. Environ. Entomol. 32, 859–865
Babendreier, D., Joller, D., Romeis, J., Bigler, F., Widmer, F. (2007) Bacterial community structures in honeybee intestines and their response to two insecticidal proteins. FEMS Microbiol. Ecol. 59, 600–610
Baumgarte, S., Tebbe, C.C. (2005) Field studies on the environmental fate of the Cry1Ab Bt-toxin produced by transgenic maize (MON810) and its effect on bacterial communities in the maize rhizosphere. Mol. Ecol. 14, 2539–2551
Blackwood, C.B., Buyer, J.S. (2004) Soil microbial communities associated with Bt and non-Bt corn in three soils. J. Environ. Qual. 33, 832–836
Clive J. (2012) Global Status of Commercialized Biotech/GM Crops: 2011. ISAAA Brief No. 43-2011
Cox-Foster, D.L., Conlan, S., Holmes, E.C., Palacios, G., Evans, J.D., Moran, N.A., Quan, P.L., Briese, T., Hornig, M., Geiser, D.M. (2007) A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318, 283–287
Dai, P.L., Zhou, W., Zhang, J., Jiang, W.Y., Wang, Q., Cui, H.J., Sun, J.H., Wu, Y.Y., Zhou, T. (2012a) The effects of Bt Cry1Ah toxin on worker honeybees (Apis mellifera ligustica and Apis cerana cerana). Apidologie 43, 384–391
Dai, P.L., Zhou, W., Zhang, J., Cui, H.J., Wang, Q., Jiang, W.Y., Sun, J.H., Wu, Y.Y., Zhou, T. (2012b) Field assessment of Bt cry1Ah corn pollen on the survival, development and behavior of Apis mellifera ligustica. Ecotoxicol. Environ. Saf. 79, 232–237
Daudu, C.K., Muchaonyerwa, P., Mnkeni, P.N.S. (2009) Litterbag decomposition of genetically modified maize residues and their constituent Bacillus thuringiensis protein (Cry1Ab) under field conditions in the central region of the Eastern Cape, South Africa. Agric. Ecosyst. Environ. 134, 153–158
Duan, J.J., Marvier, M., Huesing, J., Dively, G., Huang, Z.Y. (2008) A meta-analysis of effects of Bt crops on honey bees (Hymenoptera: Apidae). PLoS One 3, e1415
Gathmann, A., Wirooks, L., Hothorn, L.A., Bartsch, D., Schuphan, I. (2006) Impact of Bt maize pollen (MON810) on lepidopteran larvae living on accompanying weeds. Mol. Ecol. 15, 2677–2685
Gruber, H., Paul, V., Meyer, H.H.D., Müller, M.M. (2012) Determination of insecticidal Cry1Ab protein in soil collected in the final growing seasons of a nine-year field trial of Bt-maize MON810. Transgenic Res. 21, 77–78
Hendriksma, H.P., Härtel, S., Steffan-Dewenter, I. (2011) Testing pollen of single and stacked insect-resistant Bt-maize on in vitro reared honey bee larvae. PLoS One 6, e28174
Higgins, L.S., Babcock, J., Neese, P., Layton, R.J., Moellenbeck, D.J., Storer, N. (2009) Three-year field monitoring of Cry1F, event DAS-Ø15Ø7-1, maize hybrids for nontarget arthropod effects. Environ. Entomol. 38, 281–292
Lima, M.A.P., Pires, C.S.S., Guedes, R.N.C., Nakasu, E.Y.T., Lara, M.S., Fontes, E.M.G., Sujii, E.R., Dias, S.C., Campos, L.A.O. (2011) Does Cry1Ac Bt-toxin impair development of worker larvae of Africanized honey bee? J. Appl. Entomol. 135, 415–422
Malone, L.A., Todd, J.H., Burgess, E.P.J., Christeller, J.T. (2004) Development of hypopharyngeal glands in adult honey bees fed with a Bt toxin, a biotin-binding protein and a protease inhibitor. Apidologie 35, 655–664
Martinson, V.G., Danforth, B.N., Minckley, R.L., Rueppell, O., Tingek, S., Moran, N.A. (2011) A simple and distinctive microbiota associated with honey bees and bumble bees. Mol. Ecol. 20, 619–628
Mohr, K.I., Tebbe, C.C. (2006) Diversity and phylotype consistency of bacteria in the guts of three bee species (Apoidea) at an oilseed rape field. Environ. Microbiol. 8, 258–272
Mohr, K., Tebbe, C. (2007) Field study results on the probability and risk of a horizontal gene transfer from transgenic herbicide-resistant oilseed rape pollen to gut bacteria of bees. Appl. Microbiol. Biot. 75, 573–582
Nakatsu, C.H., Torsvik, V., Ovreas, L. (2000) Soil community analysis using DGGE of 16 s rDNA polymerase chain reaction products. Soil Sci. Soc. Am. J. 64, 1382–1388
Prischl, M., Hackl, E., Pastar, M., Pfeiffer, S., Sessitsch, A. (2012) Genetically modified Bt maize lines containing cry3Bb1, cry1A105 or cry1Ab2 do not affect the structure and functioning of root-associated endophyte communities. Appl. Soil Ecol. 54, 39–48
Ramirez-Romero, R., Chaufaux, J., Pham-Delegue, M. (2005) Effects of Cry1Ab protoxin, deltamethrin and imidacloprid on the foraging activity and the learning performances of the honeybee Apis mellifera, a comparative approach. Apidologie 36, 601–611
Romeis, J., Bartsch, D., Bigler, F., Candolfi, M.P., Gielkens, M.M.C., et al. (2008) Assessment of risk of insect-resistant transgenic crops to nontarget arthropods. Nat. Biotechnol. 26, 203–208
Rose, R., Dively, G.P., Pettis, J. (2007) Effects of Bt corn pollen on honey bees: emphasis on protocol development. Apidologie 38, 368–377
Shan, G., Embrey, S.K., Herman, R.A., McCormick, R. (2008) Cry1F protein not detected in soil after three years of transgenic Bt corn (1507 corn) use. Environ. Entomol. 37, 255–262
Wang Y.F. (2010) Field trails and genetic stability analysis of insect resistant transgenic Bt maize. Ph.D. dissertation, Northeast Agricultural University, Heilongjiang
Wang, Y.B., Lang, Z., Zhang, J., He, K.L., Song, F.P., Huang, D.F. (2008) Ubi1 intron-mediated enhancement of the expression of Bt cry1Ah gene in transgenic maize (Zea mays L.). Chin. Sci. Bull. 53, 3185–3190
Xue, J., Liang, G., Crickmore, N., Li, H., He, K., Song, F., Feng, X., Huang, D., Zhang, J. (2008) Cloning and characterization of a novel Cry1A toxin from Bacillus thuringiensis with high toxicity to the Asian corn borer and other lepidopteran insects. FEMS Microbiol. Lett. 280, 95–101
Yoshiyama, M., Kimura, K. (2009) Bacteria in the gut of Japanese honeybee, Apis cerana japonica, and their antagonistic effect against Paenibacillus larvae, the causal agent of American foulbrood. J. Invertebr. Pathol. 102, 91–96
Zwahlen, C., Hilbeck, A., Nentwig, W. (2007) Field decomposition of transgenic Bt maize residue and the impact on non-target soil invertebrates. Plant Soil 300, 245–257
Acknowledgments
This work was supported by grants from the Major State Basic Research Development Program of China (973 Program; No. 2009CB118902) and the National Transgenic Major Program of China (No. 2009ZX08011-016B).
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuscript editor: Monique Gauthier
L’influence du pollen de maïs-Bt transgénique sur la diversité bactérienne de l’intestin moyen d’ Apis mellifera ligustica
Maïs transgénique / Apis mellifera / flore intestinale / électrophorèse sur gel à gradient dénaturant
Der Einfluss von Bt-transgenem Maispollen auf die bakterielle Diversität im Mitteldarm von Apis mellifera ligustica
transgener Mais / Apis mellifera / Darmflora / denaturierende Gradientengelelektrophorese
Li-Li Geng and Hong-Juan Cui contributed equally to this work.
Rights and permissions
About this article
Cite this article
Geng, LL., Cui, HJ., Dai, PL. et al. The influence of Bt-transgenic maize pollen on the bacterial diversity in the midgut of Apis mellifera ligustica . Apidologie 44, 198–208 (2013). https://doi.org/10.1007/s13592-012-0171-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-012-0171-8